-
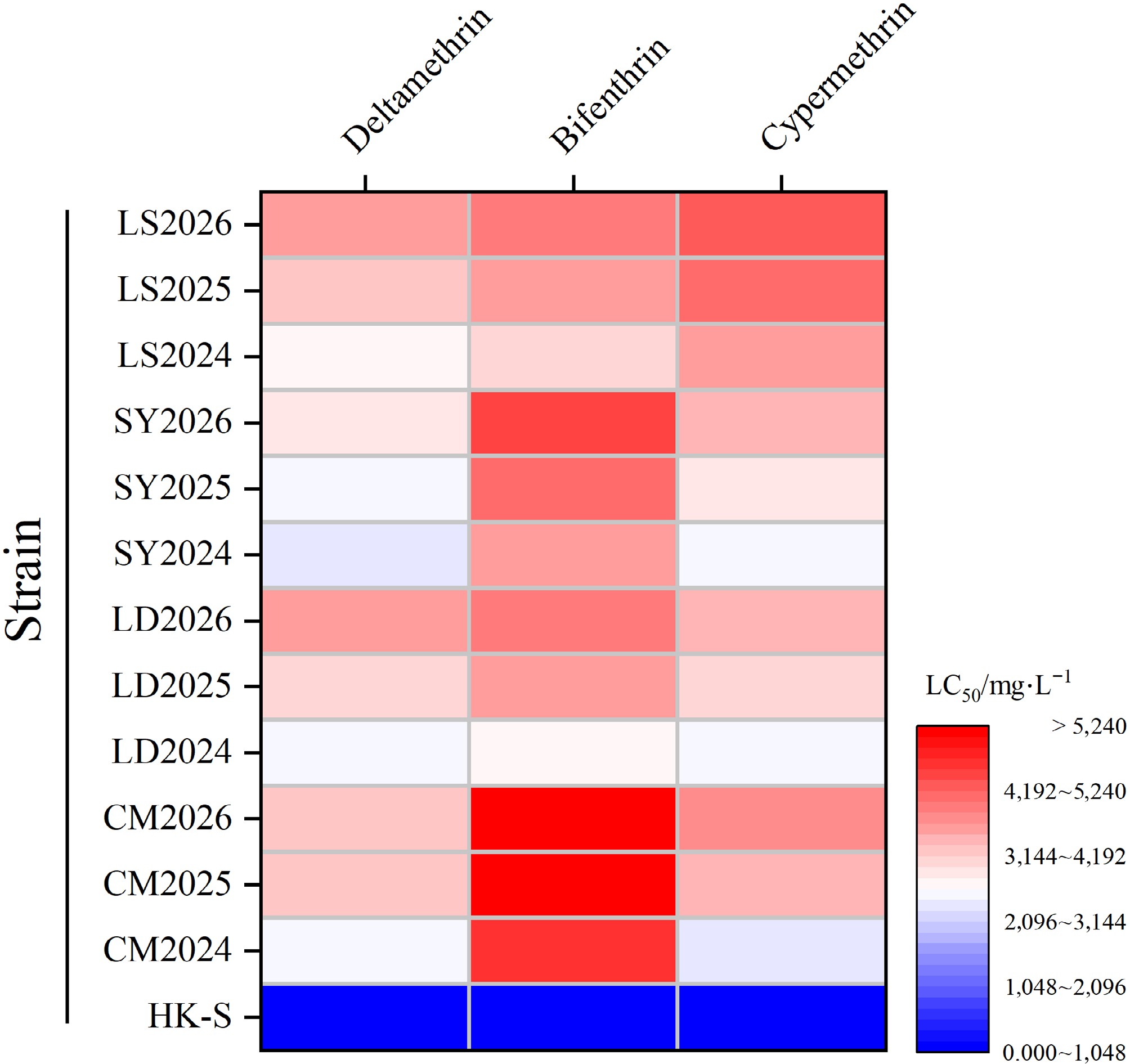
Figure 1.
2024−2026 resistance levels of M. usitatus strains.
-
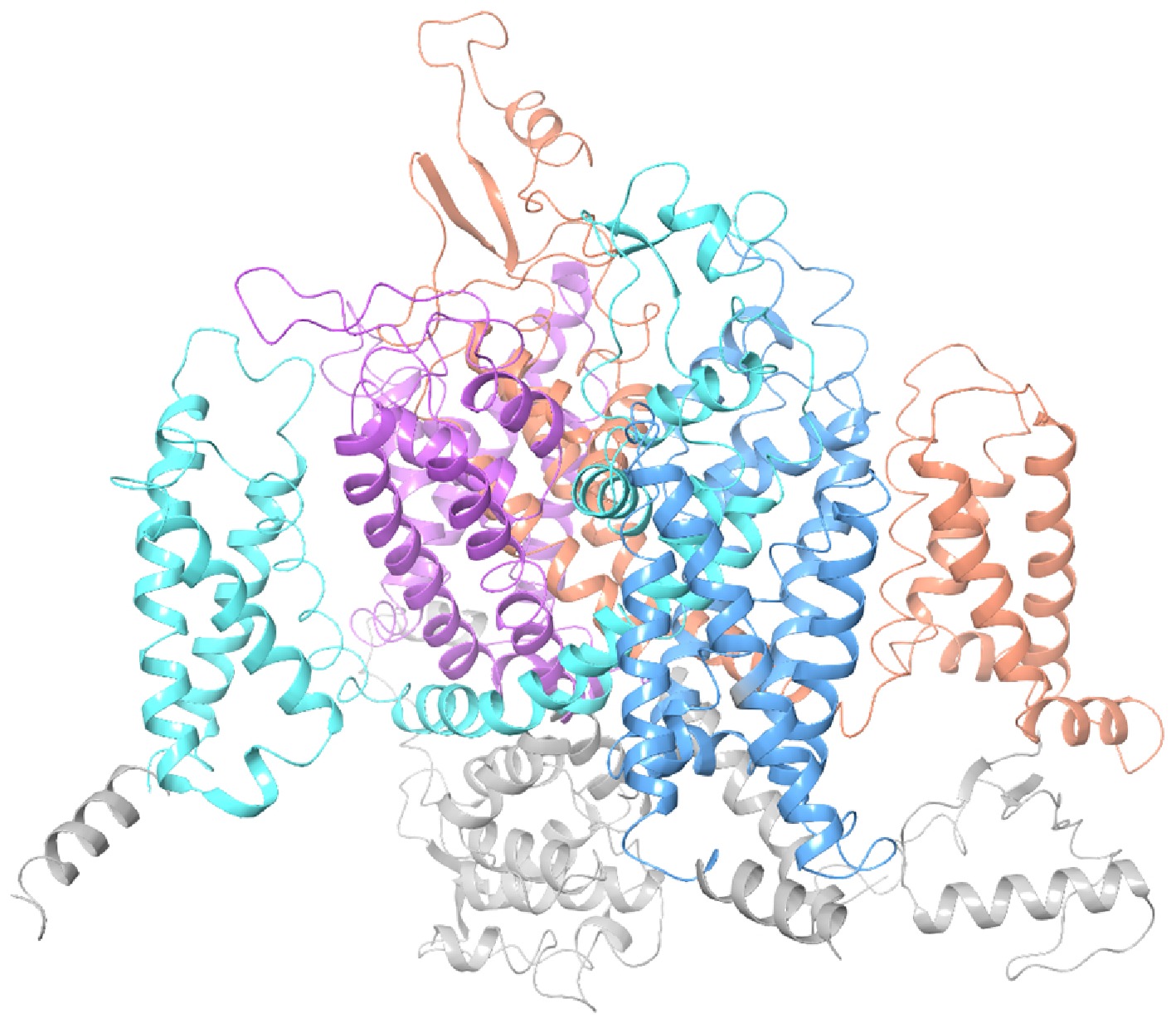
Figure 2.
3D model of M. usitatus VGSC (orange area: Domain I, blue area: Domain II, light blue area: Domain III, purple area: Domain IV).
-
Figure 3.
Topological structure diagram of the VGSC M918L/T, T929I and L1014F mutation sites in the M. usitatus.
-
Strains Collection site Year Longitude and latitude Host CM-R Chengmai County, Hainan Province 2024−2026 N 20°20'31", E 110°30'11 Cowpea LD-R Ledong Li Autonomous County, Hainan Province 2024−2026 N 18°40'03", E 109°30'25" Cowpea SY-R Yazhou District, Sanya City, Hainan Province 2024−2026 N 18°23'05", E 109°9'44" Cowpea LS-R Lingshui Li Autonomous County, Hainan Province 2024−2026 N 18°31'42", E 110°1'35" Cowpea Table 1.
Field collection information table of M. usitatus.
-
Primer name Primer sequence Usage MuNav-5(F) TTGGCAGTCAATTCCCCGGGGCCACCATGCCGAGTGTCCGGGAGTCG Full-length cloning of VGSC MuNav-3(R) AGTCGTAACCAGATCAAGCTTTCAGACATCCGCAAGGGCCGGAG Domain II-5(F) ACATTTTCTGCGTGTGGGACTGC Detection of mutation sites in Domain II Domain II-5(R) CCAGTTGATAAATCGTGATATTC Table 2.
Primers for the full-length application and mutation frequency detection of VGSC in M. usitatus.
-
Treatment Strain Sample No. Slope ± standard error Degree of freedom Chi-square LC50 (95% confidence interval) Resistance ratio Deltamethrin HK-S 444 1.975 ± 0.239 4 0.279 8.148 (5.853−10.311) 1 CM2024 461 2.545 ± 0.016 4 4.864 2,489.687 (1,473.860−4,688.216) 305.6 CM2025 450 5.197 ± 0.571 4 0.173 3,181.355 (2,890.989−3,514.066) 390.4 CM2026 449 3.920 ± 0.433 4 0.210 3,302.052 (2,945.182−3,755.924) 405.3 LD2024 445 1.162 ± 0.133 4 7.676 2,579.637 (2,469.245−3,151.628) 316.6 LD2025 441 1.096 ± 0.217 4 3.294 3,102.792 (1,902.736−5,426.599) 380.8 LD2026 450 2.947 ± 0.329 4 0.091 3,573.267 (3,090.145−4,270.751) 438.6 SY2024 438 0.905 ± 0.127 4 0.890 2,409.838 (1,472.794−3,195.164) 295.8 SY2025 446 1.477 ± 0.073 4 1.011 2,583.096 (1,401.633−5,023.782) 317.0 SY2026 447 3.102 ± 0.308 4 1.268 2,962.999 (2,131.597−4,757.661) 363.6 LS2024 443 2.190 ± 0.169 4 3.672 2,699.051 (2,730.686−2,857.614) 331.3 LS2025 440 2.174 ± 0.228 4 1.806 3,276.117 (1,396.584−6,403.851) 402.1 LS2026 448 2.952 ± 0.316 4 1.281 3,553.753 (2,280.381−4,217.518) 436.2 Resistance fold = Field population LC50/Sensitive strain (HK-S) LC50. HK-S is an indoor strain that has not been exposed to insecticides for an extended period, specifically from 2019 to 2025, and its LC50 serves as the fixed benchmark for all comparisons. Table 3.
Resistance levels of M. usitatus to Deltamethrin.
-
Treatment Strain Sample No. Slope ± standard error Degree of freedom Chi-square LC50 (95% confidence interval) Resistance ratio Bifenthrin HK-S 444 2.248 ± 0.293 4 0.461 7.610 (5.501−9.515) 1 CM2024 446 0.810 ± 0.120 4 2.060 4,642.120 (2,744.790−5,345.450) 610.0 CM2025 445 2.746 ± 0.386 4 0.704 5,154.927 (4,253.658−6,853.767) 677.4 CM2026 450 2.233 ± 0.291 4 0.461 5,222.599 (4,167.747−7,257.130) 686.3 LD2024 441 0.713 ± 0.124 4 0.154 2,710.853 (1,747.671−5,465.877) 356.2 LD2025 429 1.541 ± 0.155 4 1.045 3,508.291 (2,491.721−7,786.022) 461.0 LD2026 449 3.409 ± 0.416 4 0.428 3,964.617 (3,461.532−4,705.489) 521.0 SY2024 438 0.801 ± 0.129 4 0.025 3,602.825 (2,246.790−7,576.970) 473.4 SY2025 430 1.206 ± 0.022 4 2.964 4,127.643 (2,578.660−7,906.107) 542.4 SY2026 448 3.059 ± 0.397 4 0.495 4,440.504 (3,790.218−5,507.876) 583.5 LS2024 446 2.902 ± 0.222 4 7.844 2,999.842 (2,771.627−3,289.249) 394.2 LS2025 450 1.794 ± 0.067 4 2.454 3,601.884 (2,391.087−6,201.144) 473.3 LS2026 449 4.741 ± 0.580 4 4.453 3,946.913 (3,371.774−4,824.179) 518.7 Resistance fold = Field population LC50/Sensitive strain (HK-S) LC50. HK-S is an indoor strain that has not been exposed to insecticides for an extended period, specifically from 2019 to 2025, and its LC50 serves as the fixed benchmark for all comparisons. Table 4.
Resistance levels of M. usitatus to Bifenthrin.
-
Treatment Strain Sample No. Slope ± standard error Degree of freedom Chi-square LC50 (95% confidence interval) Resistance ratio Cypermethrin HK-S 447 1.730 ± 0.222 4 0.566 6.756 (4.387−9.029) 1 CM2024 425 2.871 ± 0.062 4 2.524 2,380.572 (1,209.860−4,688.216) 352.4 CM2025 450 2.102 ± 0.219 4 0.425 3,342.027 (2,773.125−4,222.895) 494.7 CM2026 450 1.651 ± 0.179 4 0.081 3,738.598 (2,939.492−5,142.503) 553.4 LD2024 445 2.203 ± 0.187 4 1.724 2,461.168 (2,390.935−2,937.372) 364.3 LD2025 445 2.017 ± 0.884 4 1.027 3,055.231 (1,491.087−4,923.077) 452.2 LD2026 450 2.250 ± 0.262 4 2.250 3,319.287 (2,828.367−4,040.714) 491.3 SY2024 437 0.866 ± 0.125 4 0.816 2,459.628 (1,644.078−4,515.853) 364.1 SY2025 429 2.151 ± 0.017 4 1.739 2,963.782 (1,708.851−4,996.745) 438.7 SY2026 448 2.970 ± 0.319 4 1.205 3,359.198 (2,913.821−3,984.137) 497.2 LS2024 441 0.891 ± 0.125 4 0.939 3,652.185 (2,219.346−8,724.188) 540.6 LS2025 448 2.966 ± 0.693 4 1.831 4,102.857 (3,201.822−7,012.696) 607.3 LS2026 449 3.874 ± 0.509 4 0.535 4,314.532 (3,795.844−5,094.333) 638.6 Resistance fold = Field population LC50/Sensitive strain (HK-S) LC50. HK-S is an indoor strain that has not been exposed to insecticides for an extended period, specifically from 2019 to 2025, and its LC50 serves as the fixed benchmark for all comparisons. Table 5.
Resistance levels of M. usitatus to Cypermethrin.
-
Strain Year M918L Mutation frequency M918T Mutation frequency T929I Mutation frequency L1014F Mutation frequency Total samples HK-S 2025 M/L 0 M/T 0 T/I 0 L/F 0 29 CM2024 2024 M/L 0 M/T 0 T/I 100 L/F 0 29 CM2025 2025 M/L 0 M/T 0 T/I 93.3 L/F 0 30 CM2026 2026 M/L 0 M/T 0 T/I 90.0 L/F 0 30 LD2024 2024 M/L 0 M/T 0 T/I 100 L/F 0 30 LD2025 2025 M/L 0 M/T 0 T/I 100 L/F 0 30 LD2026 2026 M/L 0 M/T 0 T/I 96.7 L/F 0 30 SY2024 2024 M/L 0 M/T 0 T/I 100 L/F 0 29 SY2025 2025 M/L 0 M/T 0 T/I 100 L/F 0 28 SY2026 2026 M/L 0 M/T 0 T/I 96.7 L/F 0 30 LS2024 2024 M/L 0 M/T 0 T/I 100.0 L/F 0 29 LS2025 2025 M/L 0 M/T 0 T/I 90.0 L/F 0 30 LS2026 2026 M/L 0 M/T 0 T/I 90.0 L/F 0 30 To ensure the reliability and accuracy of the experimental data, samples that did not meet quality standards were excluded from the final statistical analysis. Table 6.
Mutation frequency of M. usitatus M918L/T, L1014F, and T929I mutation sites.
-
Channel Segment MuNav1-1 IL45-S5 252VPGLKTIVGA VI ESVKNLRDVI ILTMFSLSVF ALMGLQIY IIL45-S5 934WPTLNLLISI MG RTMGALGNLT FVLCIIIFIF AVMGMQLF IIIL45-S5 1431MQGMRVVVNA LV QAIPSIFNVL LVCLIFWLIF AIMGVQLF IVL45-S5 1749AKGIRTLLFA LA MSLPALFNIC LLLFLVMFIF AIFGMSFF IS6 402PWHMLFFIVI IFLGSFYLVN LILAIVAMSY DE IIS6 1025WSCIPFFLAT VVIGNLVVLN LFLALLLSNF GS IIIS6 1552IYMYLYFVFF IIFGSFFTLN LFIGVIIDNF NE IIIS6 1852TIGITFLLSY LVISFLIVIN MYIAVILENY SQ Table 7.
Amino acid sequences of the Domain II-localized T929I mutation and the pyrethroid-binding site (IIL45-IIS5-IIIS6) in M. usitatus.
Figures
(3)
Tables
(7)