-
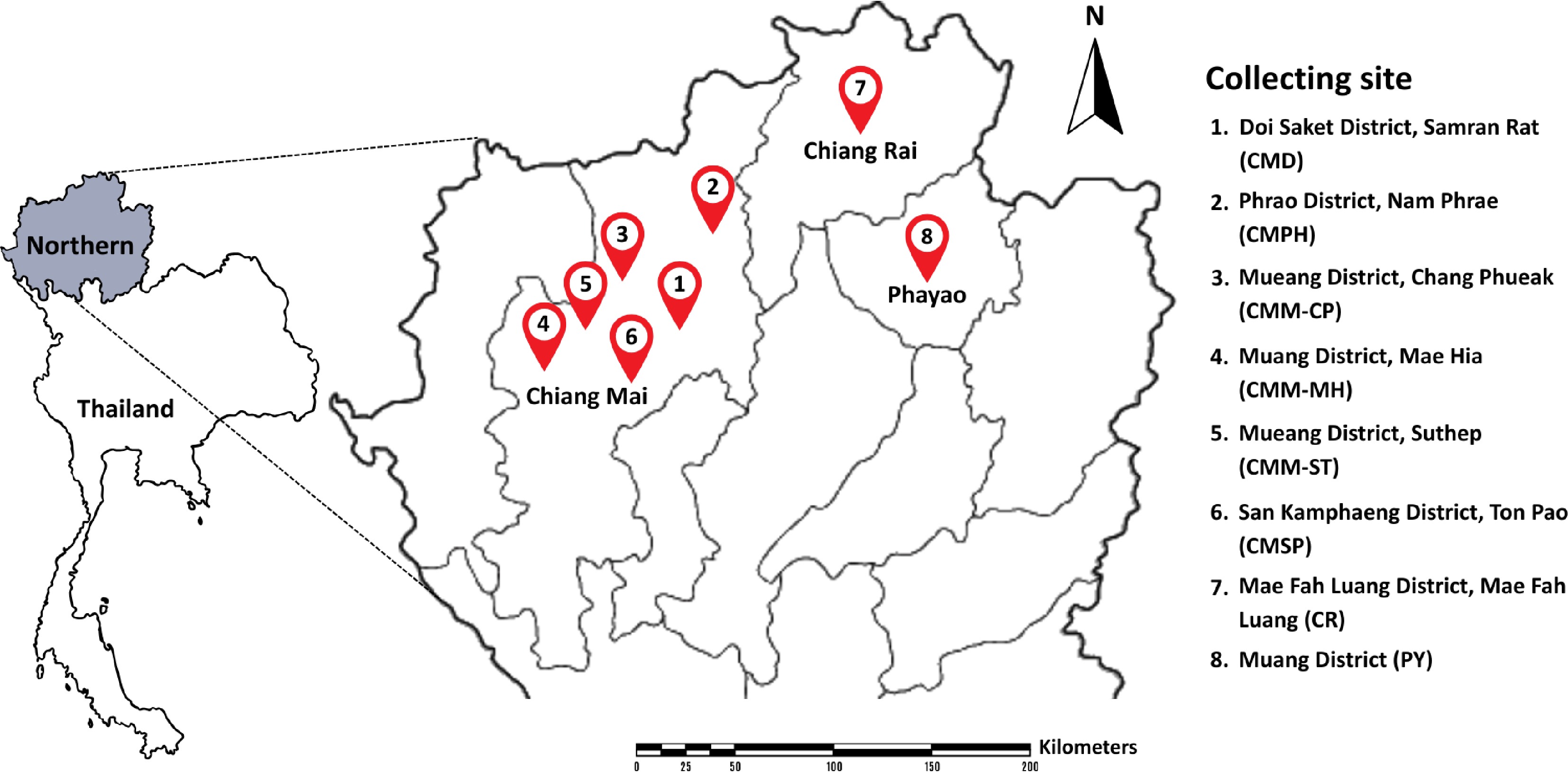
Figure 1.
Location of sampling sites in northern Thailand.
-
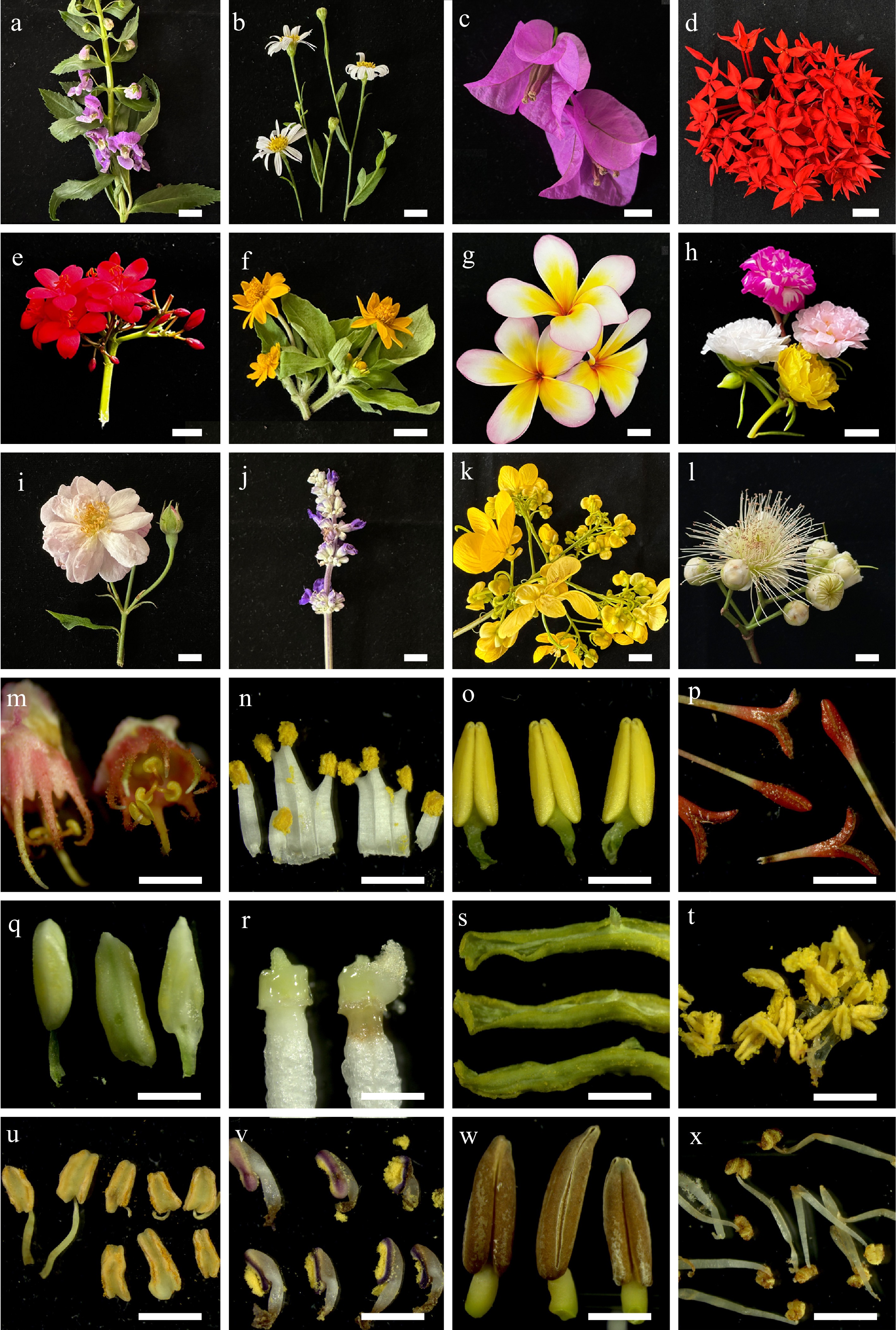

Figure 2.
(a)–(l) Some flower samples collected from northern Thailand, and their (m)–(x) internal parts. (a) Angelonia goyazensis, (b) Bellis perennis, (c) Bougainvillea hybrid, (d) Ixora chinensis, (e) Jatropha integerrima, (f) Melampodium divaricatum, (g) Plumeria rubra, (h) Portulaca grandiflora, (i) Rosa sp., (j) Salvia farinacea, (k) Senna spectabilis, (l) Syzygium jambos. The internal parts of (m) Antigonon leptopus, (n) Citrus japonica, (o) Exacum affine, (p) Ixora chinensis, (q) Jasminum sambac, (r) Nerium oleander, (s) Passiflora caerulea, (t) Portulaca grandiflora, (u) Rosa sp., (v) Salvia farinacea, (w) Senna spectabilis, and (x) Syzygium jambos. Scale bars (a)–(l) = 1 cm, and (m)–(x) = 1 mm.
-
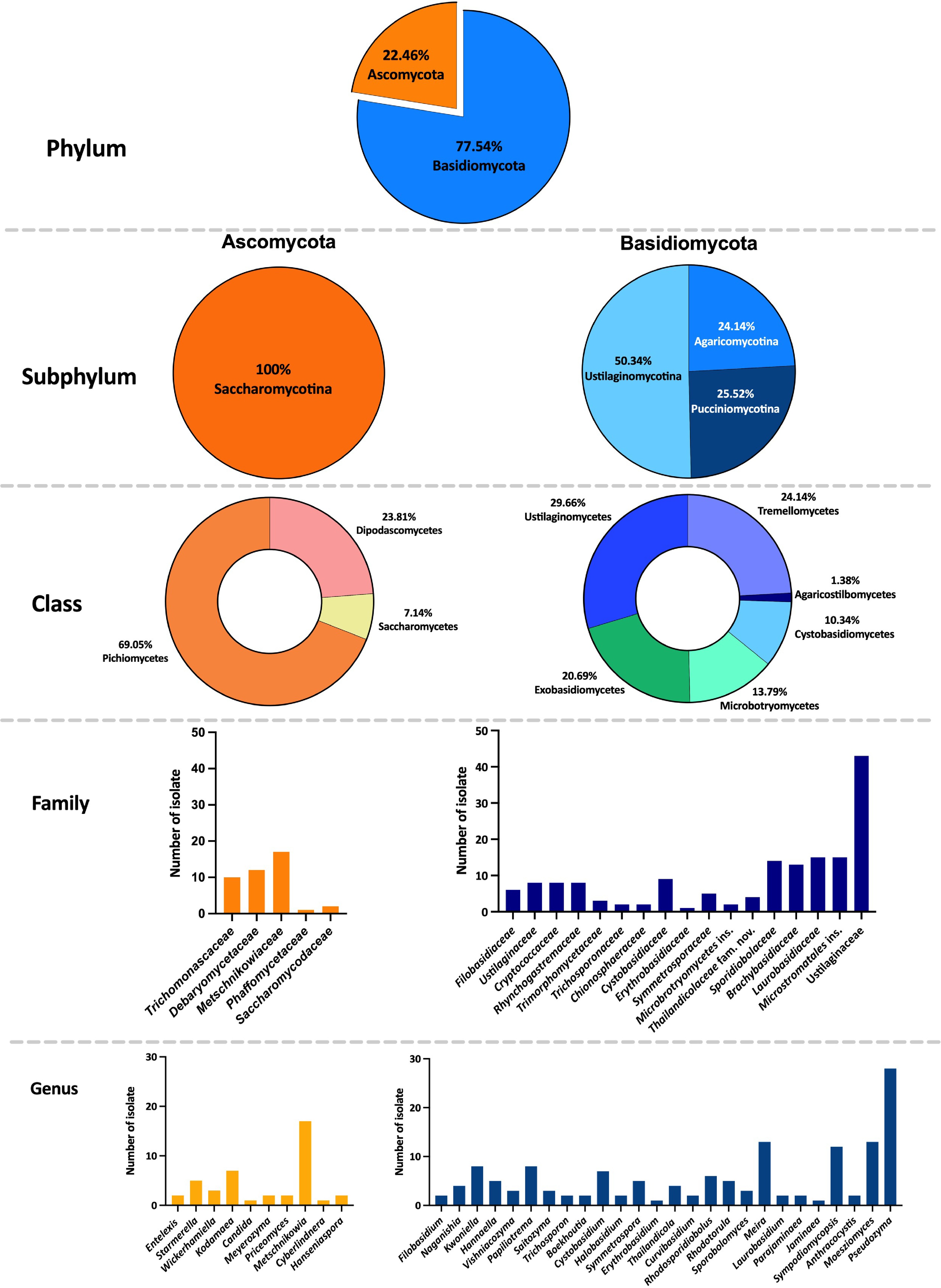
Figure 3.
Classification of anthophilous yeasts isolated from flowers collected in northern Thailand in this study.
-
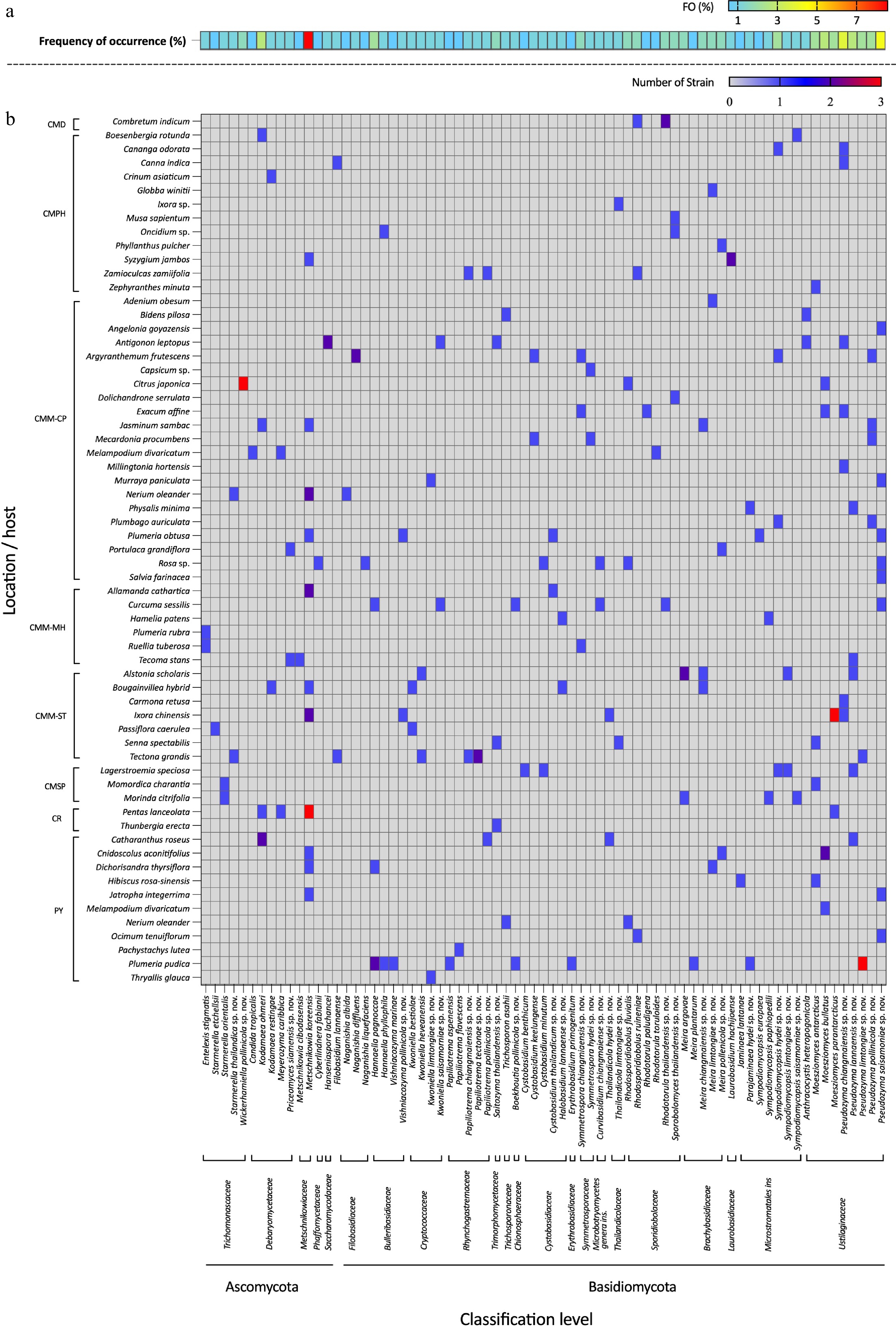
Figure 4.
Heatmap showing the consensus species diversity of anthophilous yeasts isolated from flowers collected in northern Thailand. (a) Frequency of occurrence (FO %) = number of samples, (b) where a particular species was observed, as a proportion of the total number of samples.
-
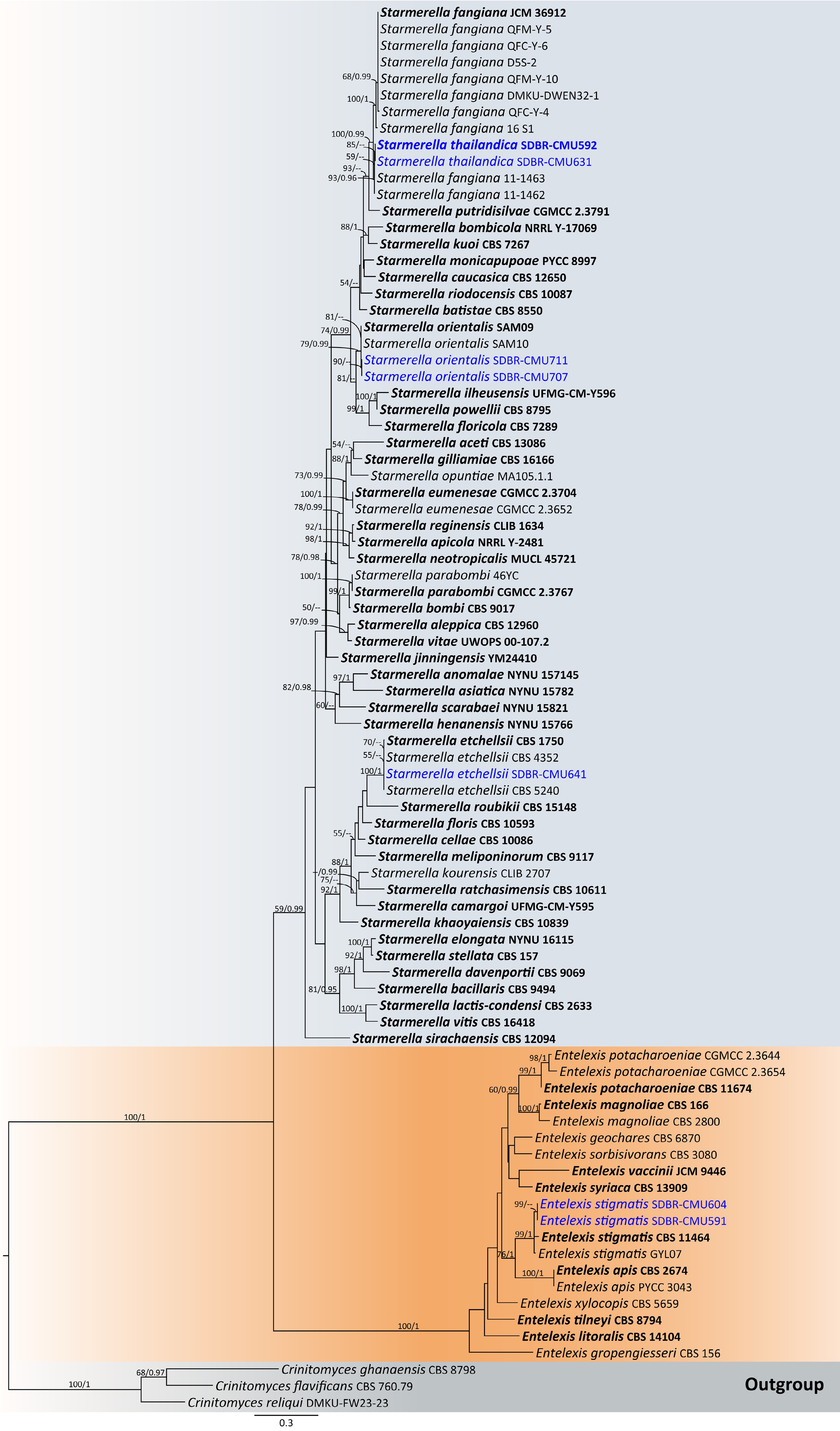
Figure 5.
Phylogram generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing genera Entelexis and Starmerella in Trichomonascaceae. The tree is rooted to Crinitomyces flavificans (CBS 760.79), C. ghanaensis (CBS 8798), and C. reliqui (DMKU-FW23-23). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Eighty-seven strains are included in the combined sequence analysis, which comprise 1,111 characters with gaps. Bootstrap support values for maximum likelihood (≥ 50%, ML, left) and Bayesian posterior probabilities (≥ 0.95, PP, right) are indicated above the nodes. Double dashes (--) denote support values below 50% ML and 0.95 PP. The scale bar represents 0.3 nucleotide substitutions per site. Ex-type strains are shown in bold, and sequences generated in this study are highlighted in blue.
-
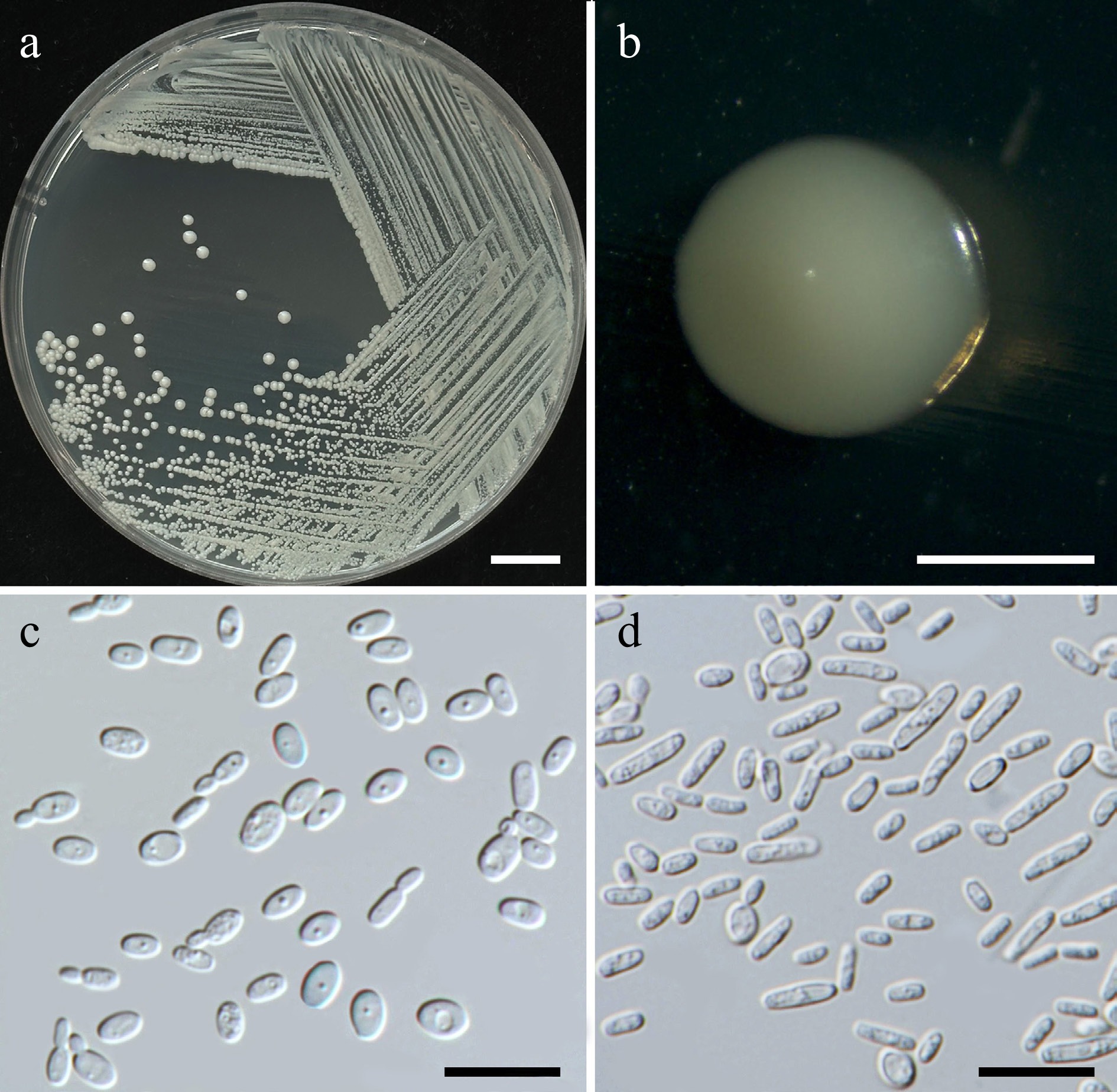
Figure 6.
Morphological characteristics of Starmerella thailandica (SDBR-CMU592, ex-type). (a) Culture, (b) single colony, (c) budding cells, on YMA at 25 °C for 5 d, and (d) elongated cells on PDA at 25 °C after 2 weeks. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
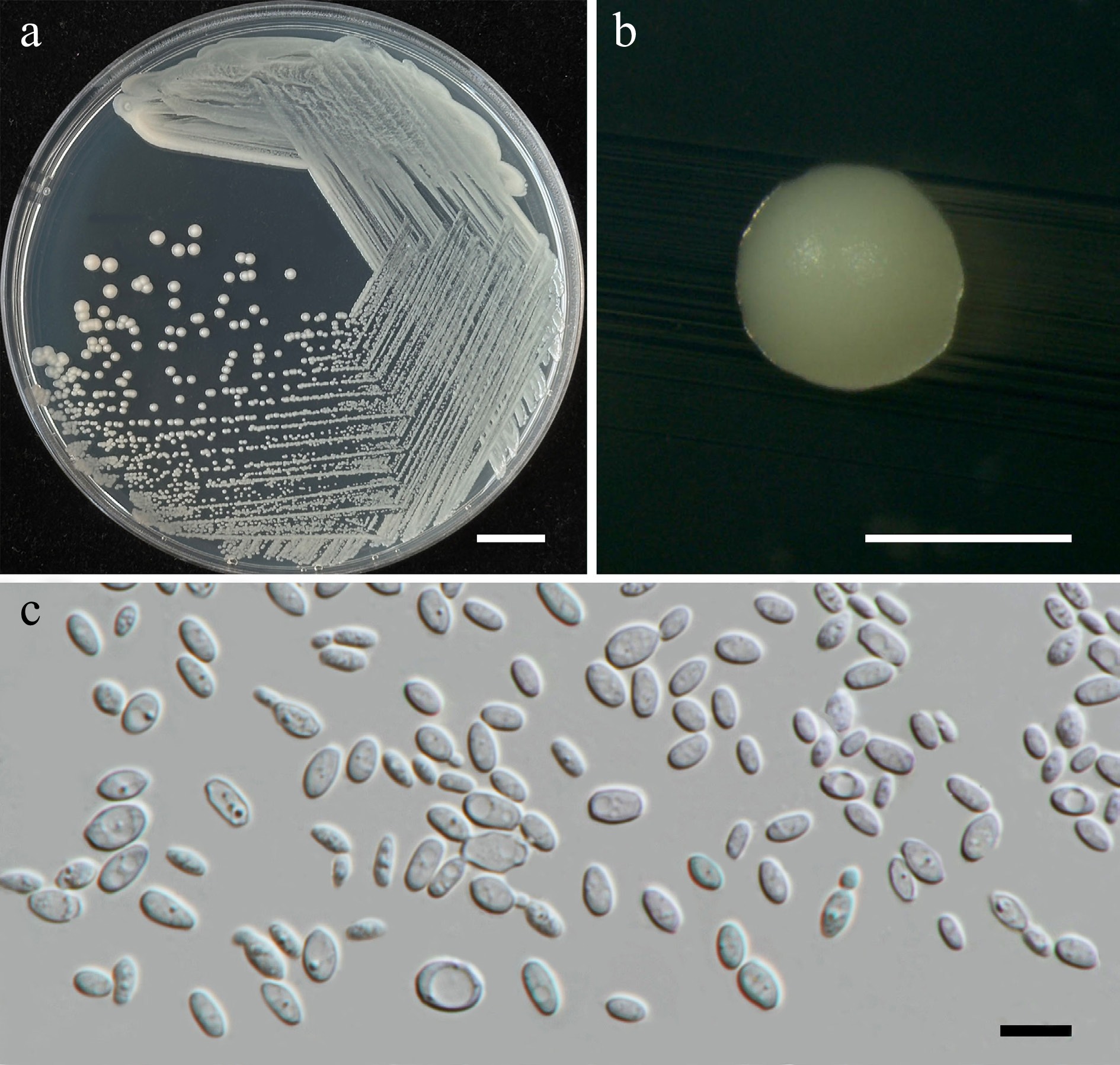
Figure 7.
Morphological characteristics of Wickerhamiella pollinicola (SDBR-CMU569, ex-type). (a) Culture, (b) single colony, (c) cells and budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, and (c) = 10 μm.
-

Figure 8.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing genus Wickerhamiella in Trichomonascaceae. The tree is rooted to Zygoascus hellenicus (CBS 5839) and Z. meyerae (CBS 4099). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Forty-nine strains are included in the combined sequence analysis, which comprise 1,190 characters with gaps. Bootstrap support values for maximum likelihood (≥ 50%, ML, left) and Bayesian posterior probabilities (≥ 0.95, PP, right) are indicated above the nodes. Double dashes (--) denote support values below 50% ML and 0.95 PP. The scale bar represents 0.1 nucleotide substitutions per site. Ex-type strains are shown in bold, and sequences generated in this study are highlighted in blue.
-
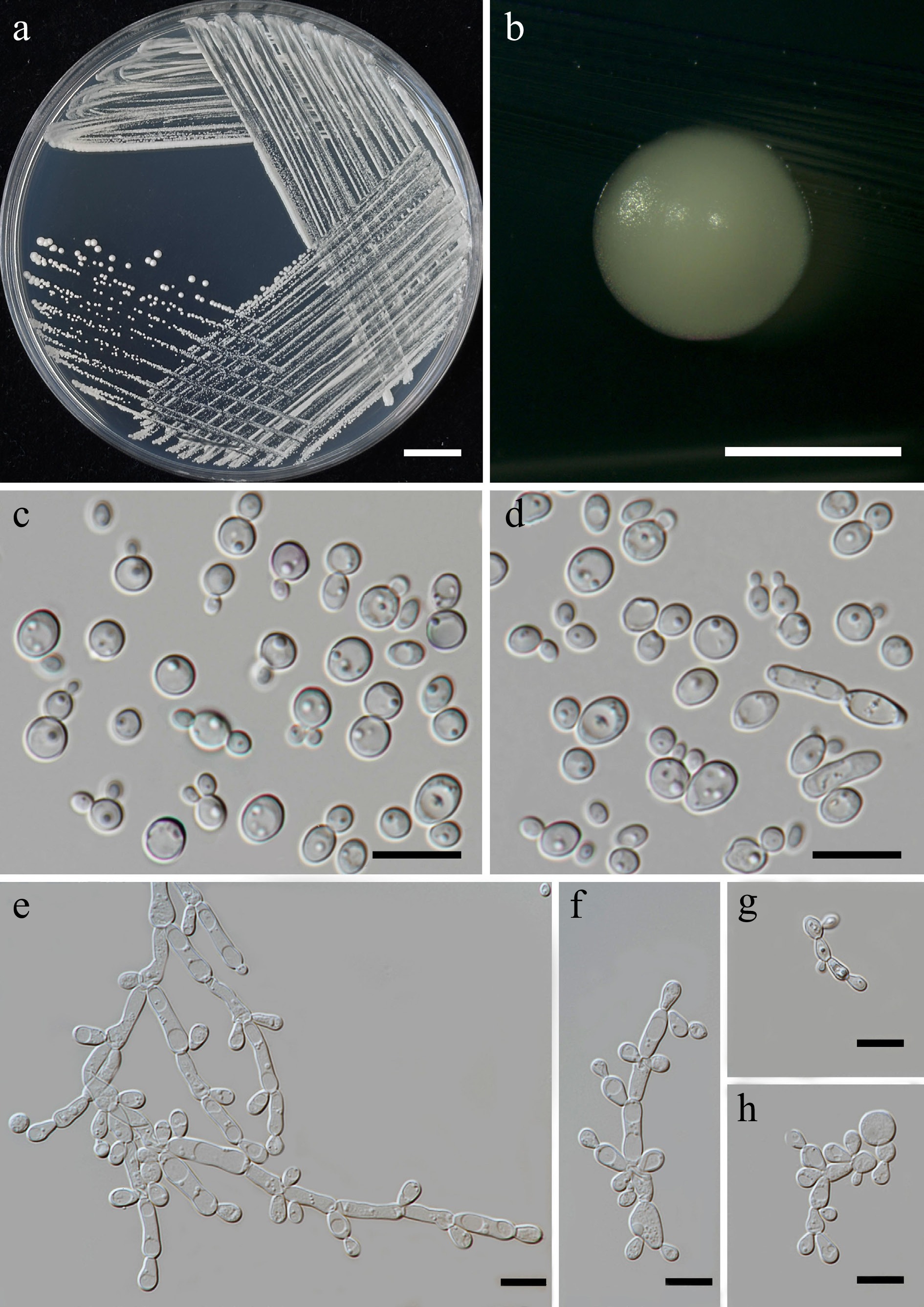
Figure 9.
Morphological characteristics of Priceomyces siamensis (SDBR-CMU614, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on YMA after 5 days at 25 °C. (e)–(h) Pseudohyphae and blastoconidia on PDA after 2 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, and (c)–(h) = 10 μm.
-
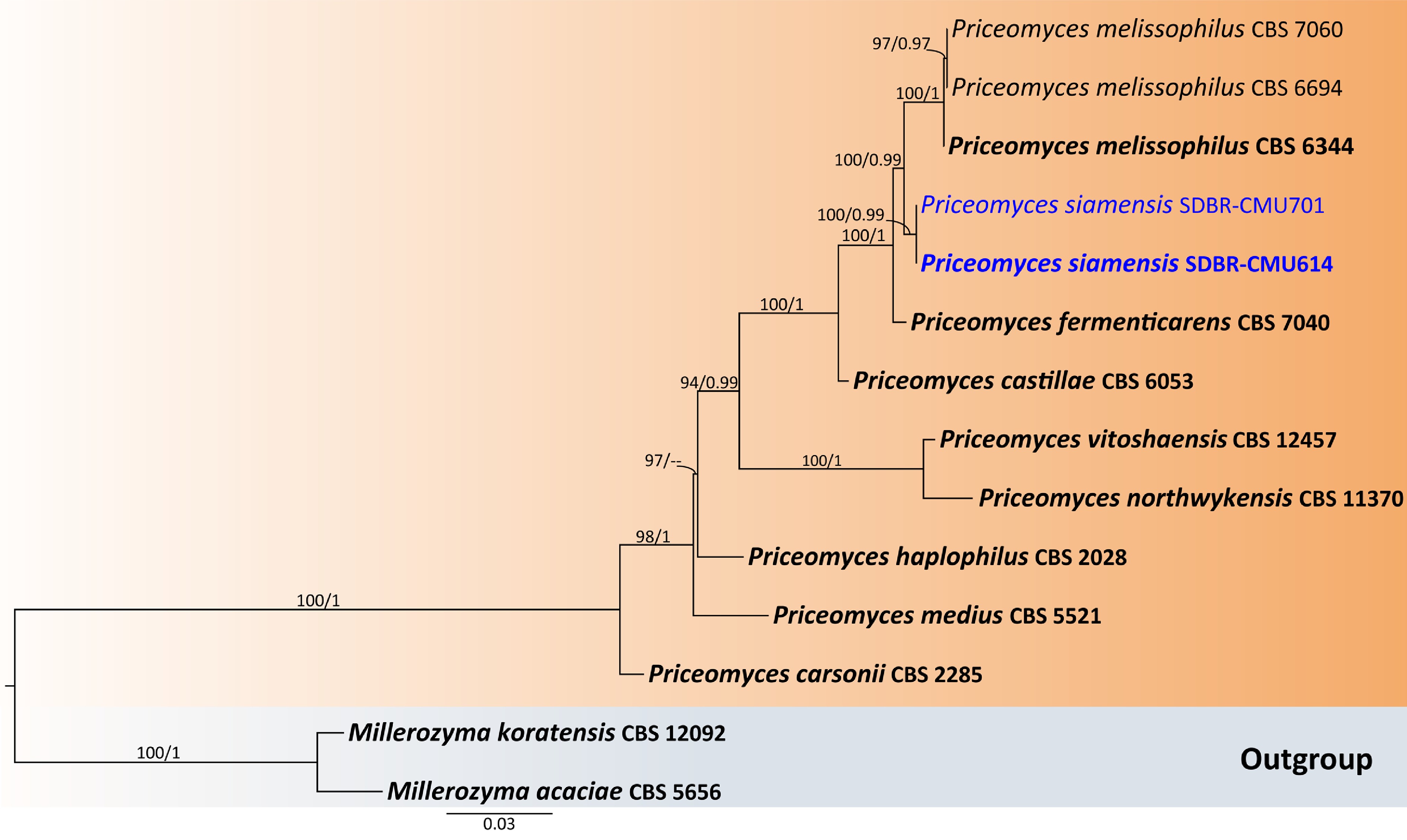
Figure 10.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Priceomyces. The tree is rooted to Millerozyma acaciae (CBS 5656) and M. koratensis (CBS 12092). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Fourteen strains are included in the combined sequence analysis, which comprise 1,121 characters with gaps. Bootstrap support values for maximum likelihood (≥ 50%, ML, left), and Bayesian posterior probabilities (≥ 0.95, PP, right) are indicated above the nodes. Double dashes (--) denote support values below 50% ML and 0.95 PP. The scale bar represents 0.03 nucleotide substitutions per site. Ex-type strains are shown in bold, and sequences generated in this study are highlighted in blue.
-
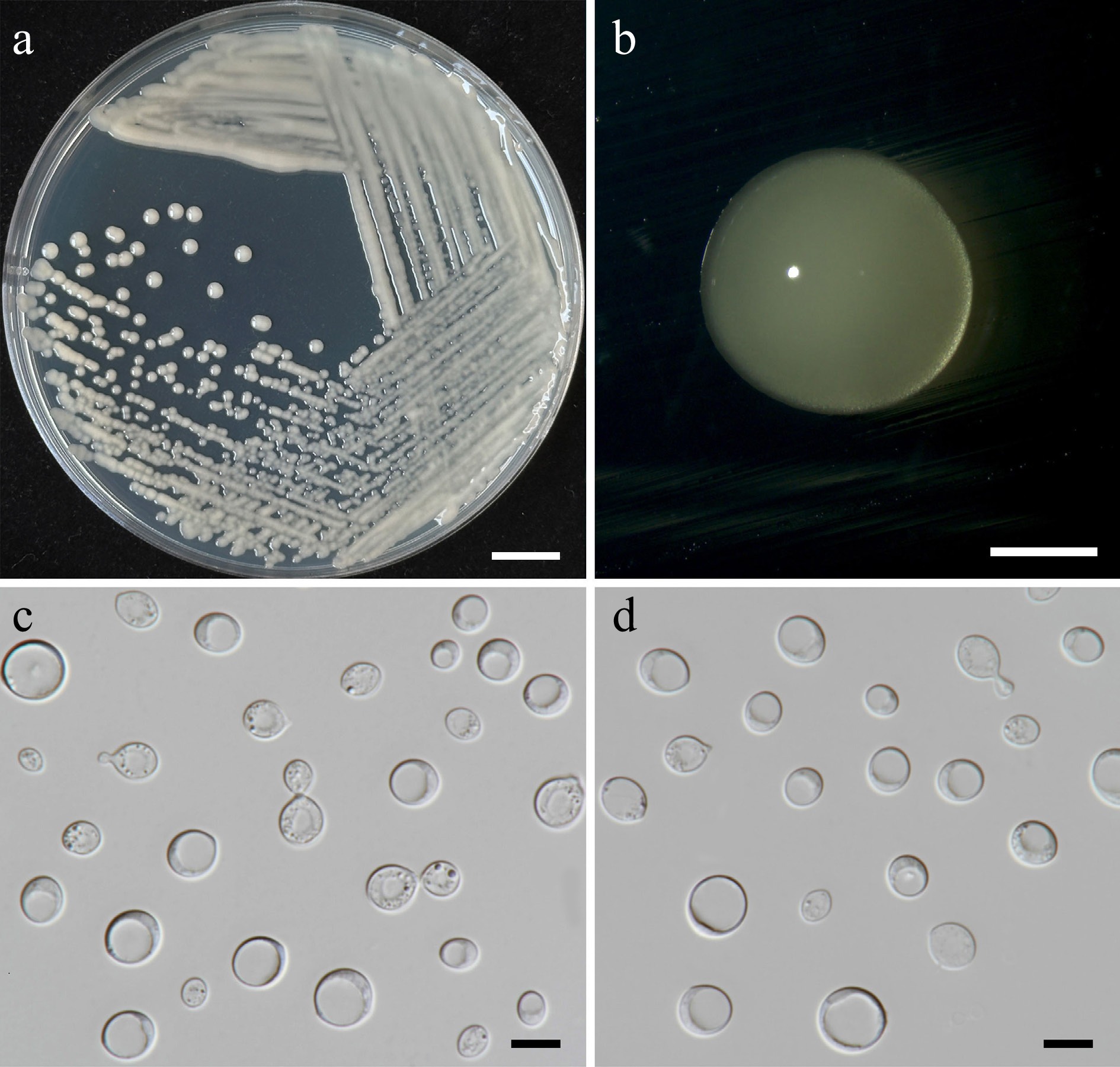
Figure 11.
Morphological characteristics of Filobasidium lannaense (SDBR-CMU628, ex-type). (a) Culture, (b) single colony, (c), (d) budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
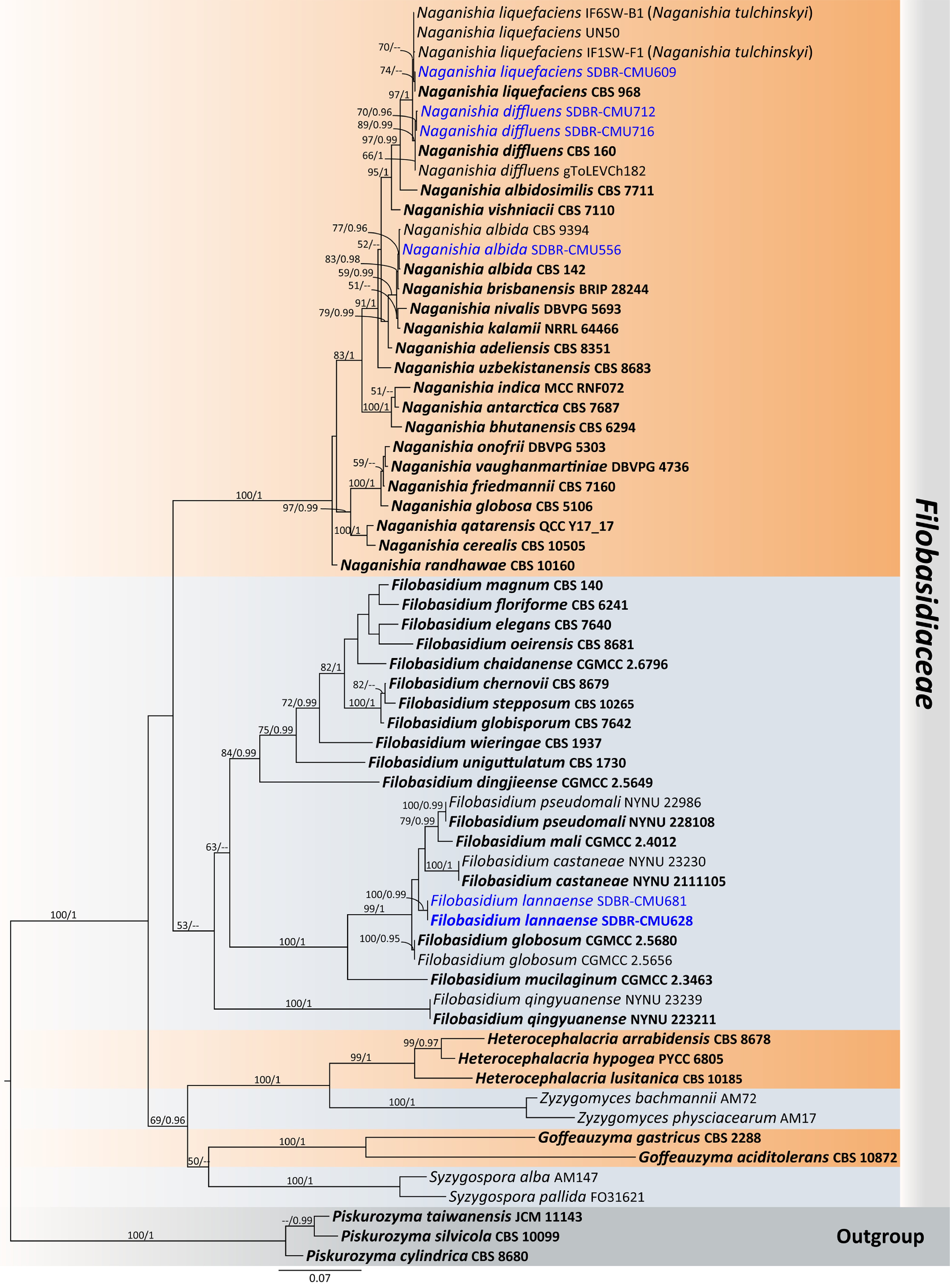
Figure 12.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Filobasidiaceae. The tree is rooted to Piskurozyma cylindrica (CBS 8680), Pi. silvicola (CBS 10099), and Pi. taiwanensis (JCM 11143). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Sixty-four strains are included in the combined sequence analysis, which comprise 1,398 characters with gaps. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
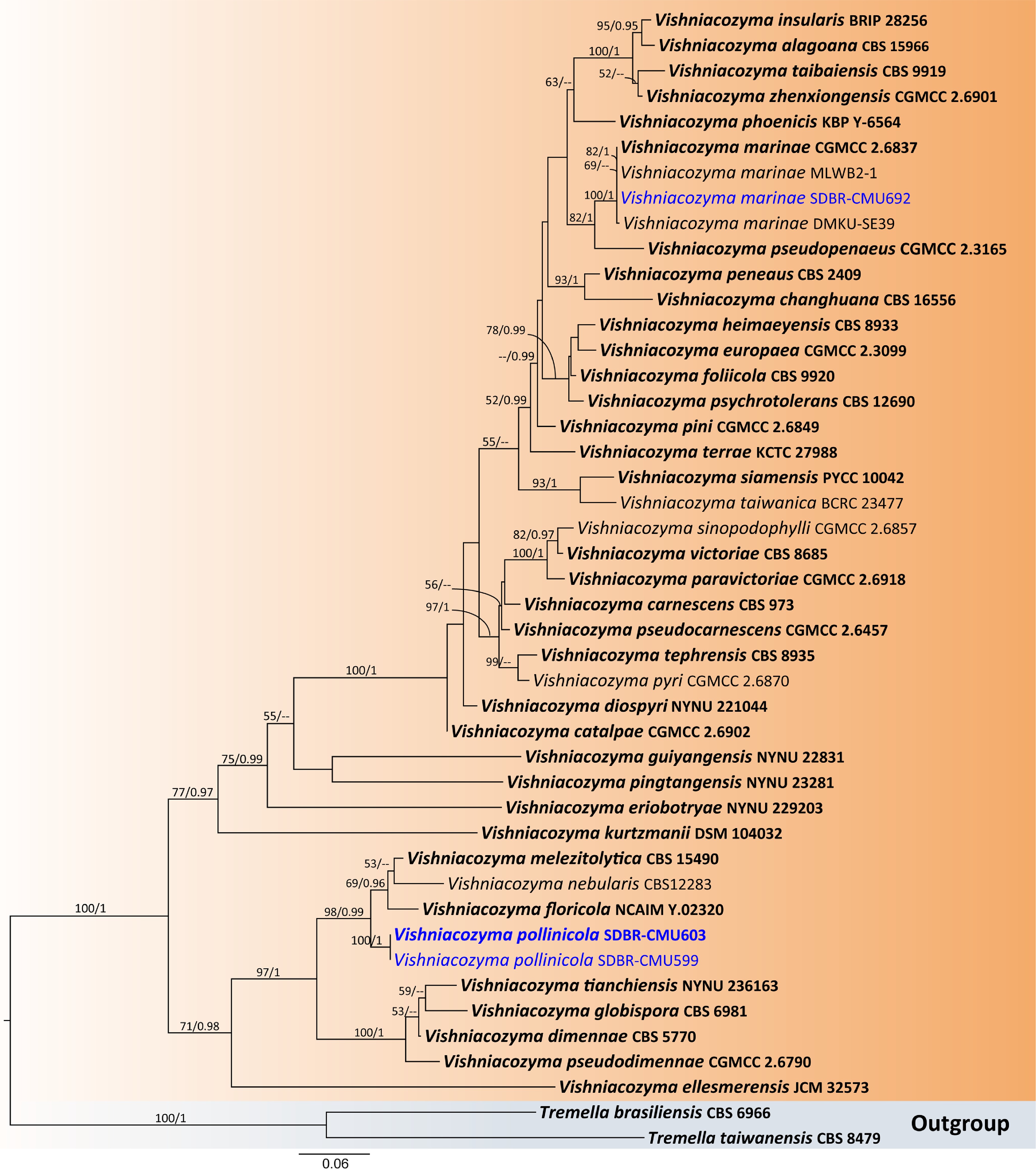
Figure 13.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Vishniacozyma. The tree is rooted to Tremella brasiliensis (CBS 6966) and T. taiwanensis (CBS 8479). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Fourty-five strains are included in the combined sequence analysis, which comprise 1,204 characters with gaps. The average standard deviation of the split frequencies of the BI analysis was 0.004360. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
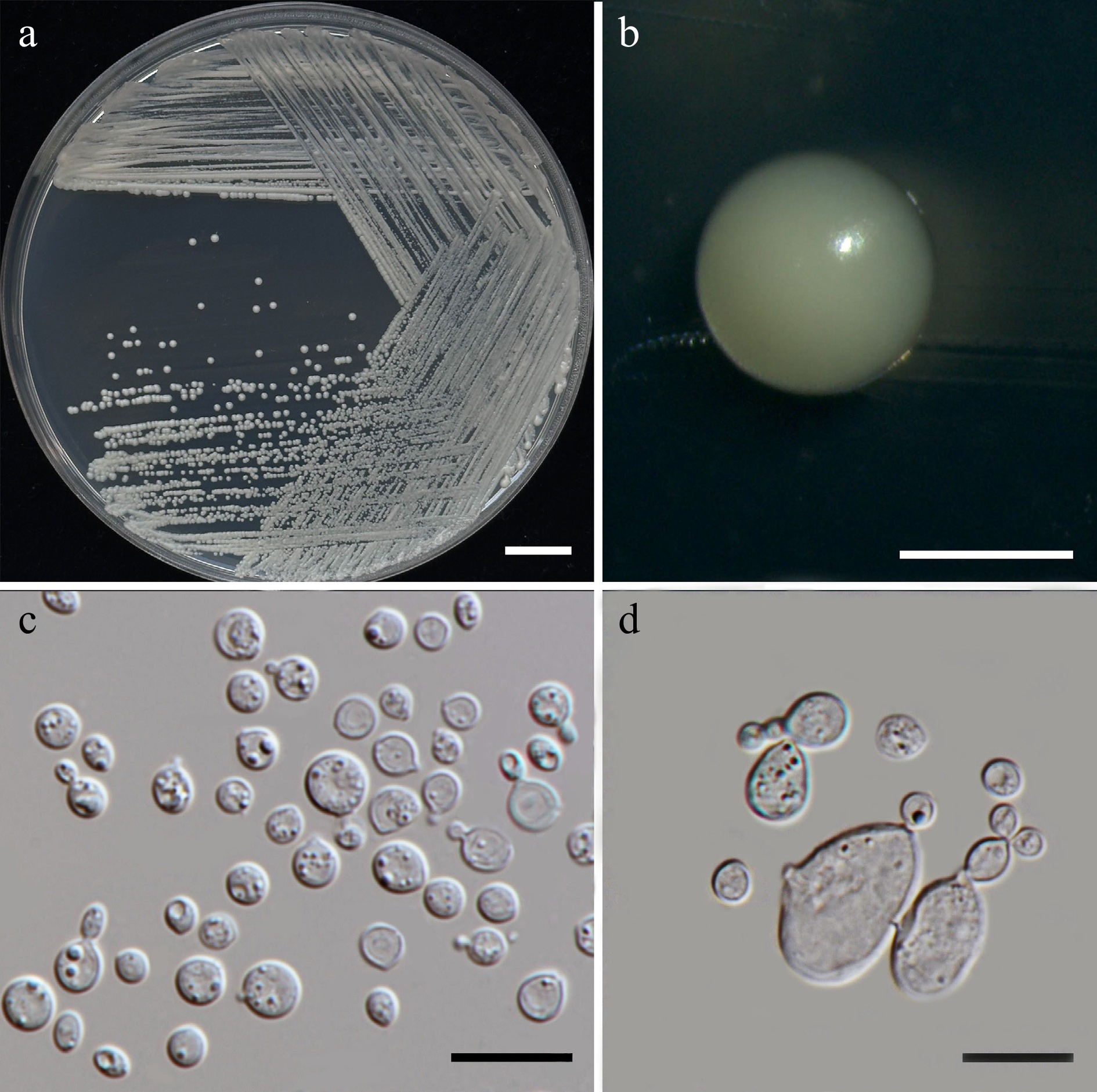
Figure 14.
Morphological characteristics of Vishniacozyma pollinicola (SDBR-CMU603, ex-type). (a) Culture, (b) single colony, (c) cells and budding cells on YMA after 5 d at 25 °C. (d) Rudimentary pseudohypha on YMA after 3 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
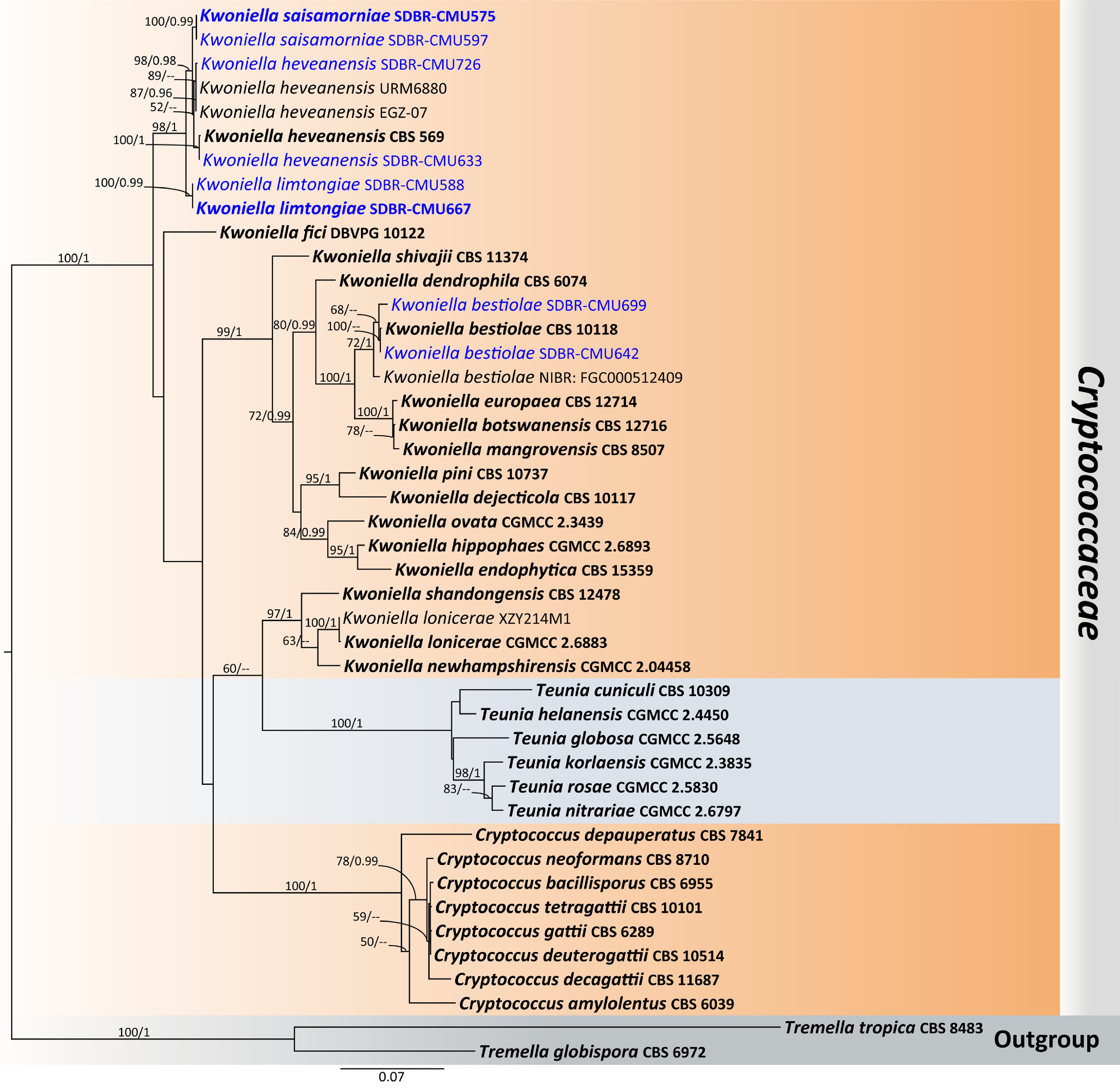
Figure 15.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Cryptococcaceae. The tree is rooted to Tremella globispora (CBS 6972) and T. tropica (CBS 8483). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Fourty-four strains are included in the combined sequence analysis, which comprise 1,274 characters with gaps. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
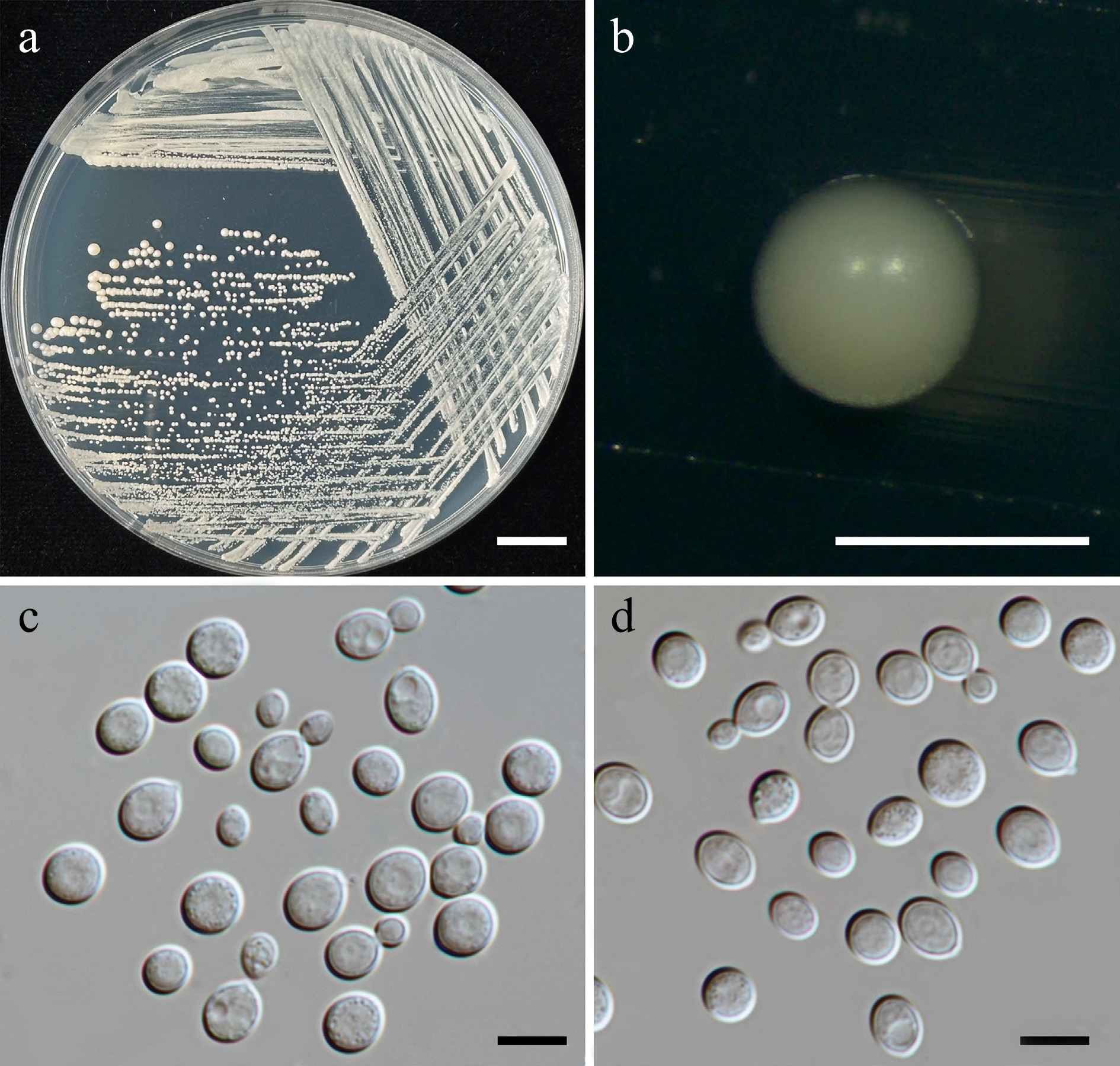
Figure 16.
Morphological characteristics of Kwoniella limtongiae (SDBR-CMU667, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
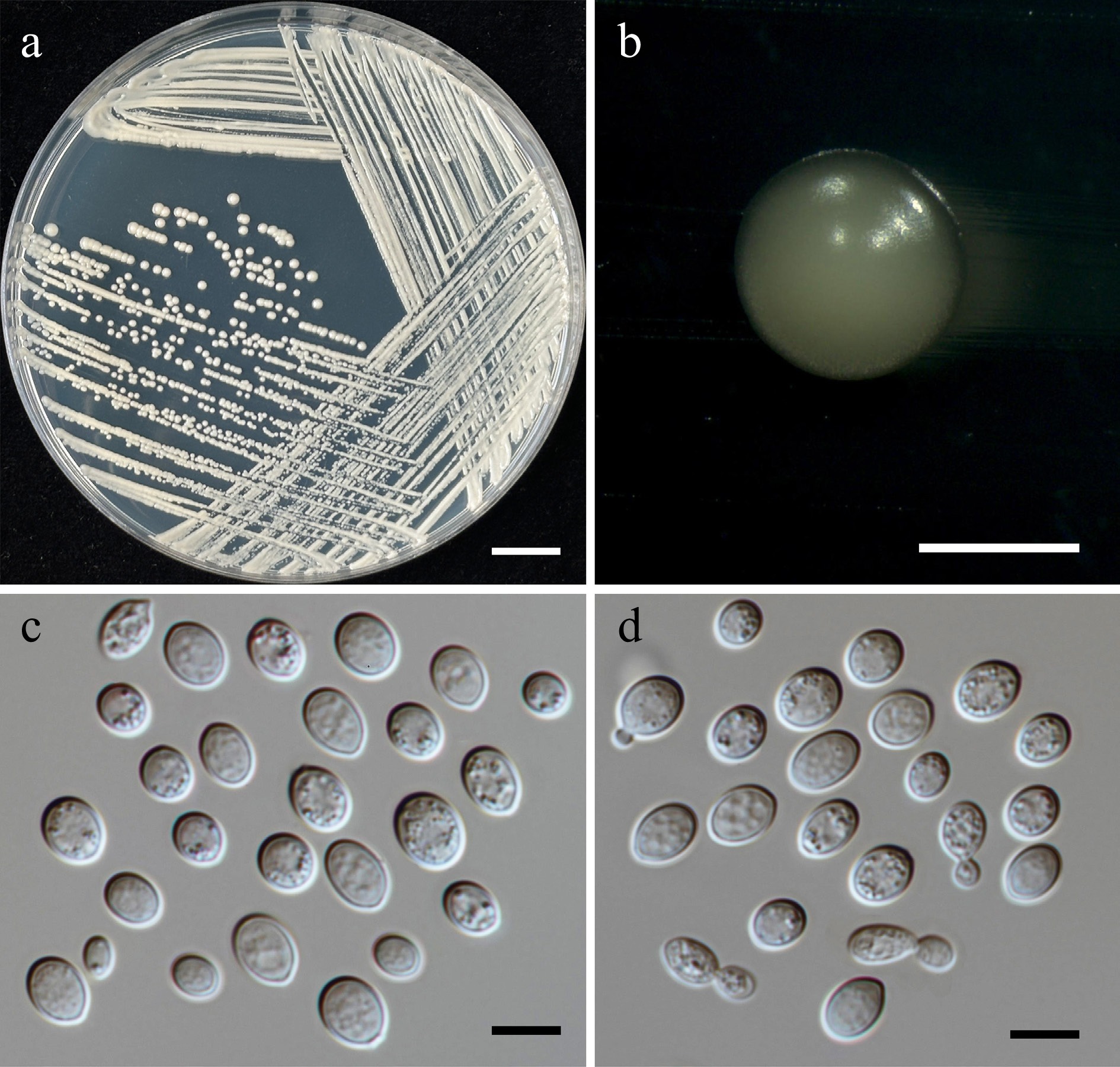
Figure 17.
Morphological characteristics of Kwoniella saisamorniae (SDBR-CMU575, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
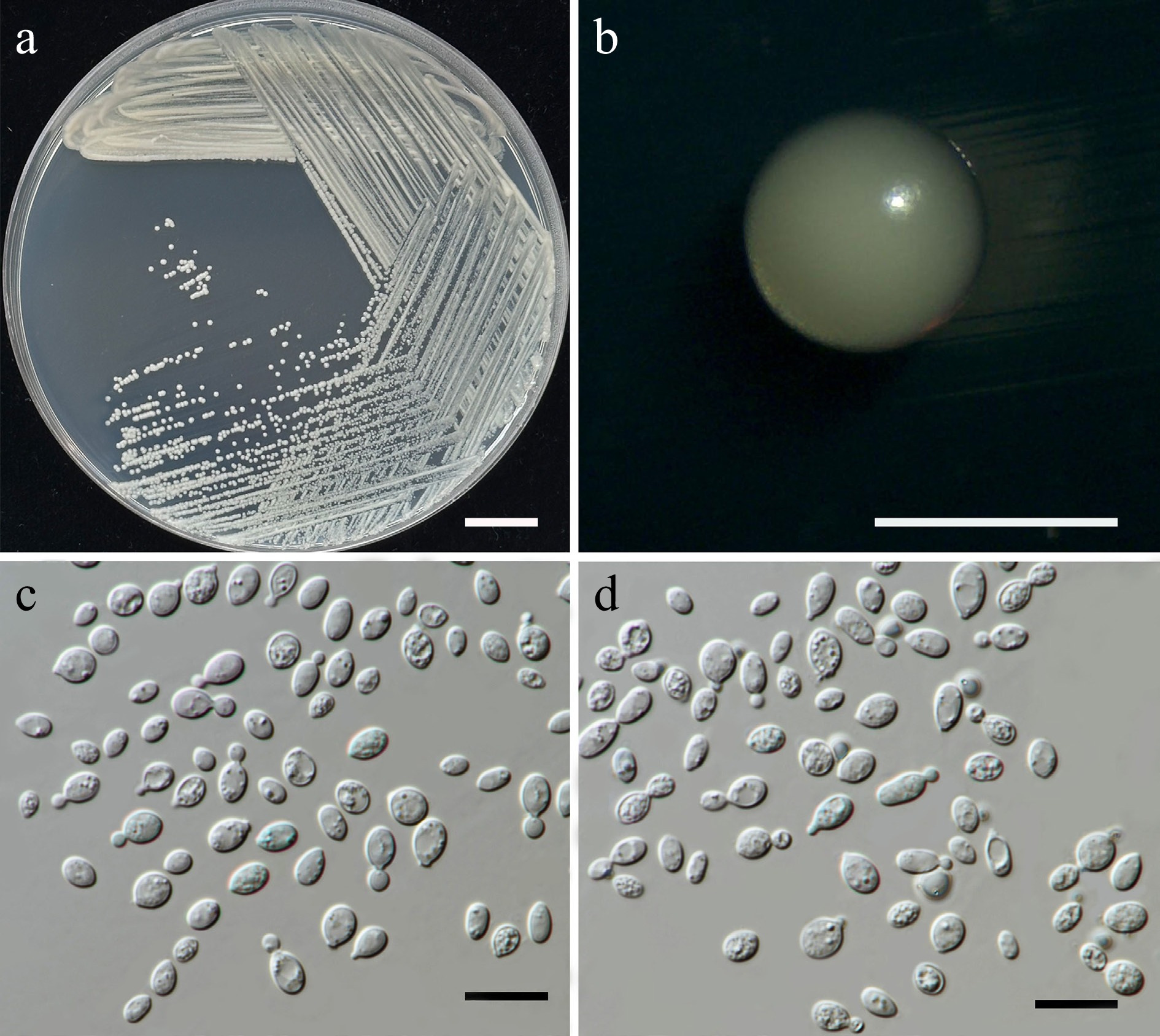
Figure 18.
Morphological characteristics of Papiliotrema chiangmaiensis (SDBR-CMU594, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
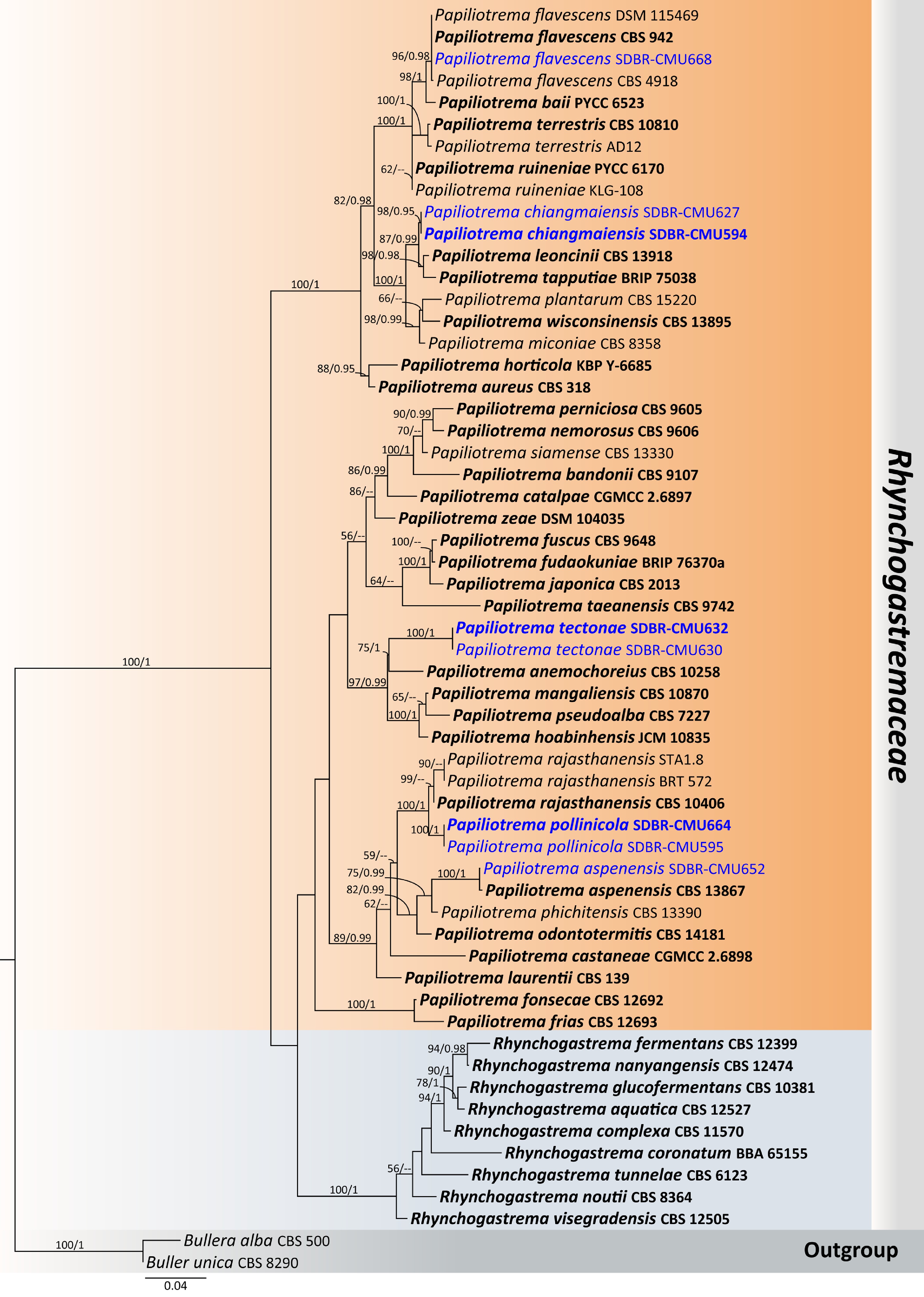
Figure 19.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Rhynchogastremaceae. The tree is rooted to Bullera alba (CBS 500) and B. unica (CBS 8290). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Fifty-eight strains are included in the combined sequence analysis, which comprise 1,218 characters with gaps. The average standard deviation of the split frequencies of the BI analysis was 0.009286. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
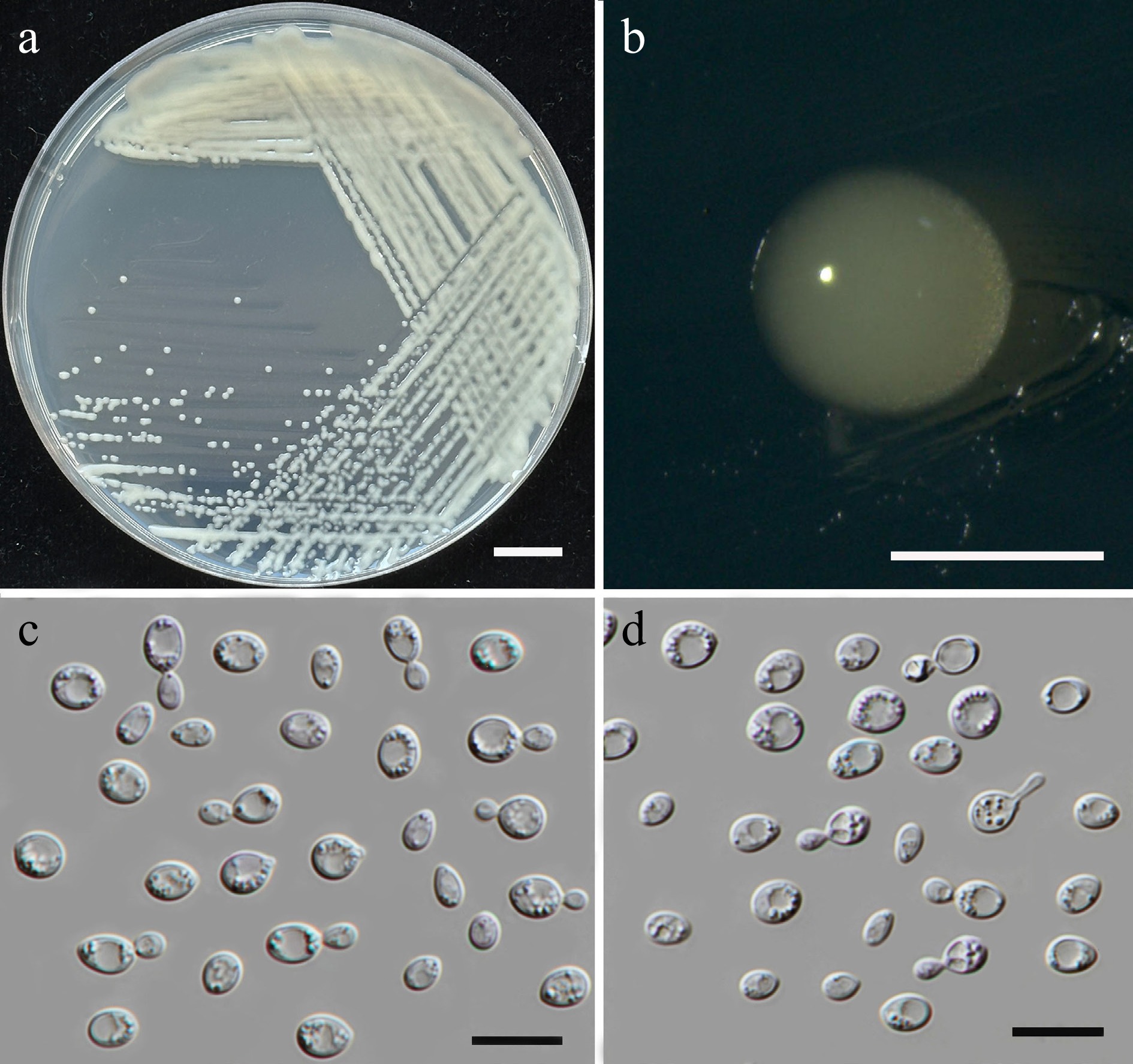
Figure 20.
Morphological characteristics of Papiliotrema pollinicola (SDBR-CMU664, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
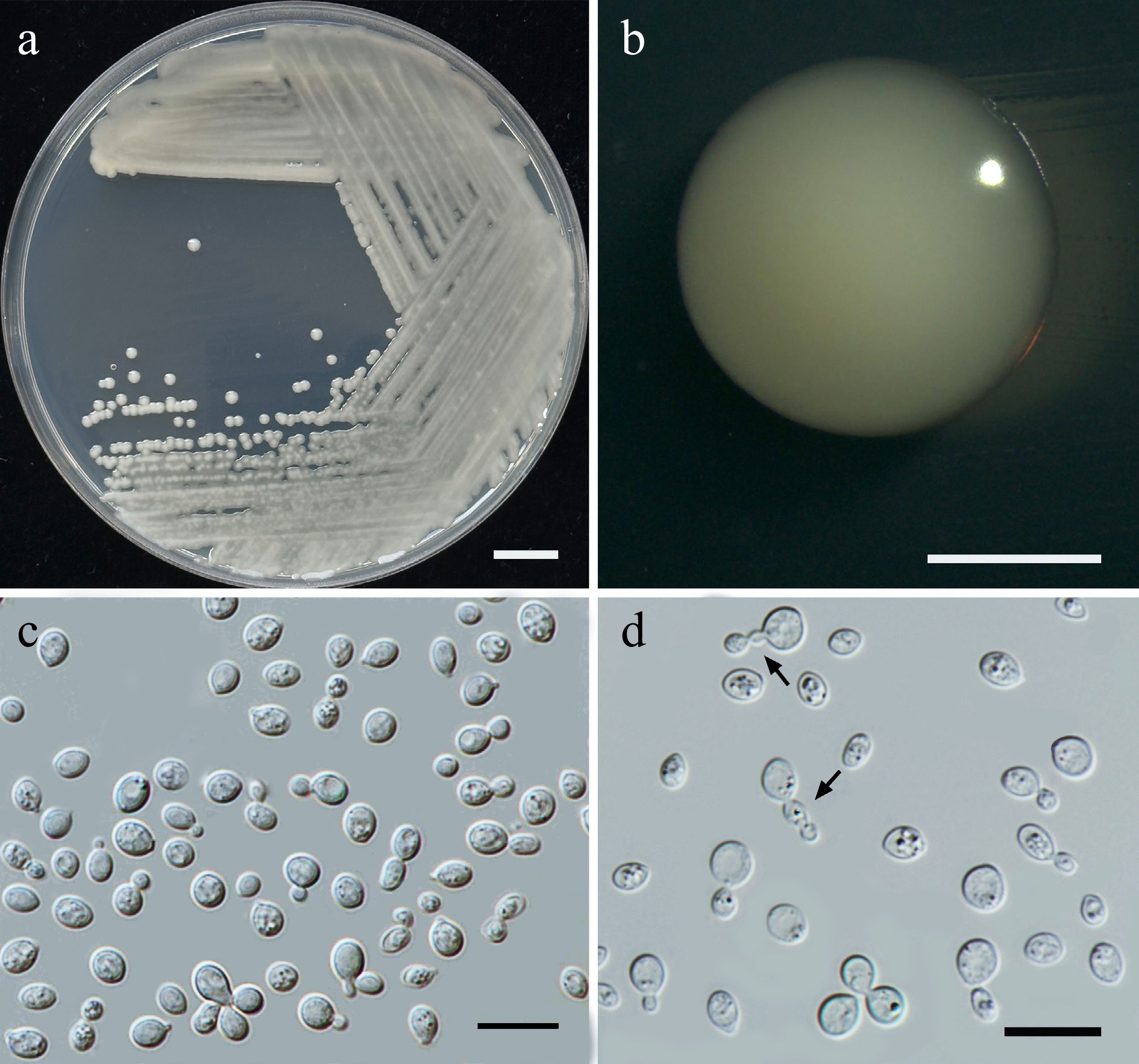
Figure 21.
Morphological characteristics of Papiliotrema tectonae (SDBR-CMU632, ex-type). (a) Culture, (b) single colony, (c), (d) budding cells, and chains of cells (indicated by arrows) on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
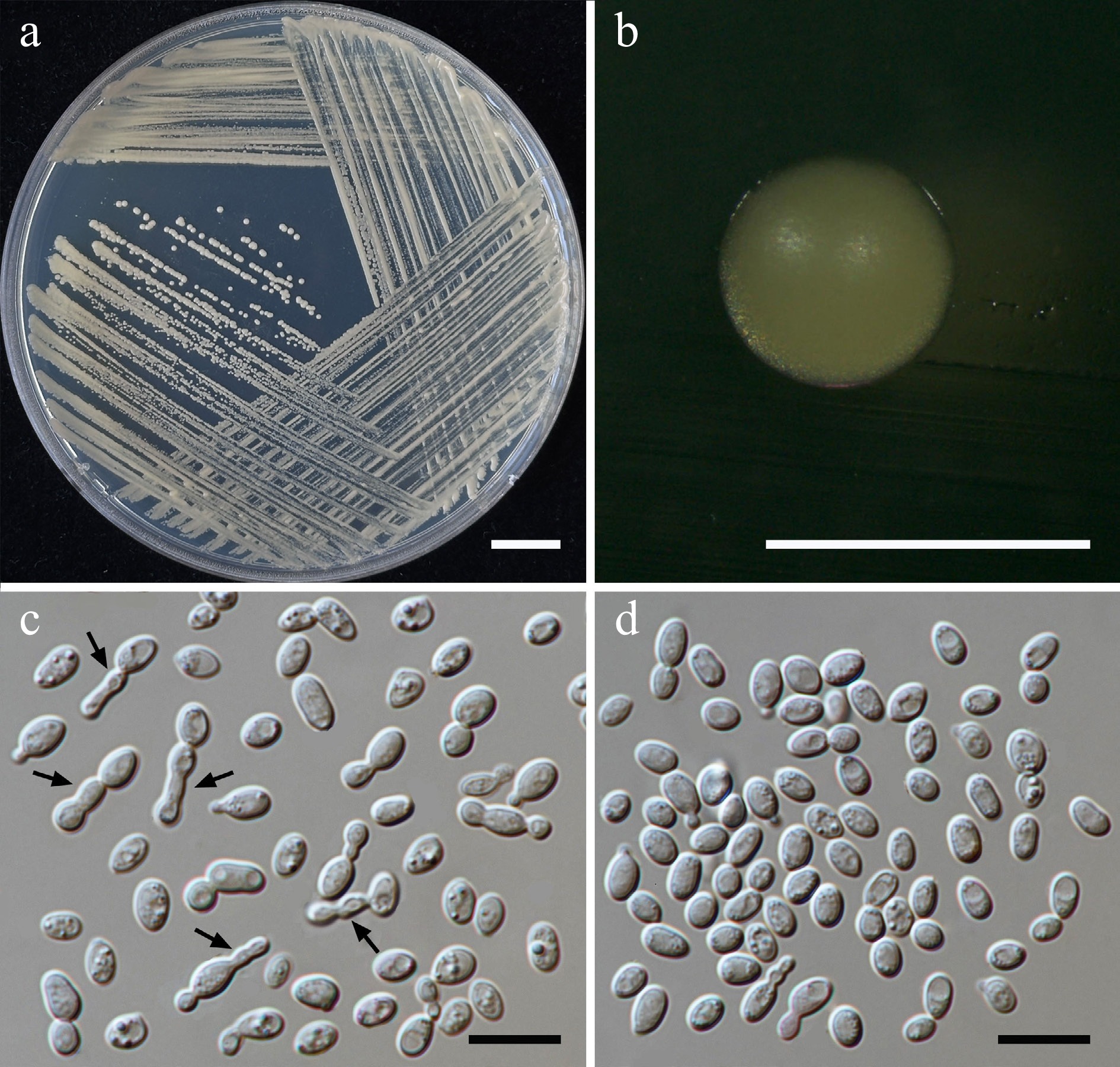
Figure 22.
Morphological characteristics of Saitozyma thailandensis (SDBR-CMU590, ex-type). (a) Culture, (b) single colony, (c),chains of cells (indicated by arrows), (d) budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
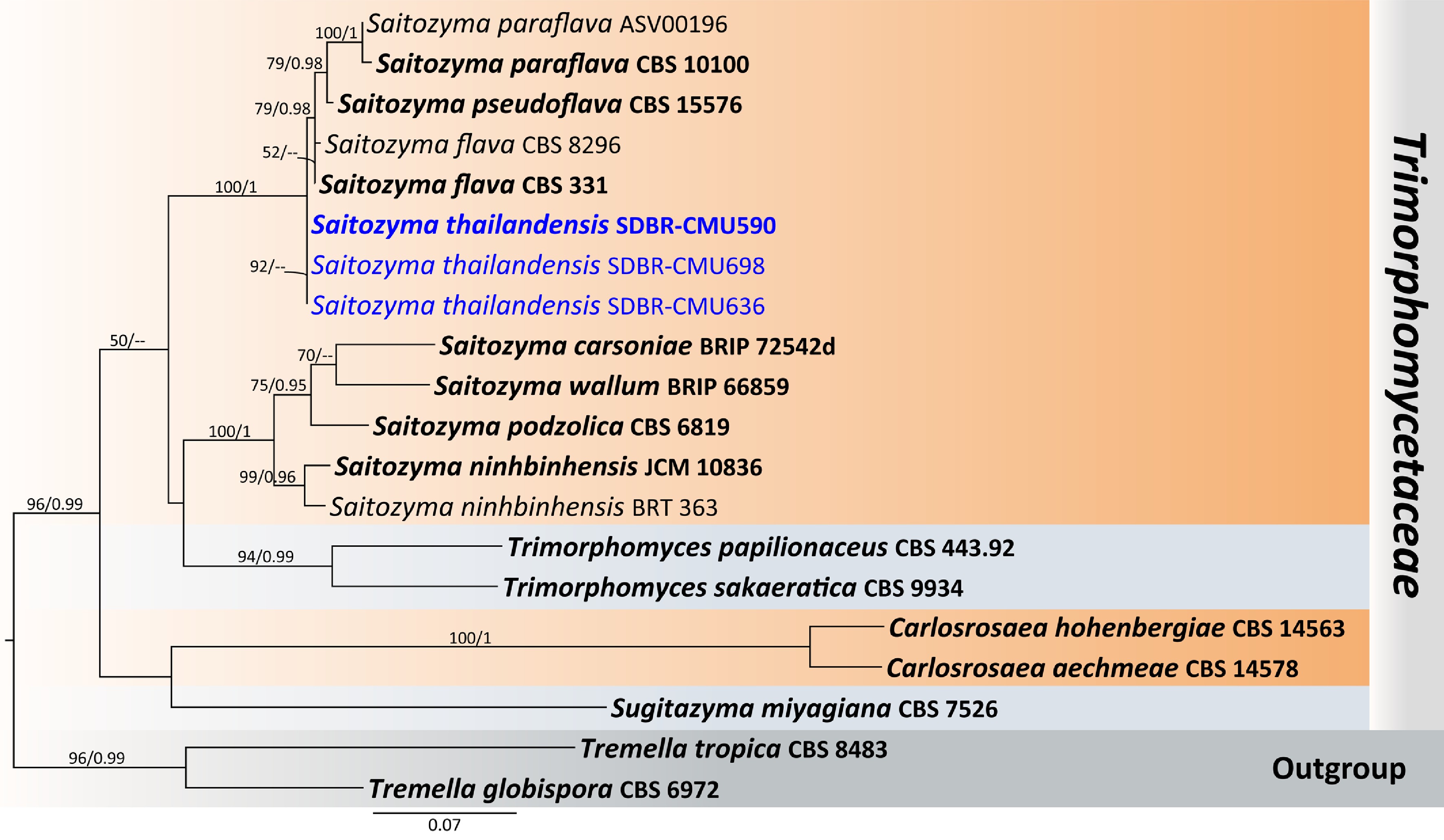
Figure 23.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Trimorphomycetaceae. The tree is rooted to Tremella globispora (CBS 6972) and T. tropica (CBS 8483). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Twenty strains are included in the combined sequence analysis, which comprise 1,149 characters with gaps. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
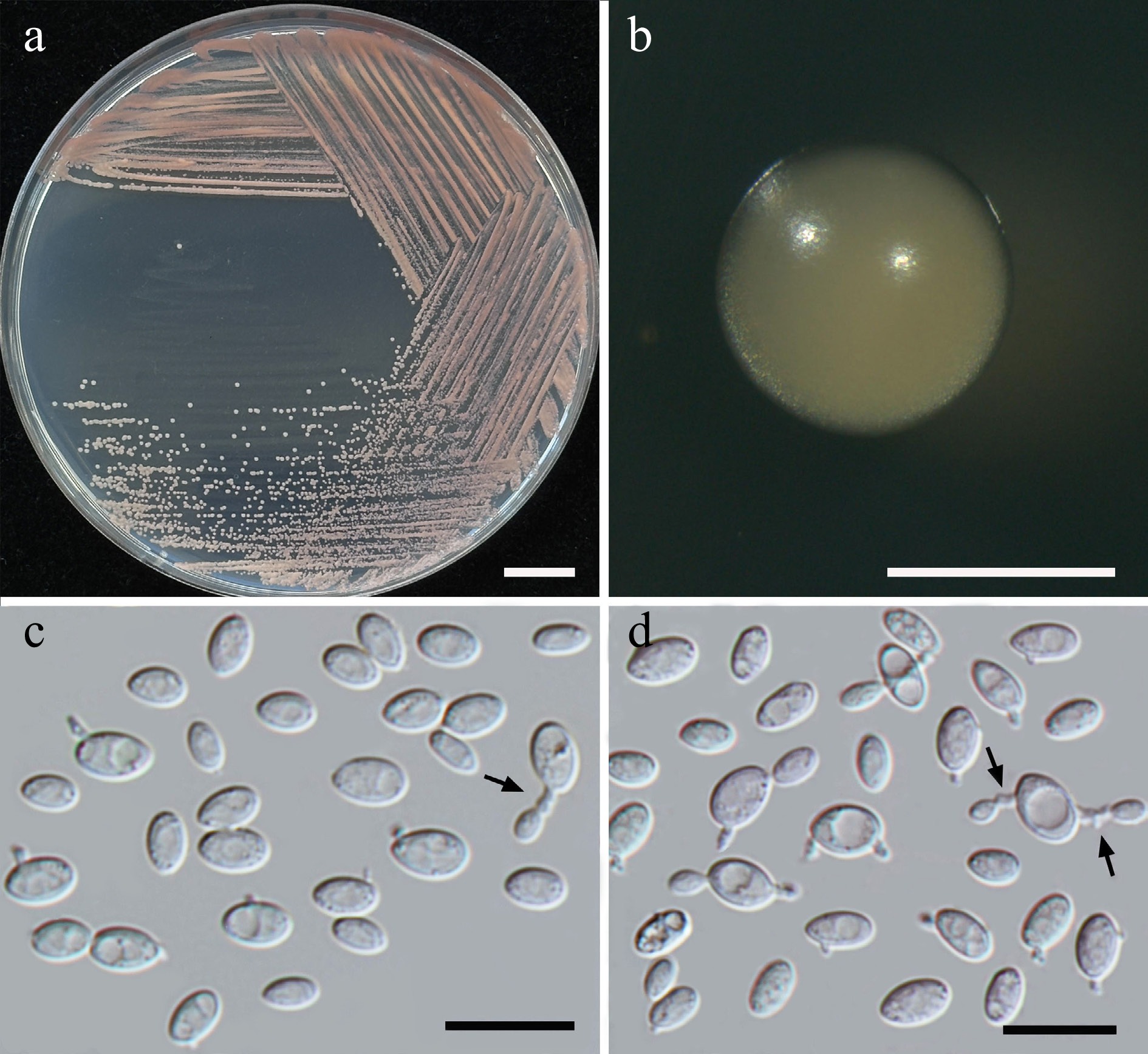
Figure 24.
Morphological characteristics of Boekhoutia pollinicola (SDBR-CMU684, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on sterigma (indicated by arrows) on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
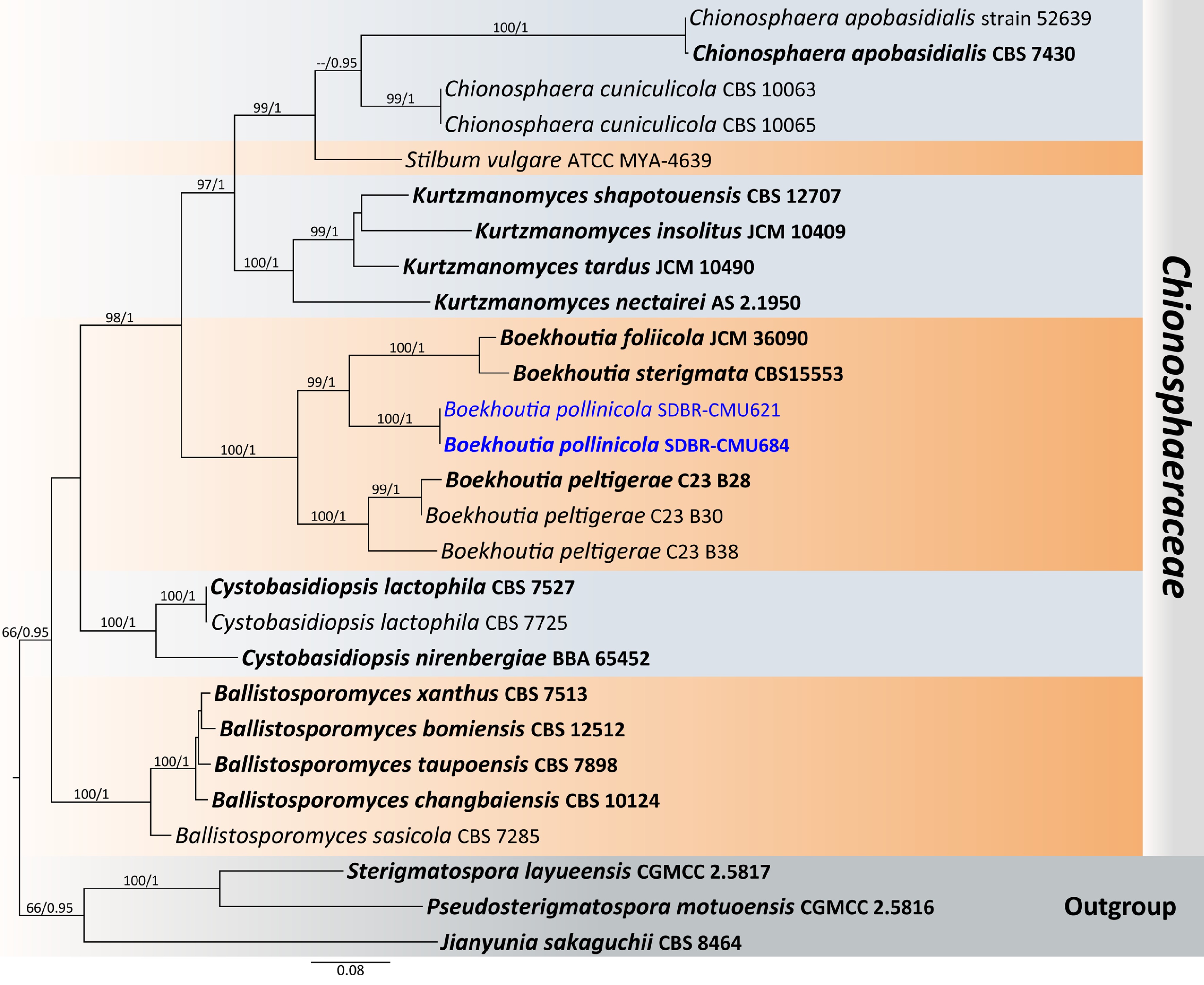
Figure 25.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Chionosphaeraceae. The tree is rooted to Jianyunia sakaguchii (CBS 8464), Pseudosterigmatospora motuoensis (CGMCC 2.5816), and Sterigmatospora layueensis (CGMCC 2.5817). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Twenty-seven strains are included in the combined sequence analysis, which comprise 1,594 characters with gaps. The average standard deviation of the split frequencies of the BI analysis was 0.001890. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
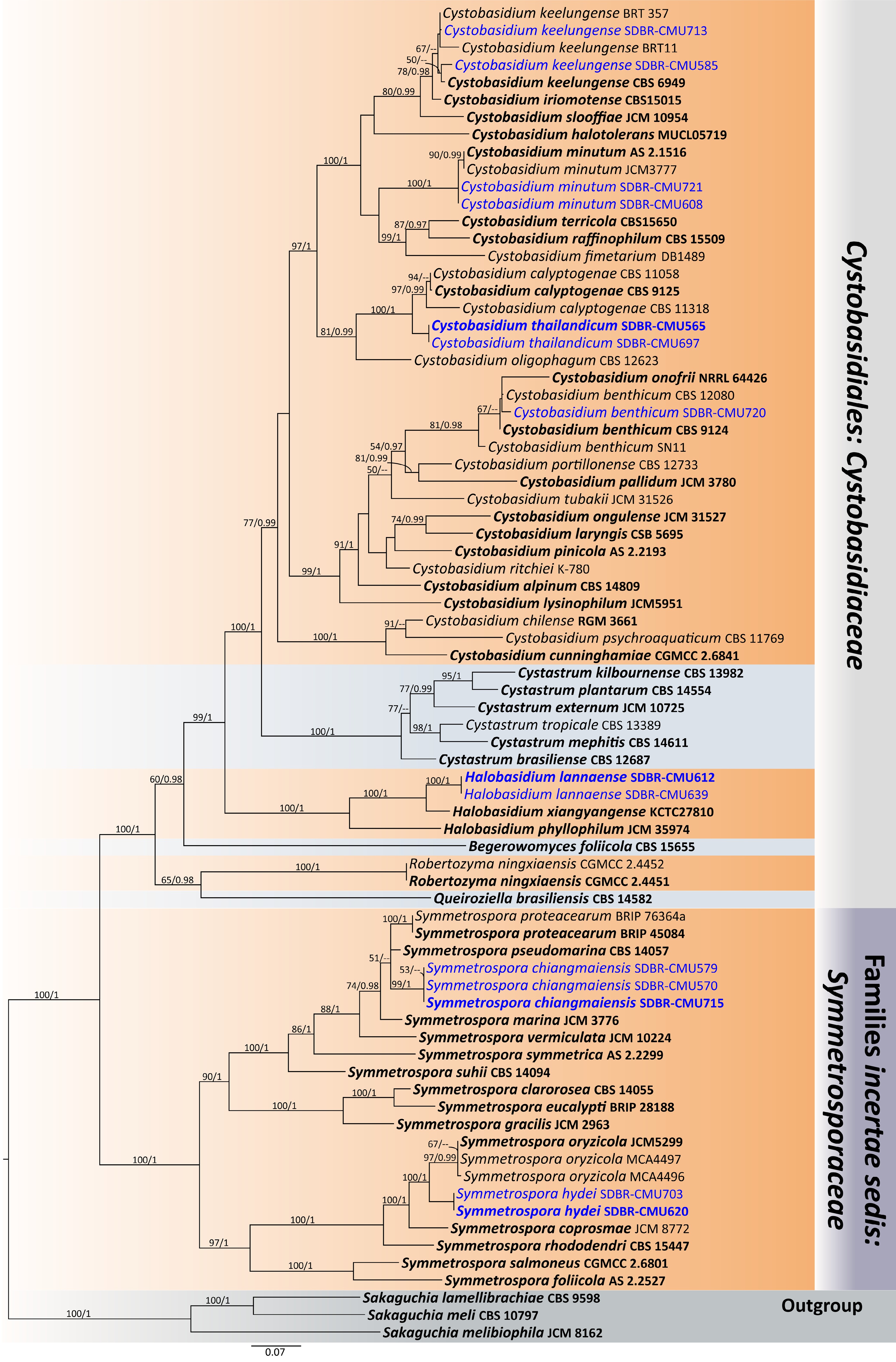
Figure 26.
Phylogram generated by maximum likelihood analysis of the combined D1/D2 domain of LSU, ITS, SSU, rpb1, rpb2, and tef1-α sequence data representing Cystobasidiaceae (Cystobasidiales) and Symmetrosporaceae (Incertae sedis) in Cystobasidiomycetes. The tree is rooted to Sakaguchia lamellibrachiae (CBS 9598), S. meli (CBS 10797), and S. melibiophila (JCM 8162). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Seventy-seven strains are included in the combined sequence analysis, which comprise 7,211 characters with gaps. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
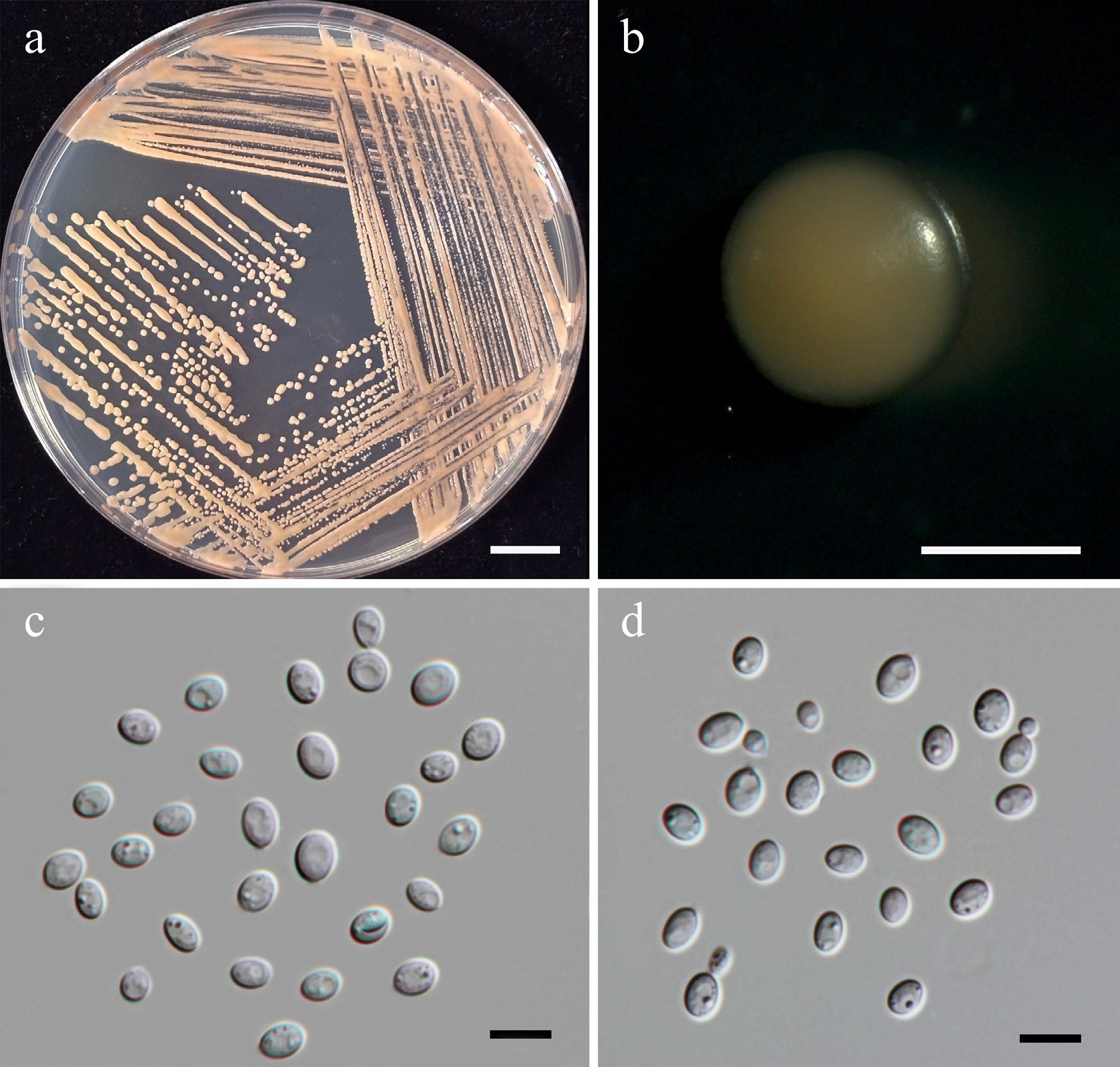
Figure 27.
Morphological characteristics of Cystobasidium thailandicum (SDBR-CMU697, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
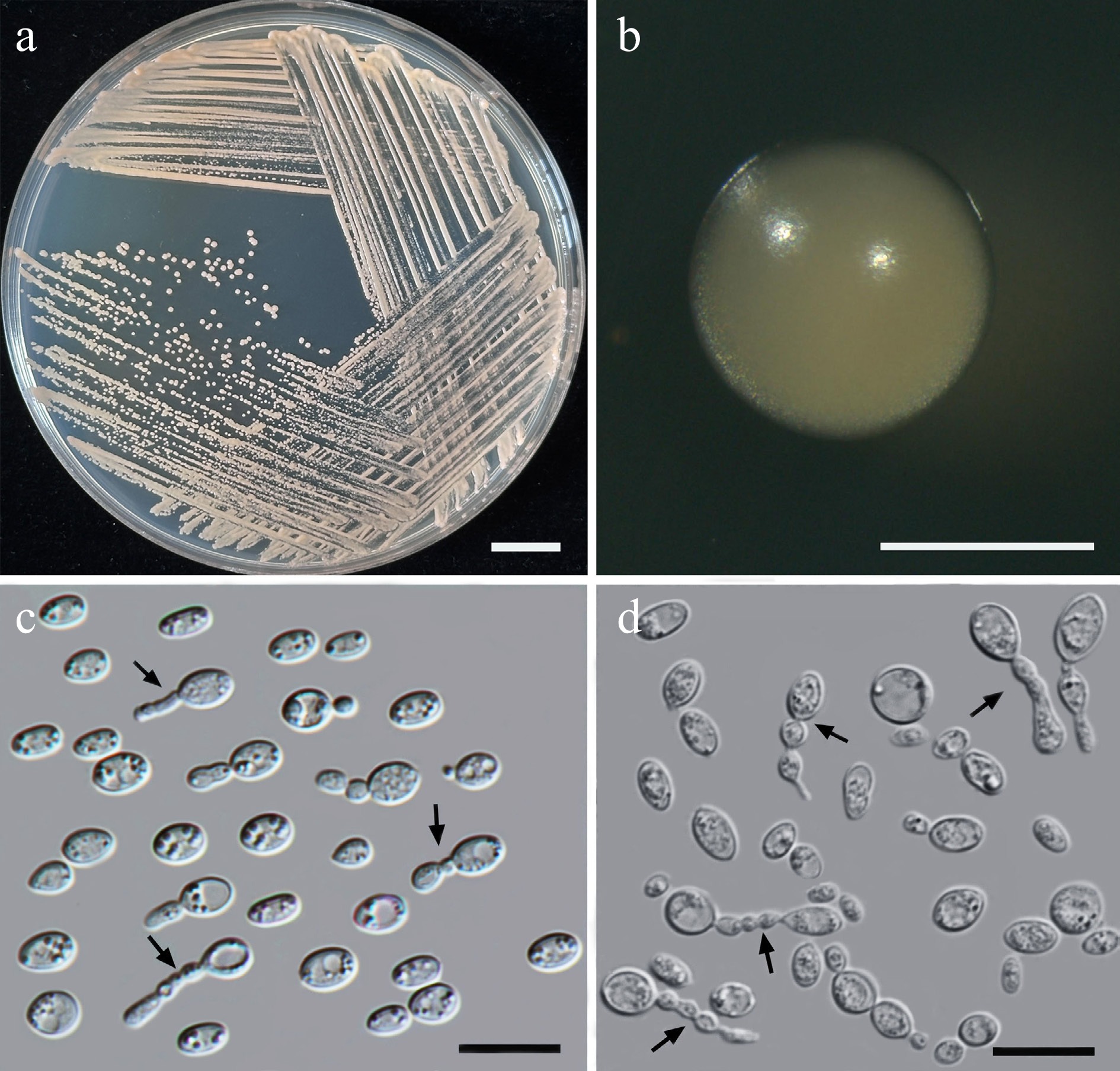
Figure 28.
Morphological characteristics of Halobasidium lannaense (SDBR-CMU612, ex-type). (a) Culture, (b) single colony, (c) chains of cells on YMA after 5 d at 25 °C, and (d) chains of cells (indicated by arrows) on YMA after 3 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
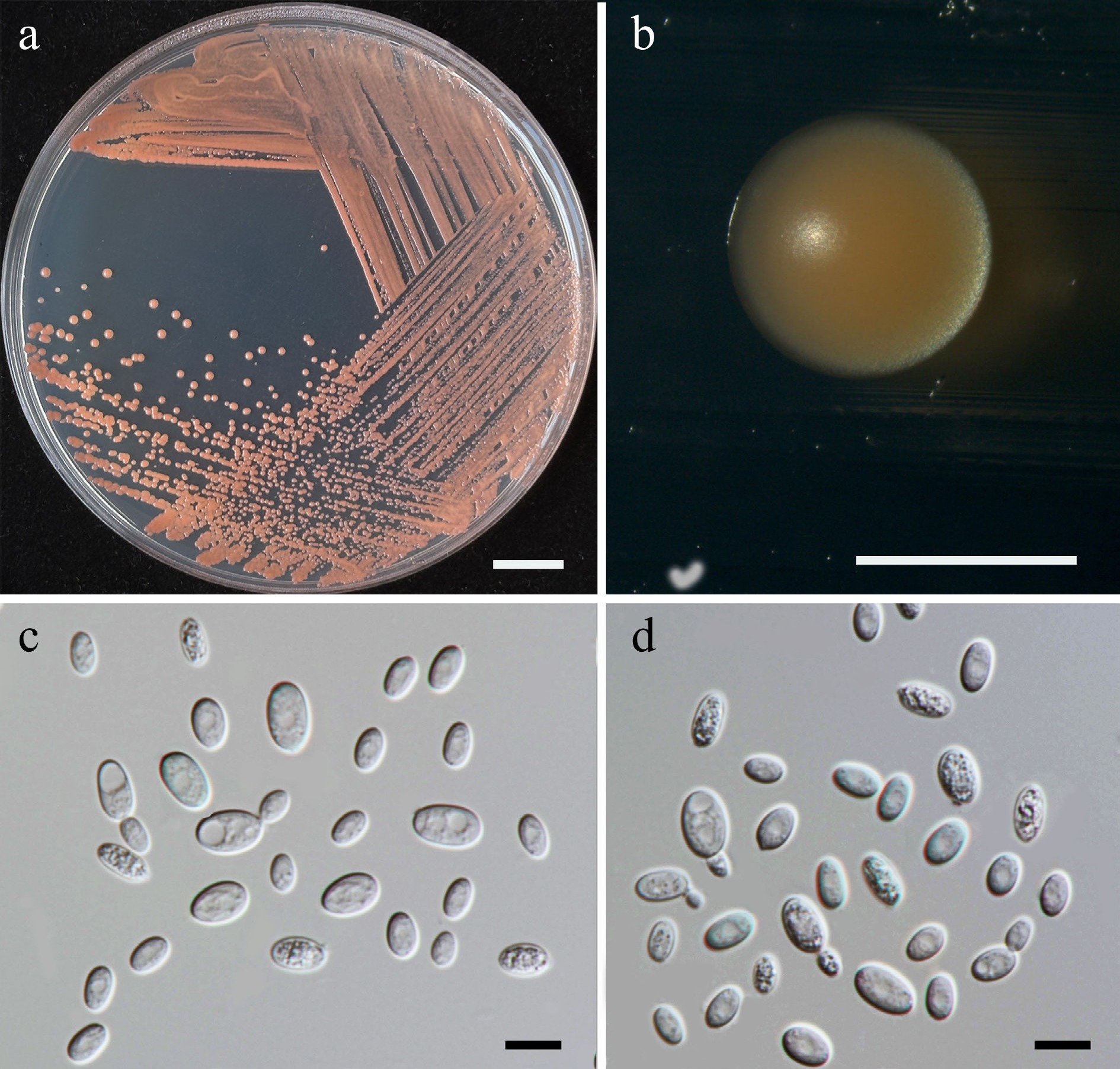
Figure 29.
Morphological characteristics of Symmetrospora chiangmaiensis (SDBR-CMU715, ex-type). (a) Culture, (b) single colony, and (c), (d) cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
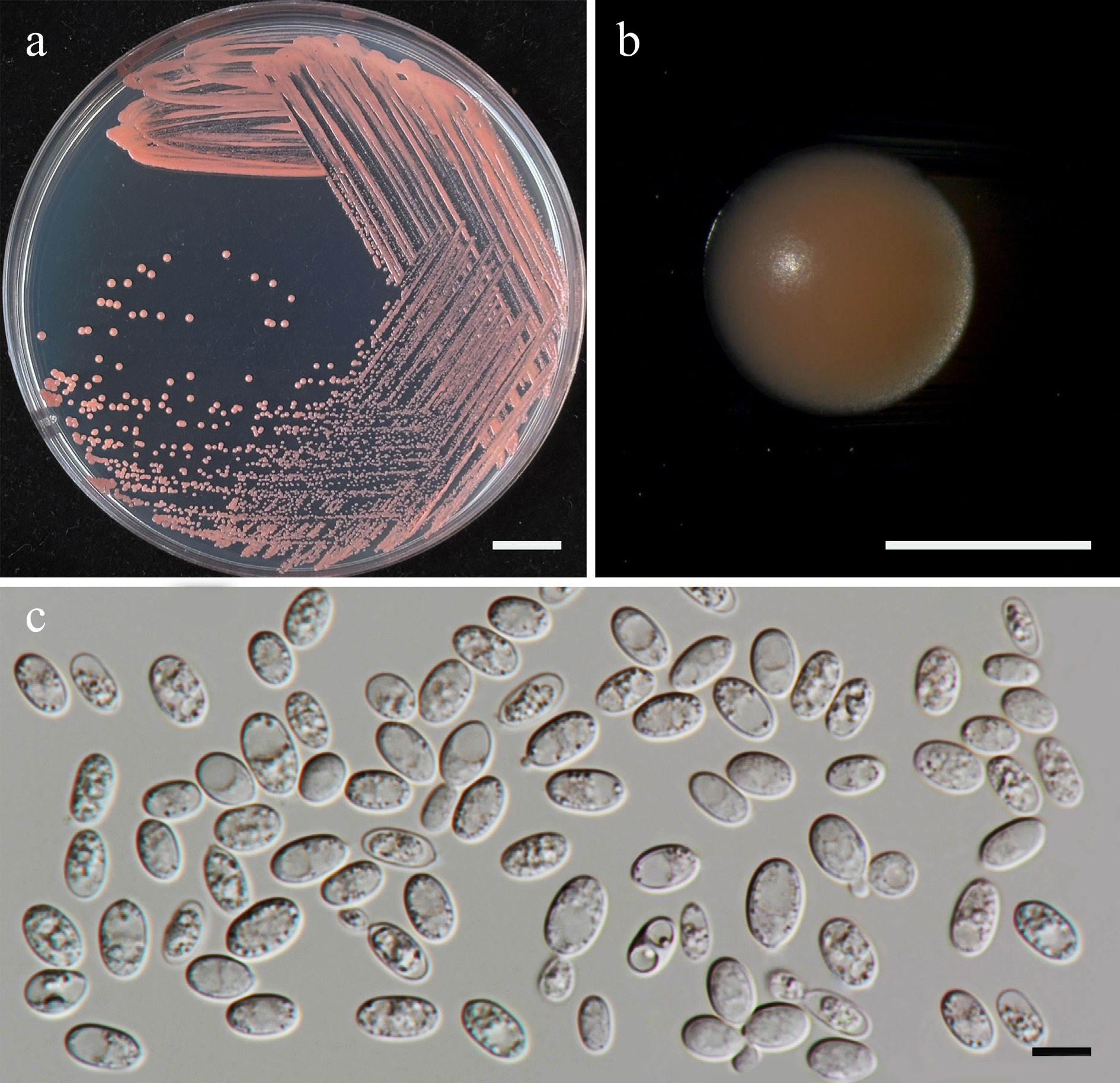
Figure 30.
Morphological characteristics of Symmetrospora hydei (SDBR-CMU620, ex-type). (a) Culture, (b) single colony, and (c) cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, and (c) = 10 μm.
-
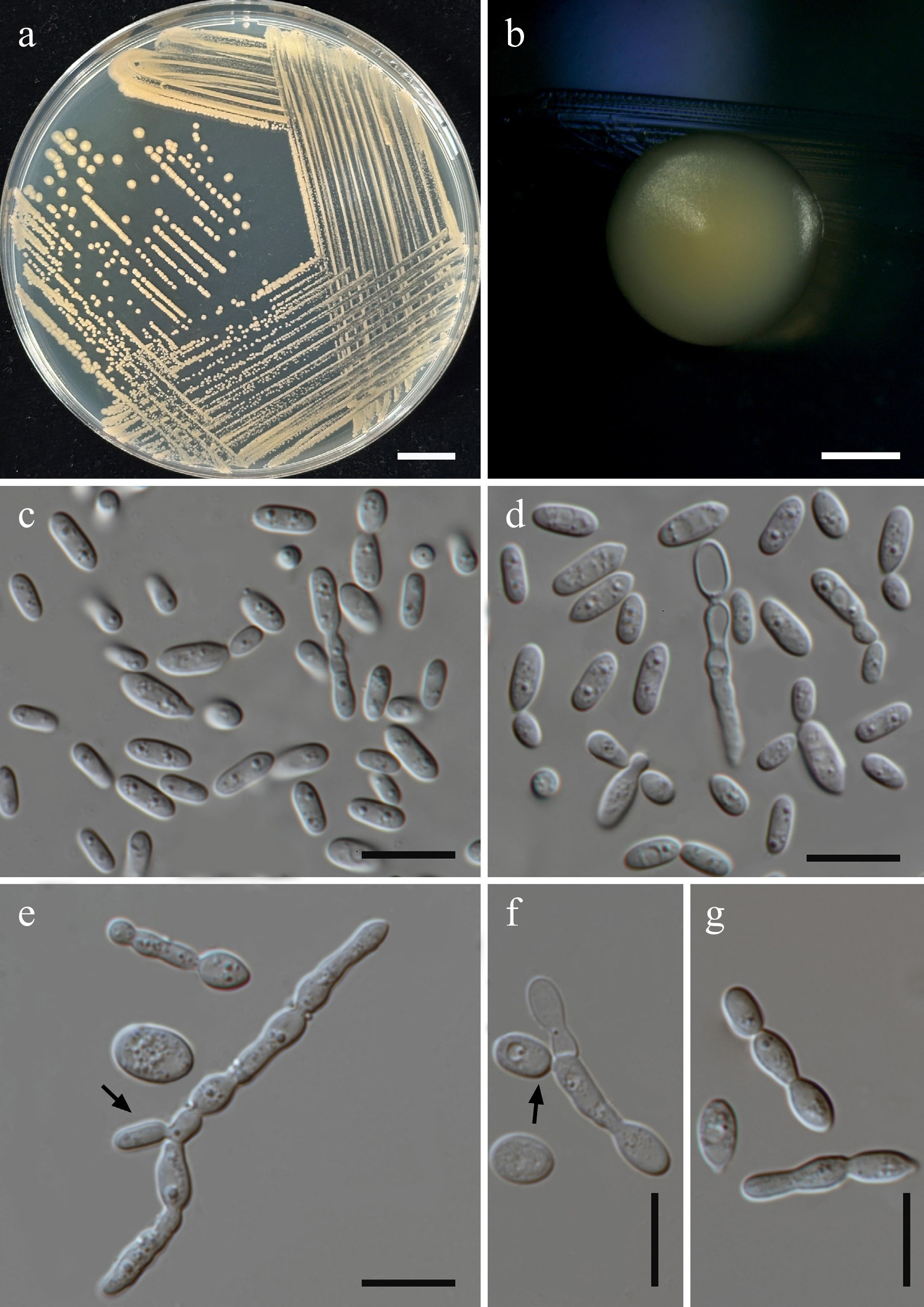
Figure 31.
Morphological characteristics of Curvibasidium chiangmaiense (SDBR-CMU574, ex-type). (a) Culture, (b) single colony, and (c), (d) cells on YMA after 5 d at 25 °C. (e), (f) Pseudohyphae and blastoconidia (indicated by arrows) on PDA after 2 weeks at 25 °C, and (g) elongate cells. Scale bars: (a) = 10 mm, (b) = 1 mm, and (c)–(g) = 10 μm.
-
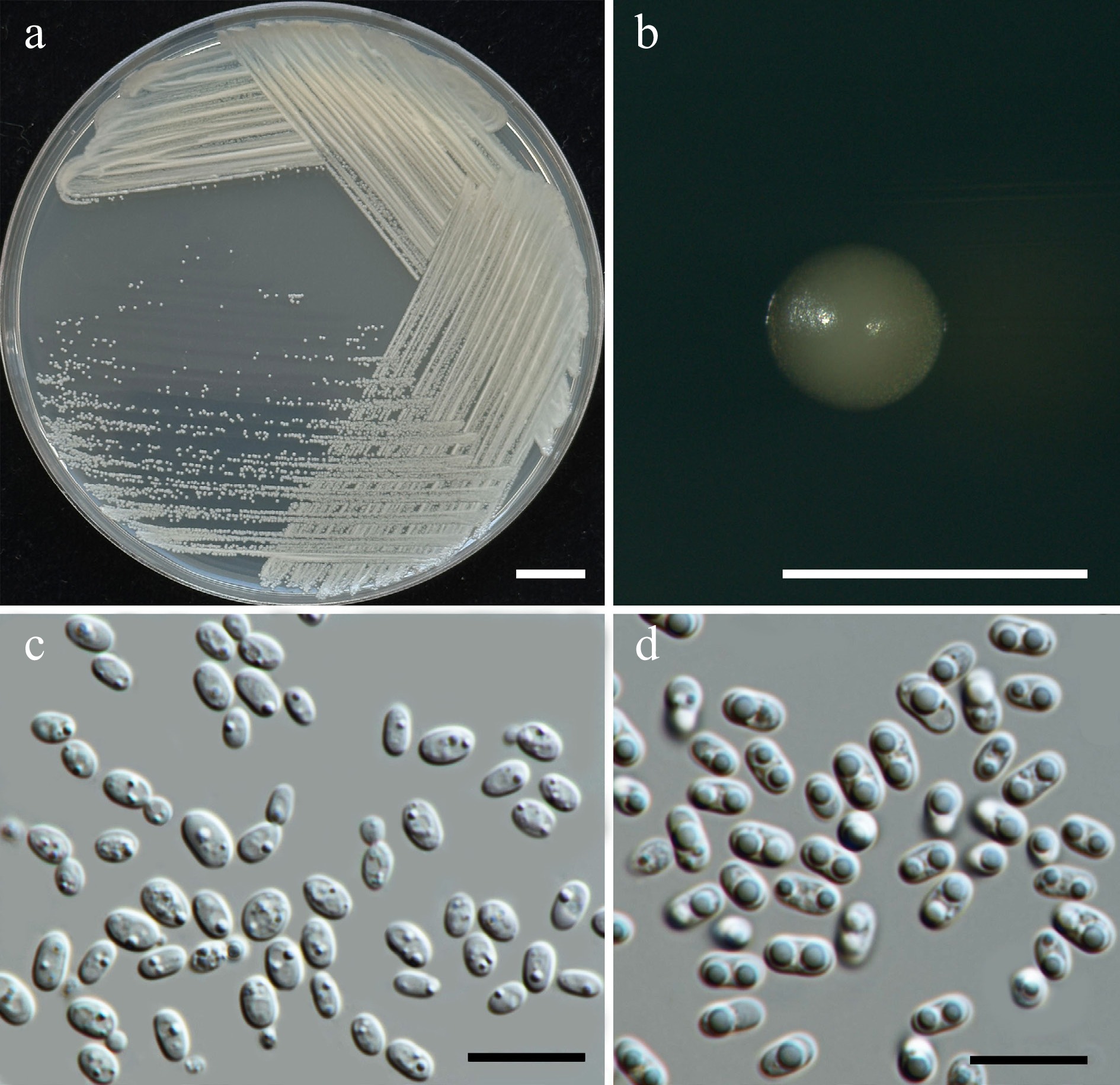
Figure 33.
Morphological characteristics of Thailandicola hydei (SDBR-CMU662, ex-type). (a) Culture, (b) single colony, (c), (d) cells and budding cells on YMA after 5 d at 25 °C. (d) Vegetative cells with one or two large globules on PDA at 25 °C for 2 weeks. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
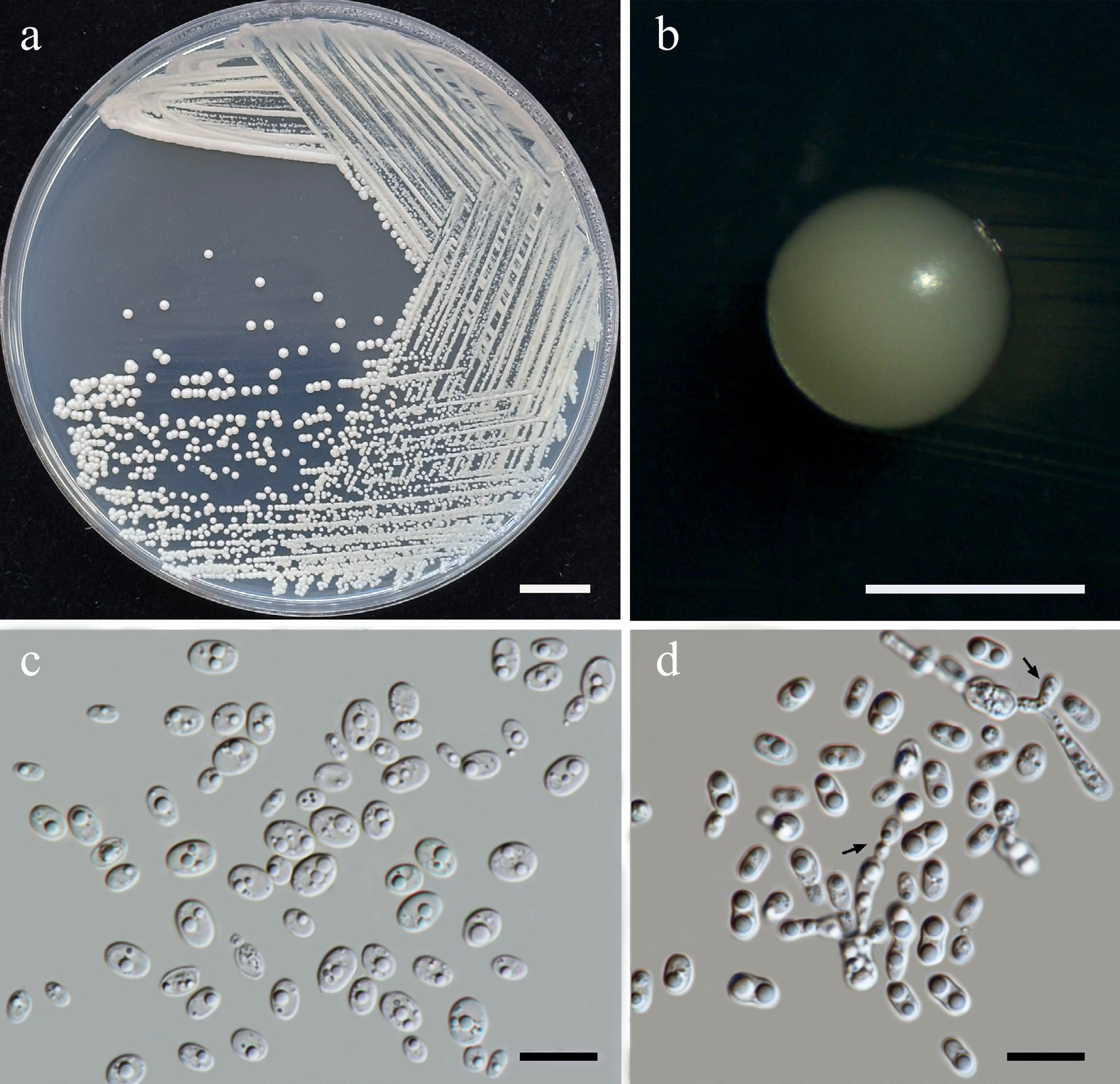
Figure 34.
Morphological characteristics of Thailandicola limtongiae sp. nov. (SDBR-CMU635, ex-type). (a) Culture, (b) single colony, (c) cells and budding cells on YMA after 5 d at 25 °C. (d) Vegetative cells with one or two large globules and pseudohyphae (indicated by arrows) on PDA at 25 °C for 2 weeks. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
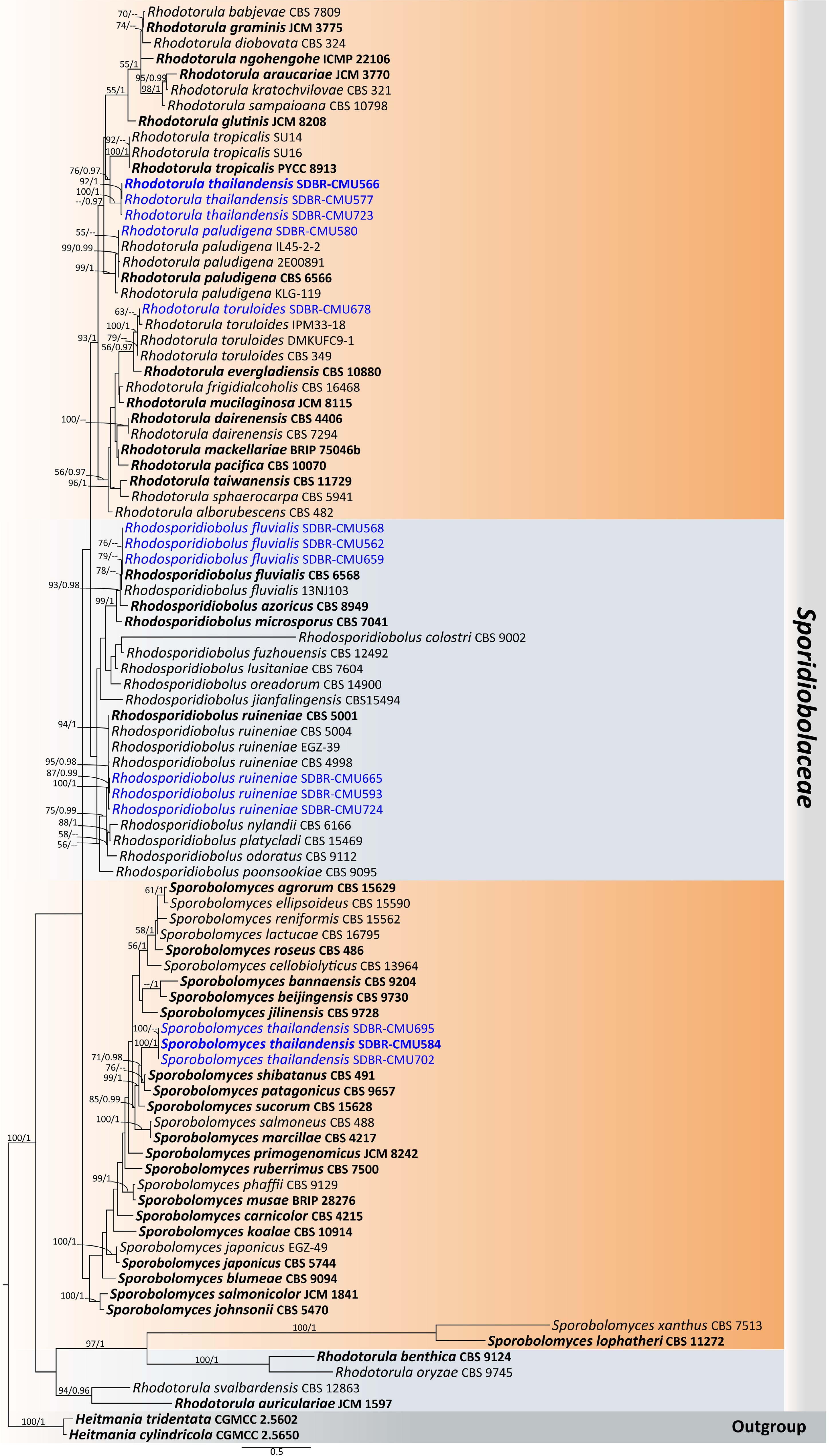
Figure 35.
Phylogram generated by maximum likelihood analysis of the combined D1/D2 domain of LSU, ITS, SSU, rpb1, rpb2, and tef1-α sequence data representing Sporidiobolaceae. The tree is rooted to Heitmania cylindrica (CGMCC 2.5650) and H. tridentata (CGMCC 2.5602). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Ninety-two strains are included in the combined sequence analysis, which comprise 7,114 characters with gaps. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
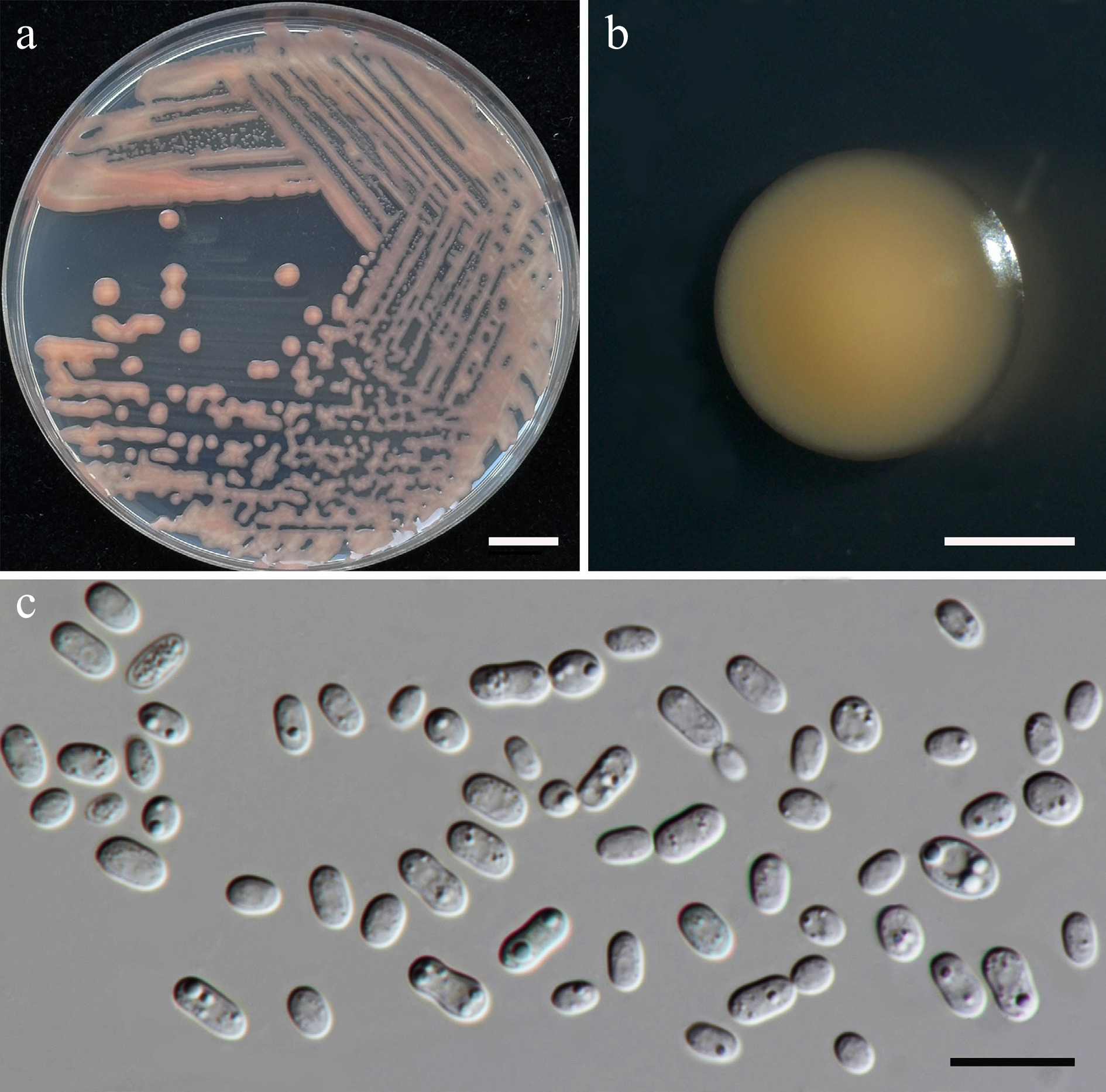
Figure 36.
Morphological characteristics of Rhodotorula thailandensis (SDBR-CMU566, ex-type). (a) Culture, (b) single colony, (c) cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, and (c) = 10 μm.
-
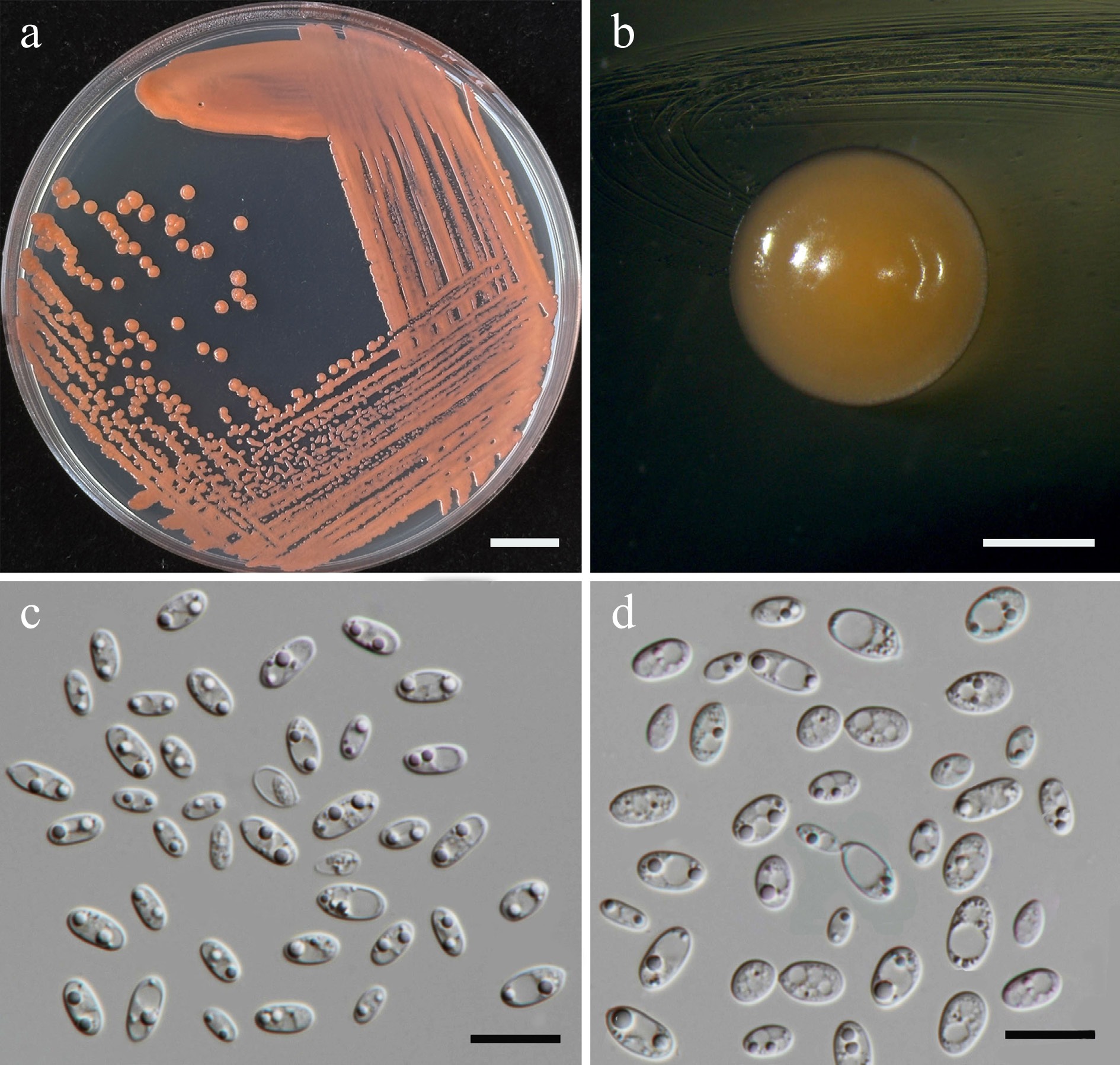
Figure 37.
Morphological characteristics of Sporobolomyces thailandensis (SDBR-CMU584, ex-type). (a) Culture, (b) single colony, (c) cells, and (d) budding cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
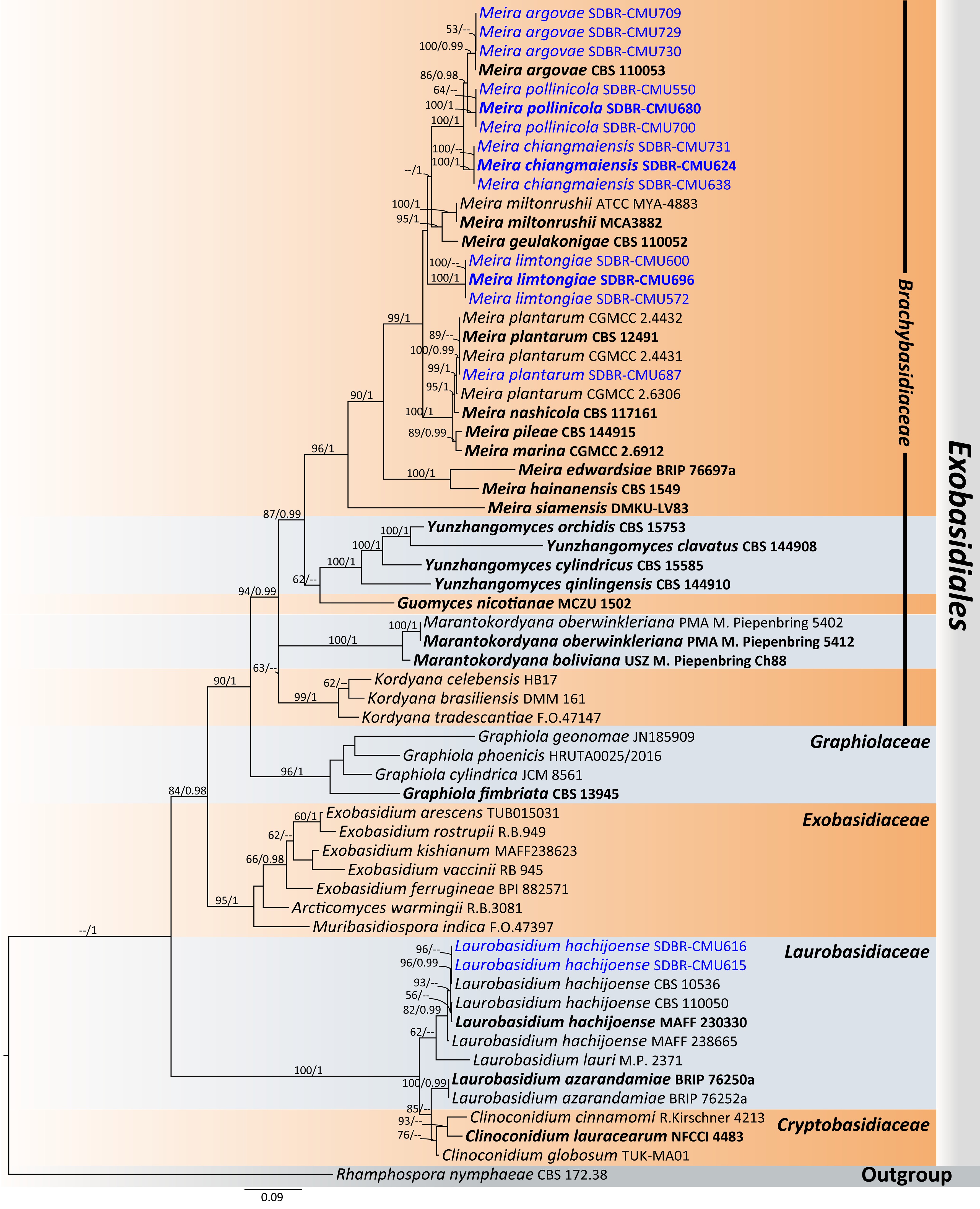
Figure 38.
Phylogram generated by maximum likelihood analysis of the combined D1/D2 domain of LSU, ITS, SSU, rpb1, rpb2, and tef1-α sequence data representing Meira and Laurobasidium in Exobasidiales. The tree is rooted to Rhamphospora nymphaeae (CBS 172.38). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Sixty-two strains are included in the combined sequence analysis, which comprise 5,633 characters with gaps. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
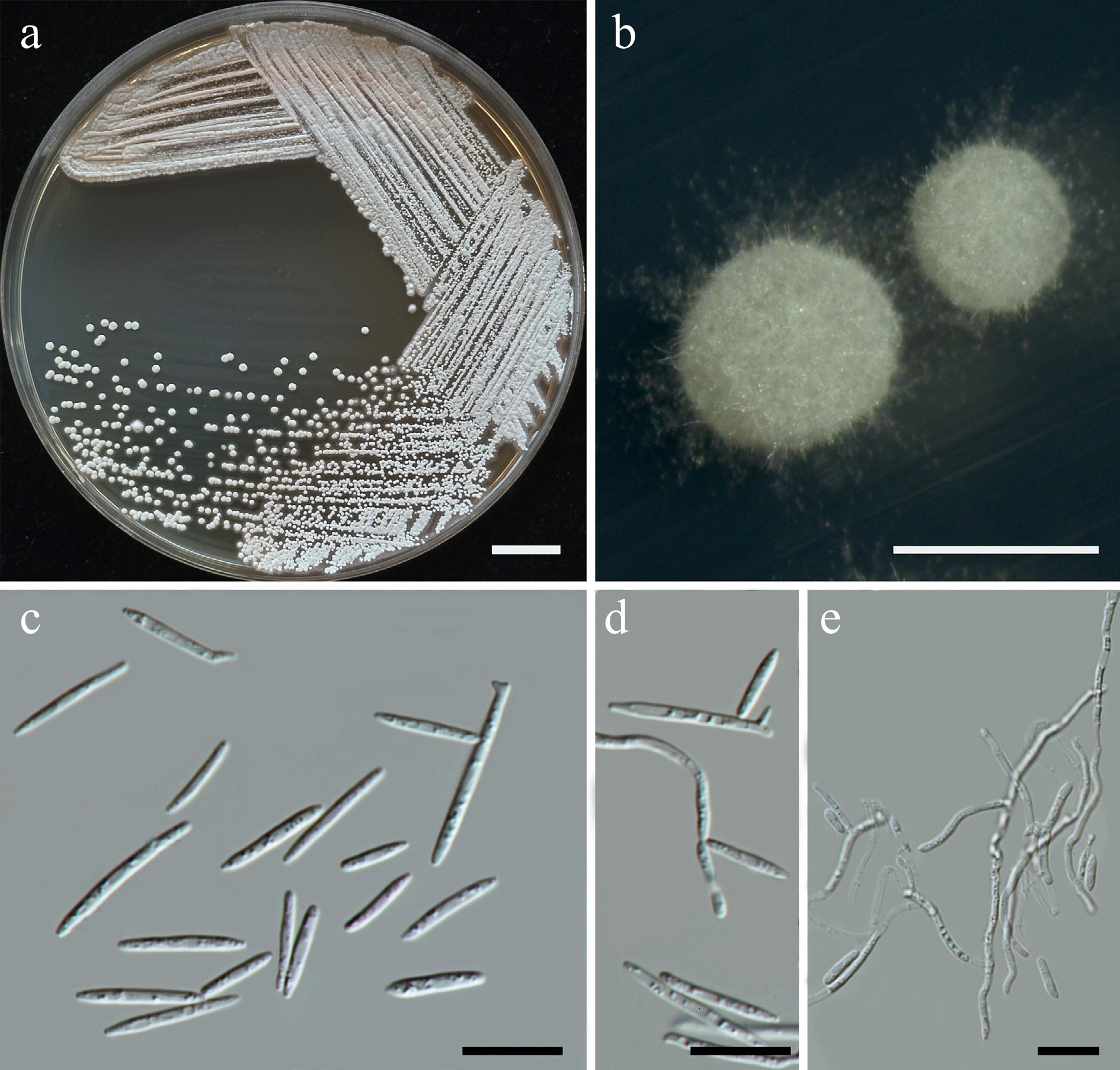
Figure 39.
Morphological characteristics of Meira chiangmaiensis (SDBR-CMU624, ex-type). Culture (a), single colony (b) and cells (c, d) on YMA after five days at 25 °C. Pseudohyphae on PDA after 2 weeks at 25 °C (d, e). Scale bars a = 10 mm, b = 1 mm, c and d = 10 μm.
-
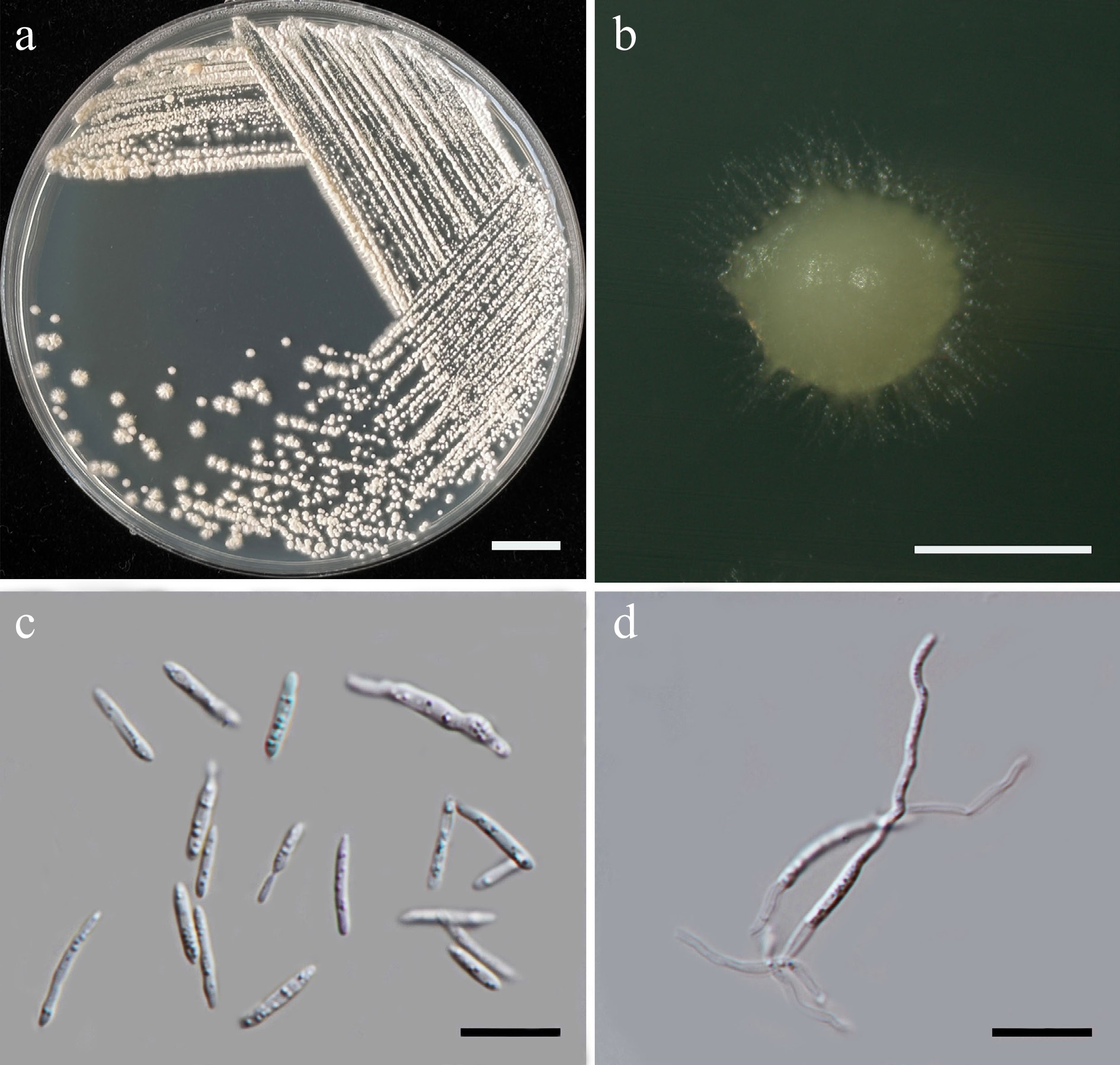
Figure 40.
Morphological characteristics of Meira limtongiae (SDBR-CMU696, ex-type). (a) Culture, (b) single colony, and (c) cells on YMA after 5 d at 25 °C. (d) Pseudohyphae on PDA after 2 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
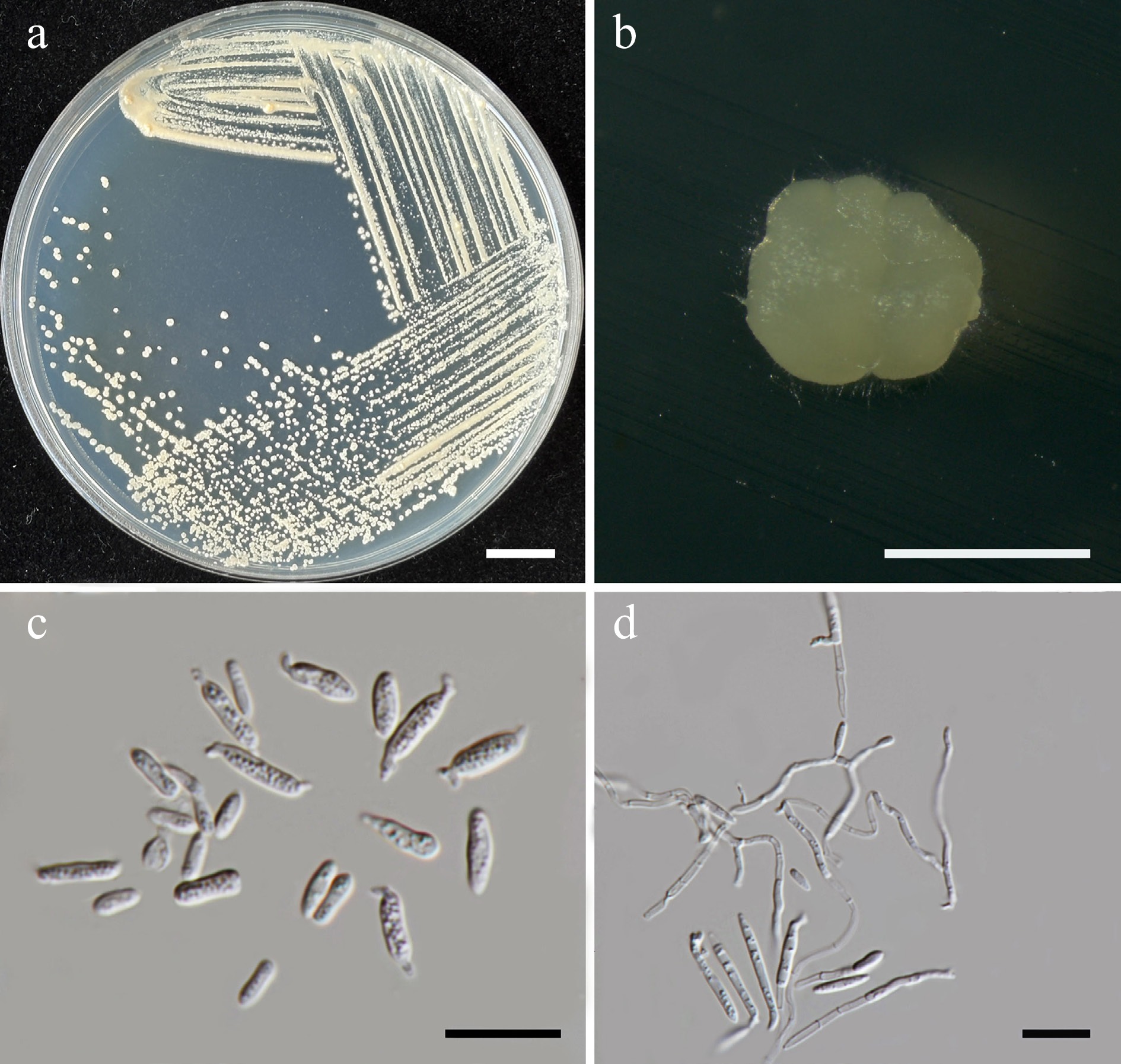
Figure 41.
Morphological characteristics of Meira pollinicola (SDBR-CMU680, ex-type). (a) Culture, (b) single colony, and (c) cells on YMA after 5 d at 25 °C. (d) Pseudohyphae on PDA after 2 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
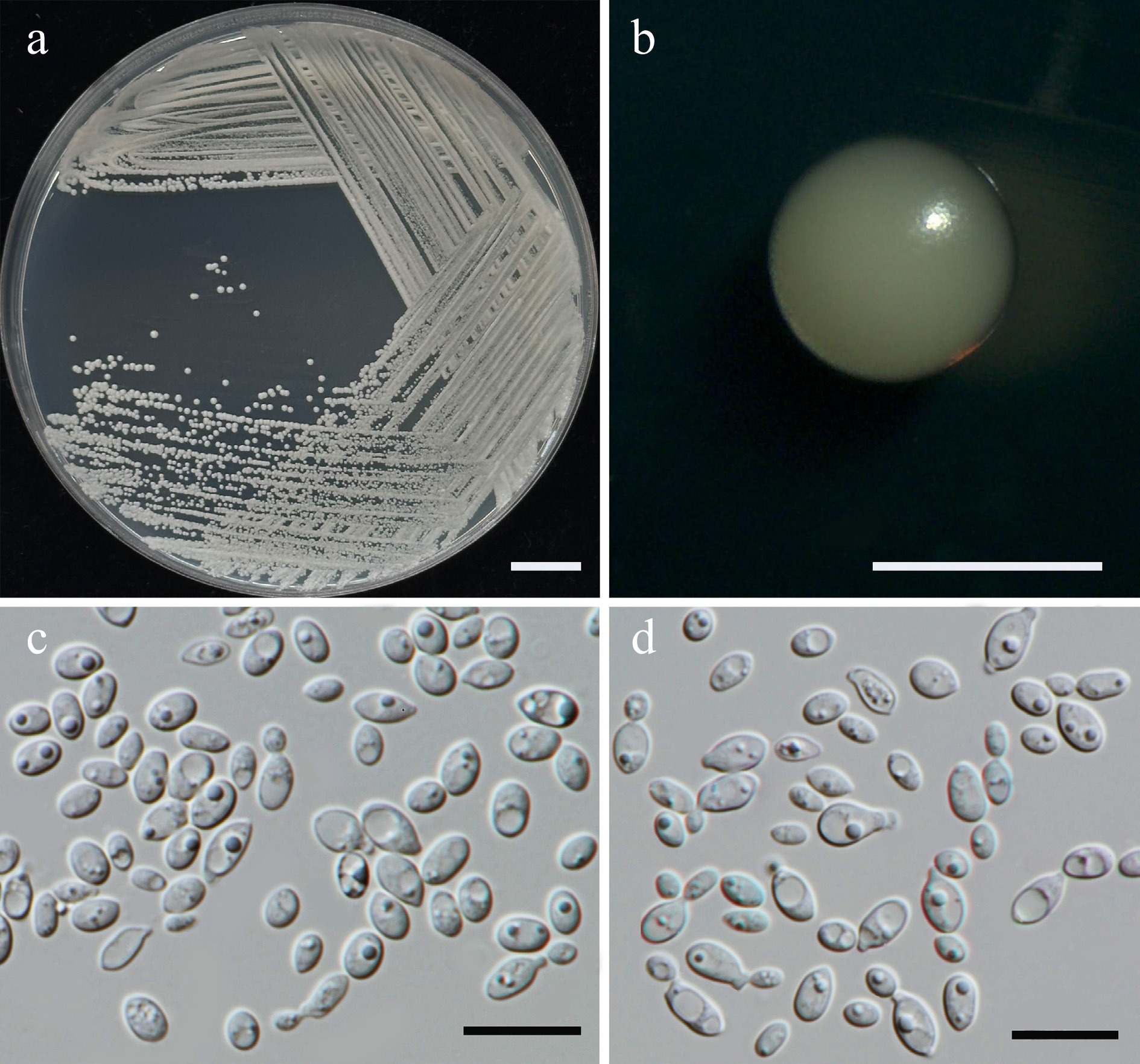
Figure 42.
Morphological characteristics of Parajaminaea hydei (SDBR-CMU601, ex-type). (a) Culture, (b) single colony, and (c), (d) cells on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
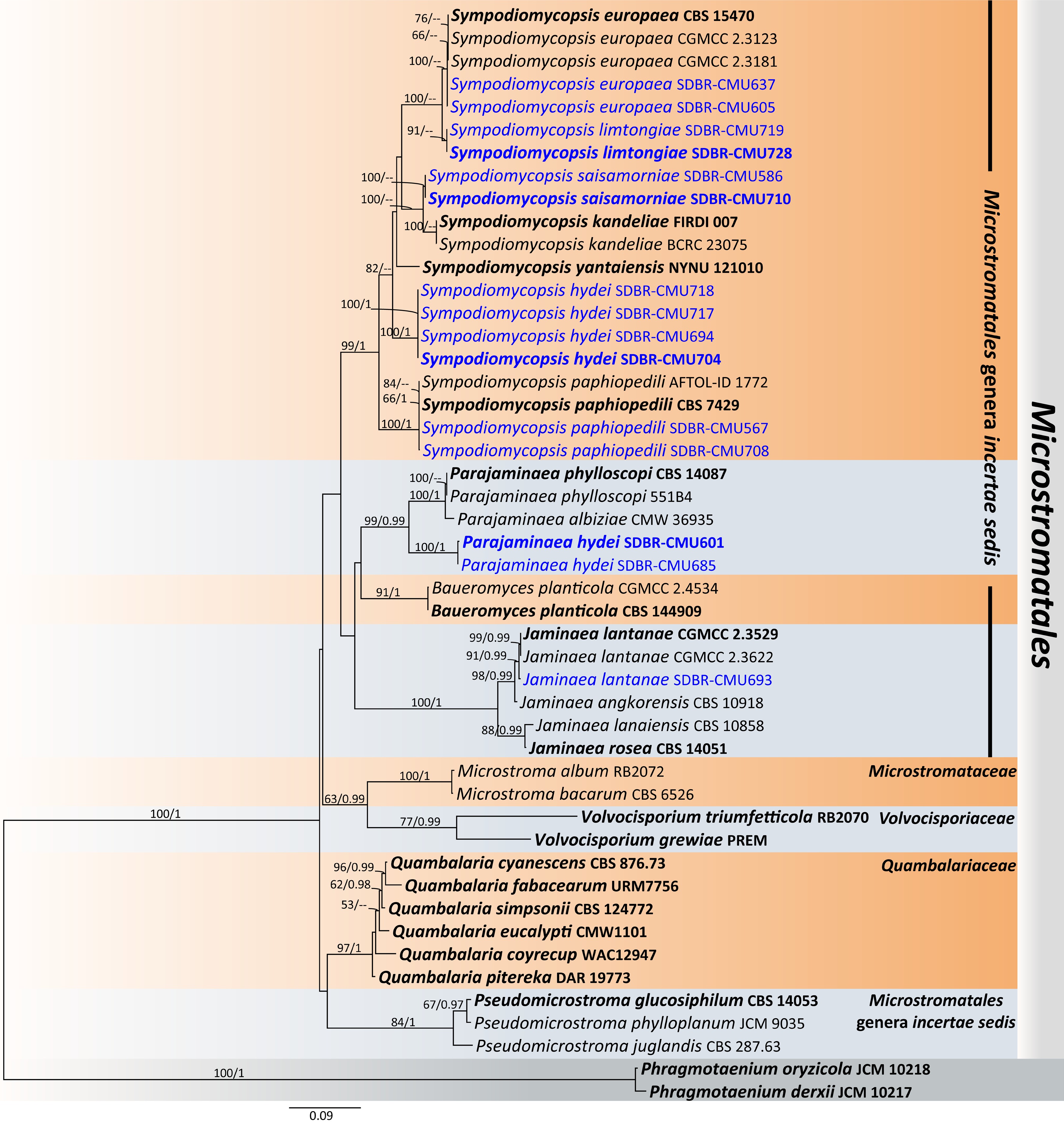
Figure 43.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Microstromatales. The tree is rooted to Phragmotaenium derxii (JCM 10217) and P. oryzicola (JCM 10218). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Fourty-eight strains are included in the combined sequence analysis, which comprise 1,326 characters with gaps. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.95 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.95 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
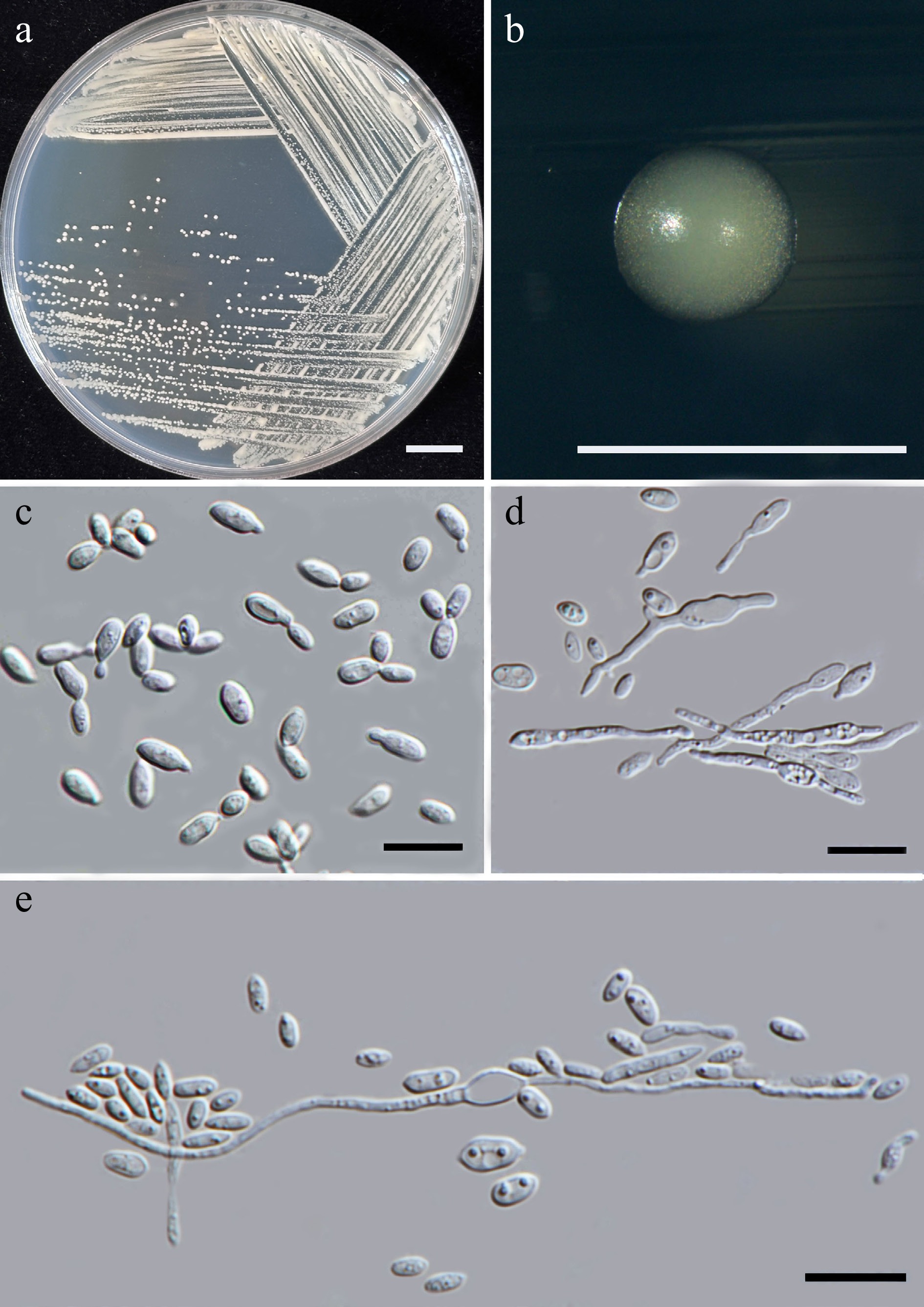
Figure 44.
Morphological characteristics of Sympodiomycopsis hydei (SDBR-CMU704, ex-type). (a) Culture, (b) single colony, and (c) cells on YMA at 25 °C for 5 d. (d), (e) Pseudohyphae on PDA after 2 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, and (c)–(e) = 10 μm.
-
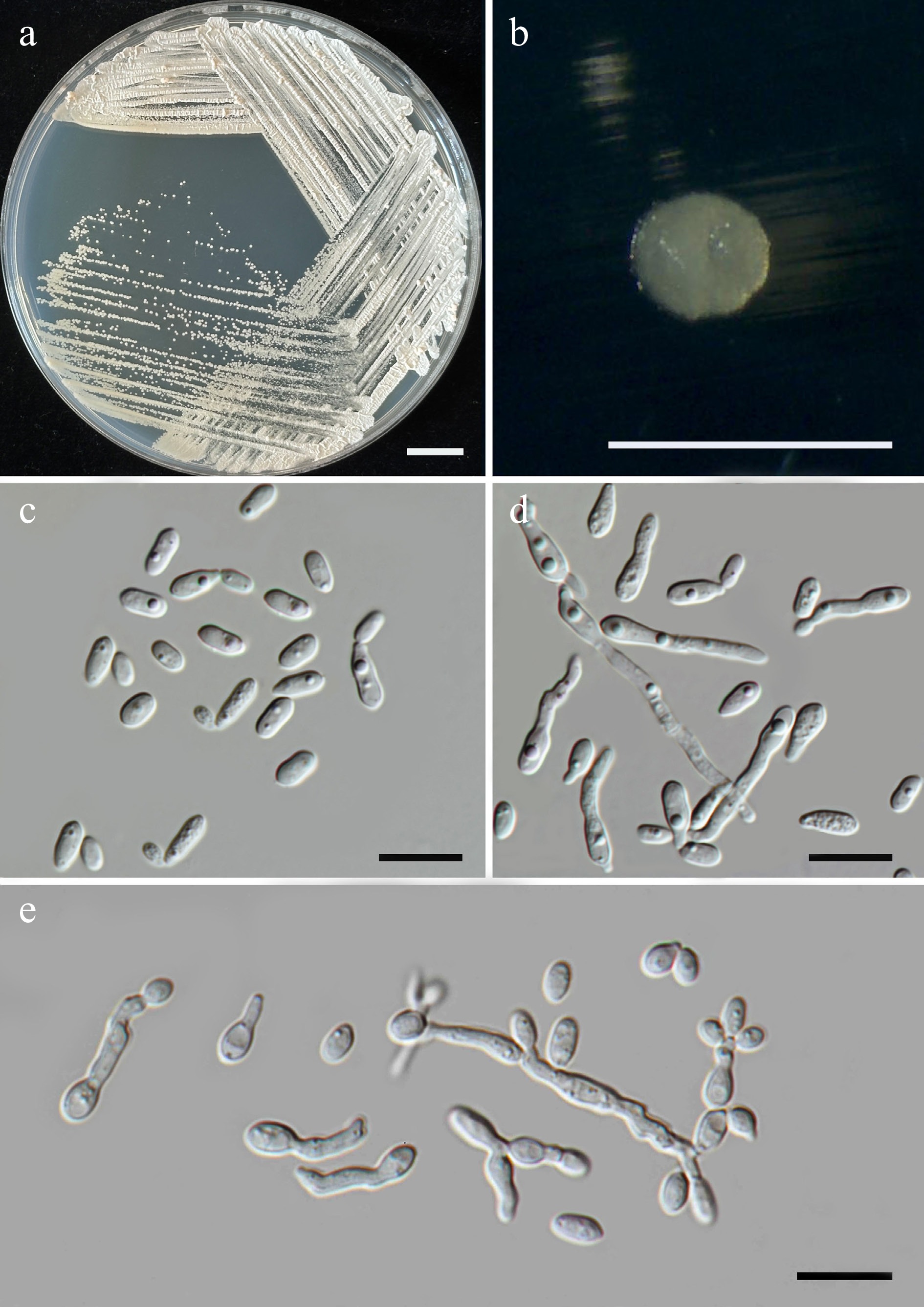
Figure 45.
Morphological characteristics of Sympodiomycopsis limtongiae (SDBR-CMU728, ex-type). (a) Culture, (b) single colony, and (c) cells on YMA at 25 °C for 5 d. (d), (e) Pseudohyphae on PDA after 2 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, and (c)–(e) = 10 μm.
-
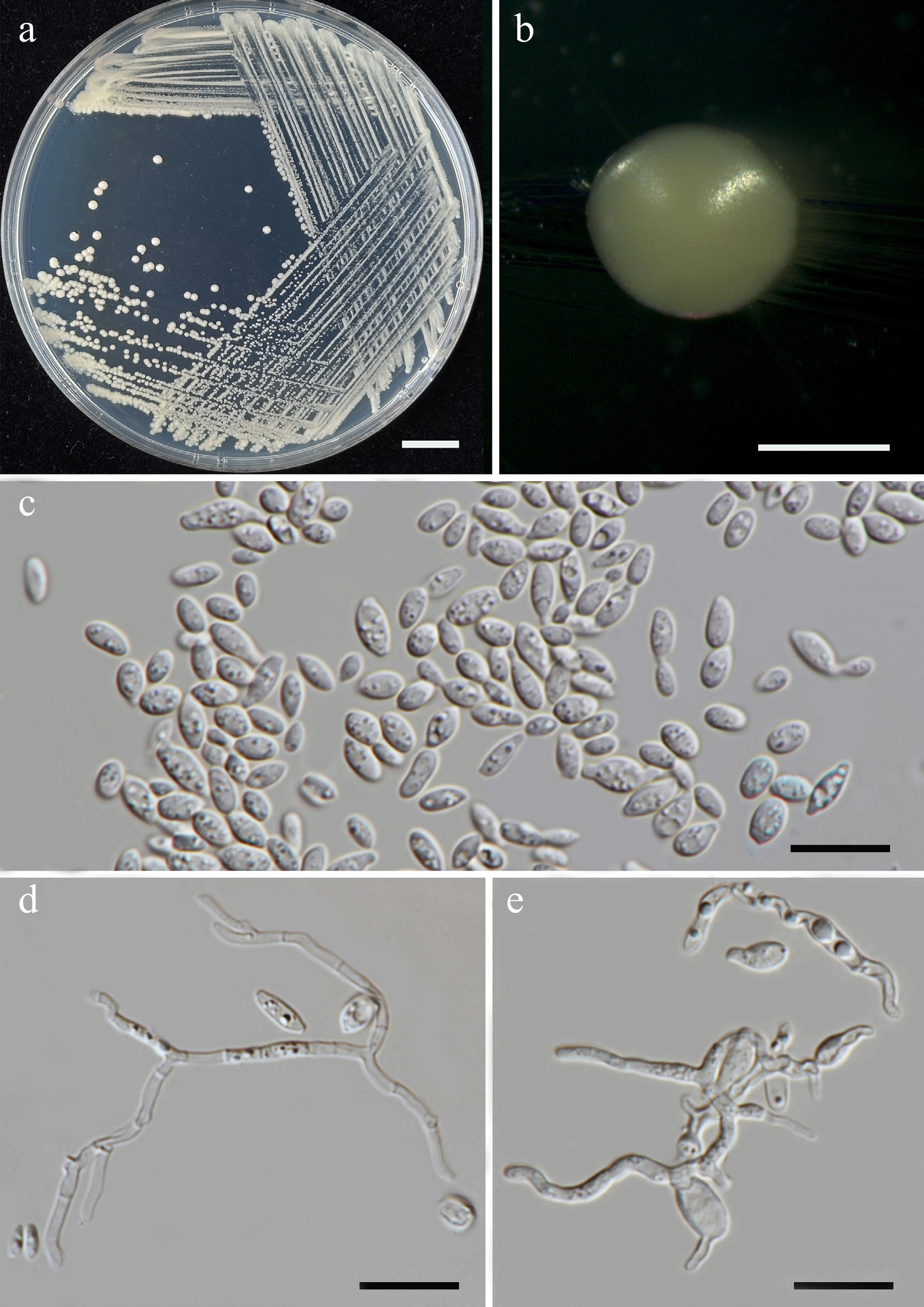
Figure 46.
Morphological characteristics of Sympodiomycopsis saisamorniae (SDBR-CMU710, ex-type). (a) Culture, (b) single colony, (c) cells and budding cells on YMA at 25 °C for 5 d. (d), (e) True hyphae and pseudohyphae on PDA after 2 weeks at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c)–(e) = 10 μm.
-
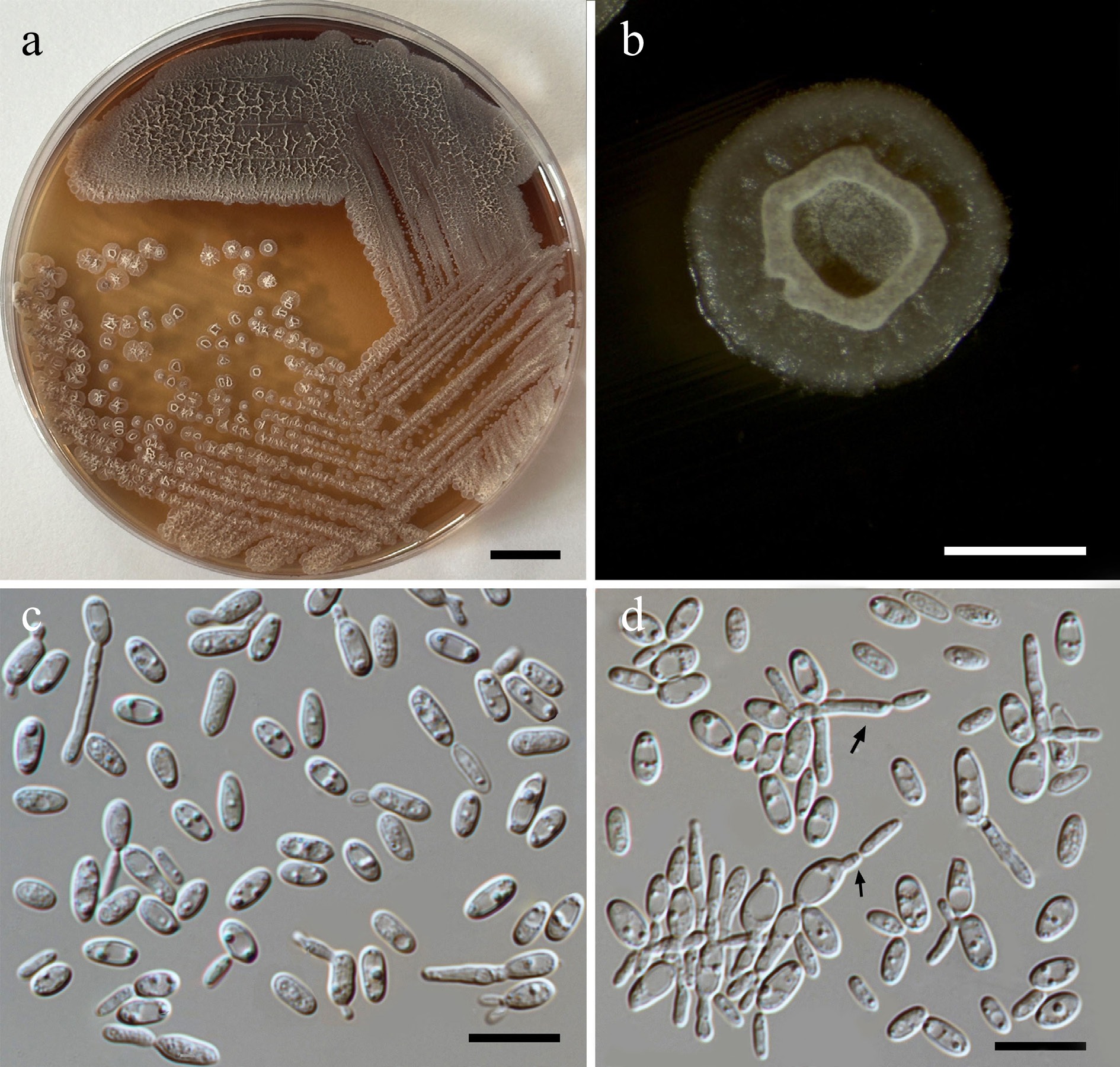
Figure 47.
Morphological characteristics of Pseudozyma chiangmaiensis (SDBR-CMU611, ex-type). (a) Culture, (b) single colony, (c), (d) cells rudimentary pseudohyphae (indicated by arrows) on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
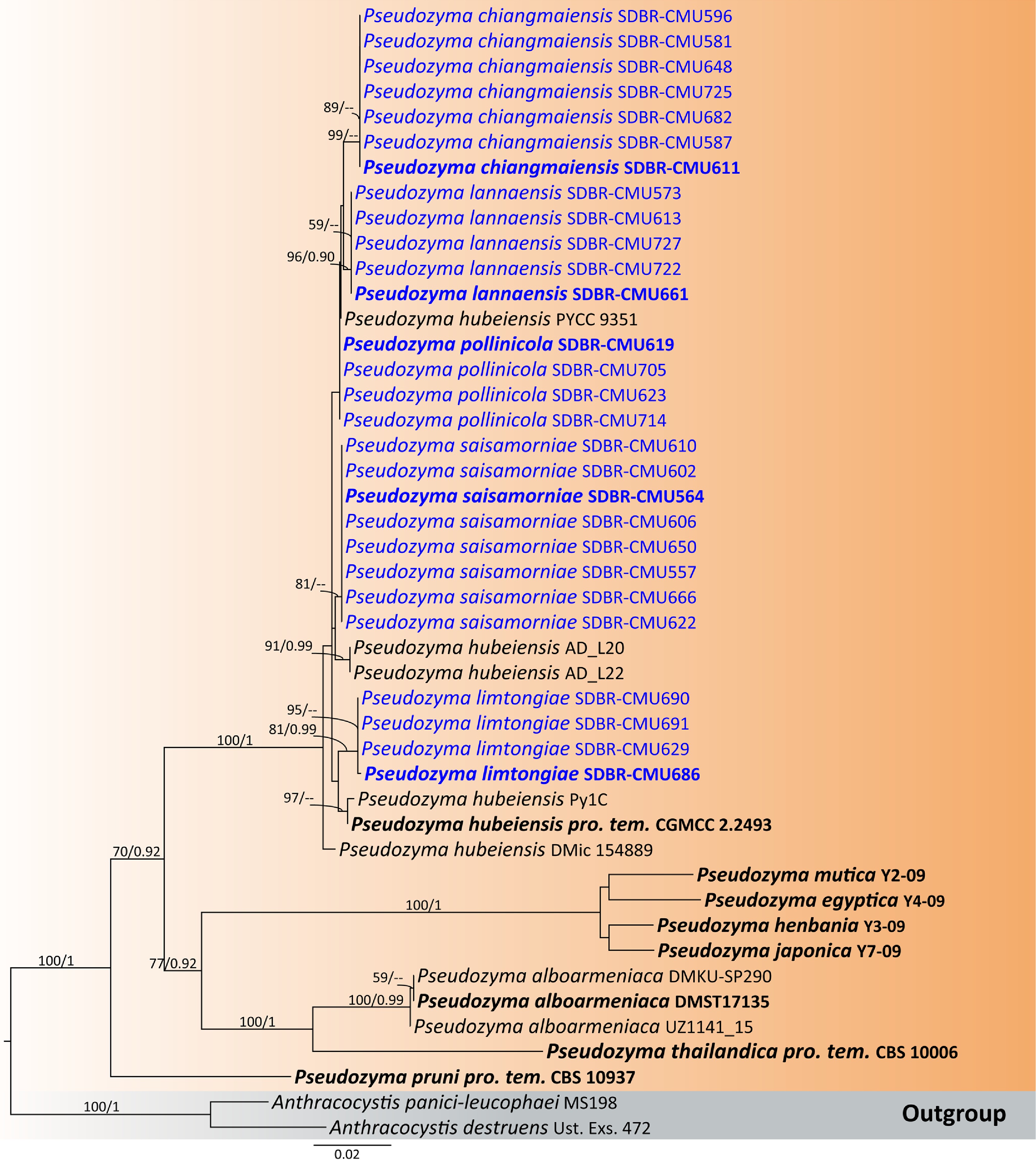
Figure 48.
Phylogenetic tree generated by maximum likelihood analysis of the combined D1/D2 domain of LSU and ITS sequence data representing Pseudozyma. The tree is rooted to Anthracocystis destruens (Ust. Exs. 472) and A. panici-leucophaei (MS198). Single-locus analyses were also performed, and topology and clade stability were compared from combined gene analyses. Fourty-five strains are included in the combined sequence analysis, which comprise 1,704 characters with gaps. The average standard deviation of the split frequencies of the BI analysis was 0.004541. Bootstrap support values for maximum likelihood ≥ 50% (ML, left) and Bayesian posterior probabilities ≥ 0.90 (PP, right) are indicated above the node. Double dashes (--) represent support values less than 50% ML/0.90 PP. The scale bar represents the expected number of nucleotide substitutions per site. The ex-type strains are in bold, and the newly generated sequences in this study are in blue.
-
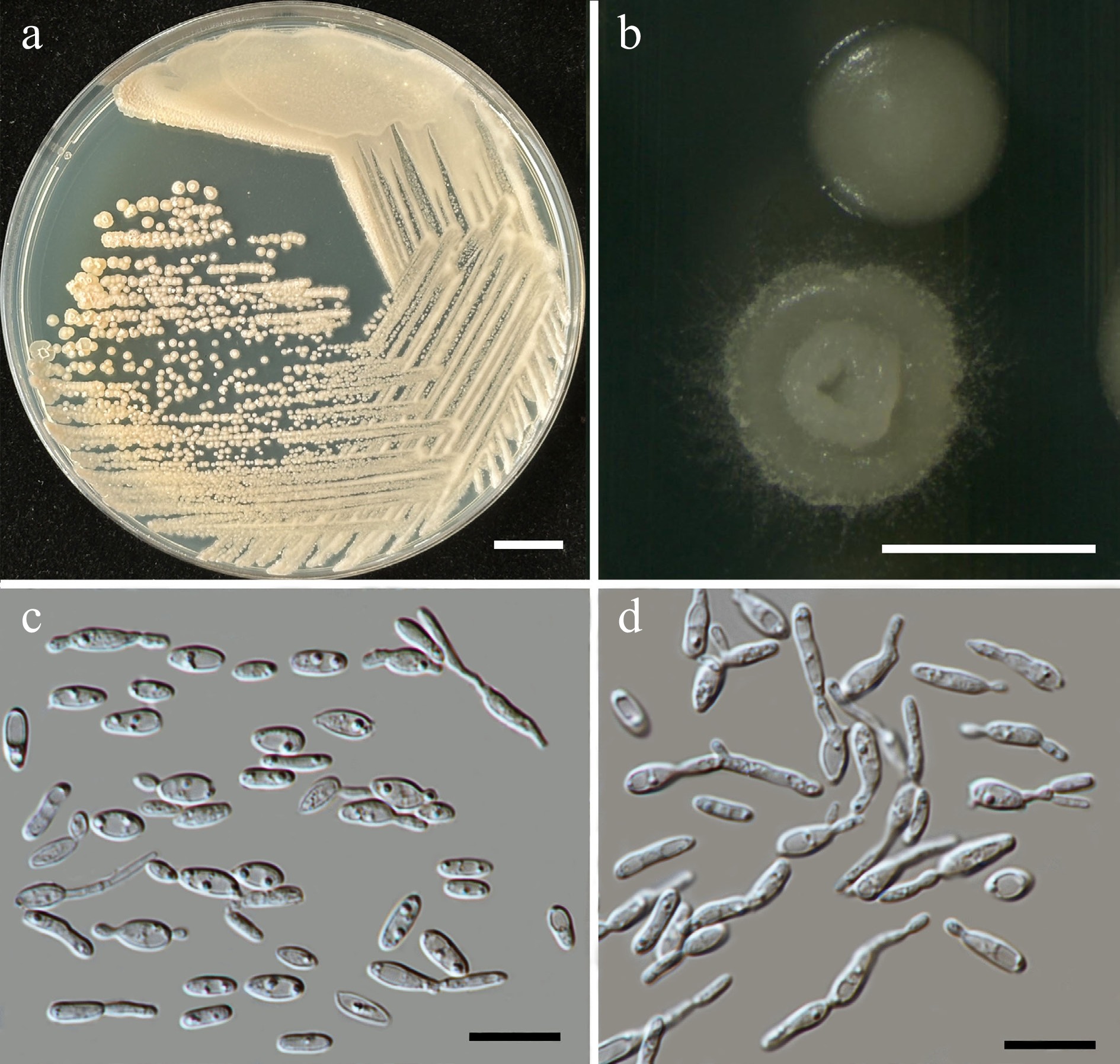
Figure 49.
Morphological characteristics of Pseudozyma lannaensis (SDBR-CMU661, ex-type). (a) Culture, (b) single colony, (c) cells, and (d) rudimentary pseudohyphae on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
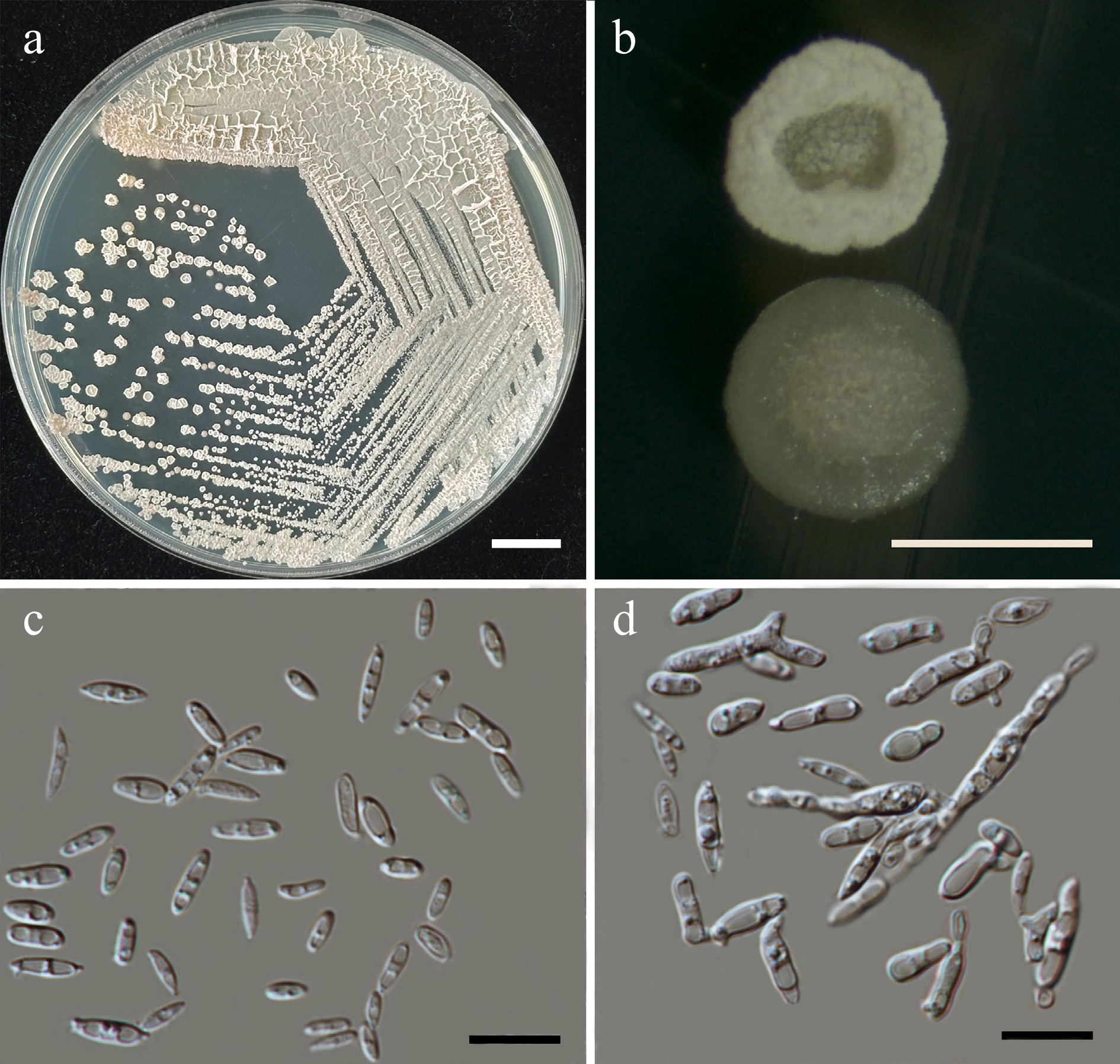
Figure 50.
Morphological characteristics of Pseudozyma limtongiae (SDBR-CMU686, ex-type). (a) Culture, (b) single colony, (c) cells, and (d) rudimentary pseudohyphae on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
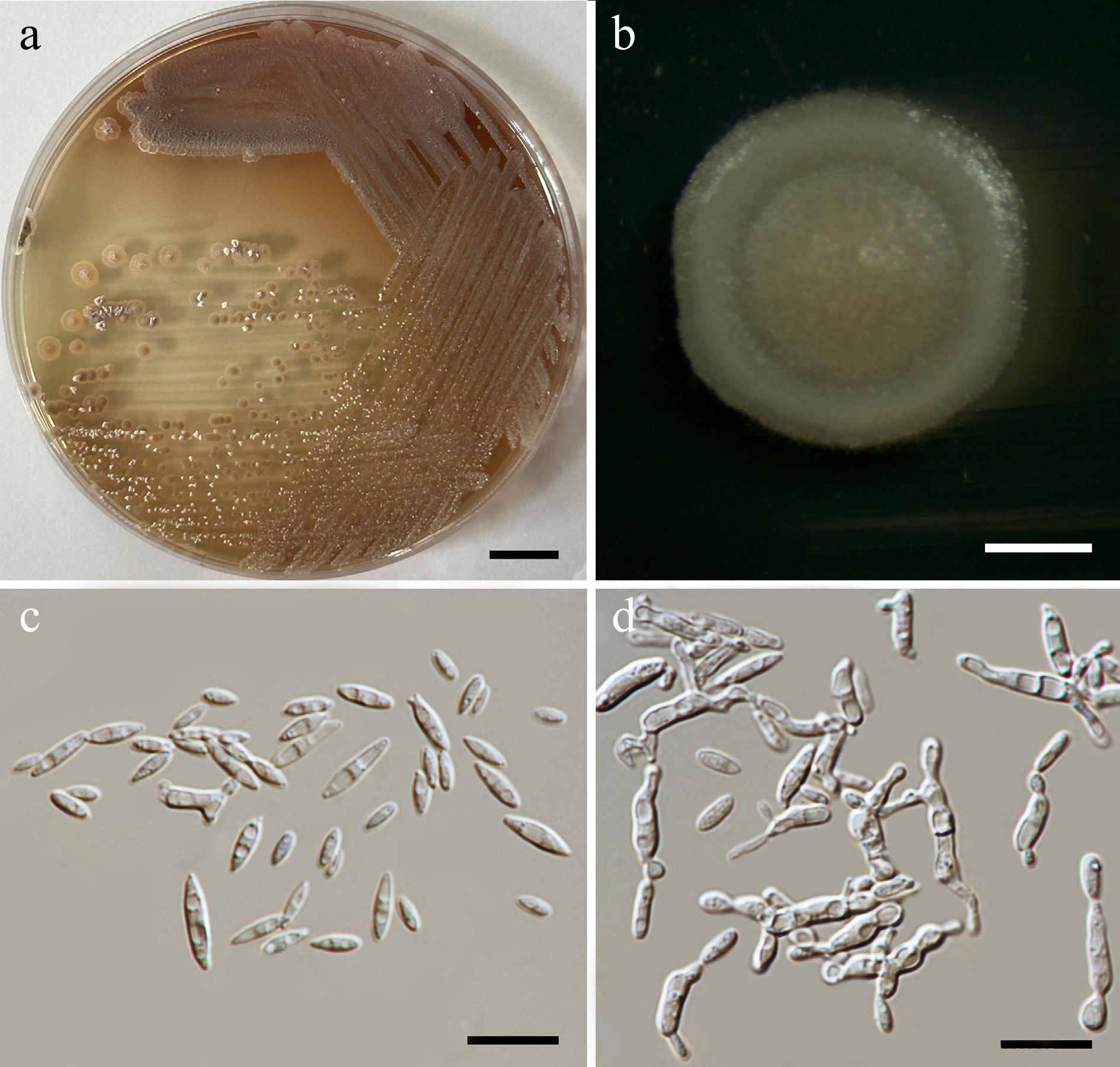
Figure 51.
Morphological characteristics of Pseudozyma pollinicola (SDBR-CMU619, ex-type). (a) Culture, (b) single colony, (c) cells, and (d) rudimentary pseudohyphae on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
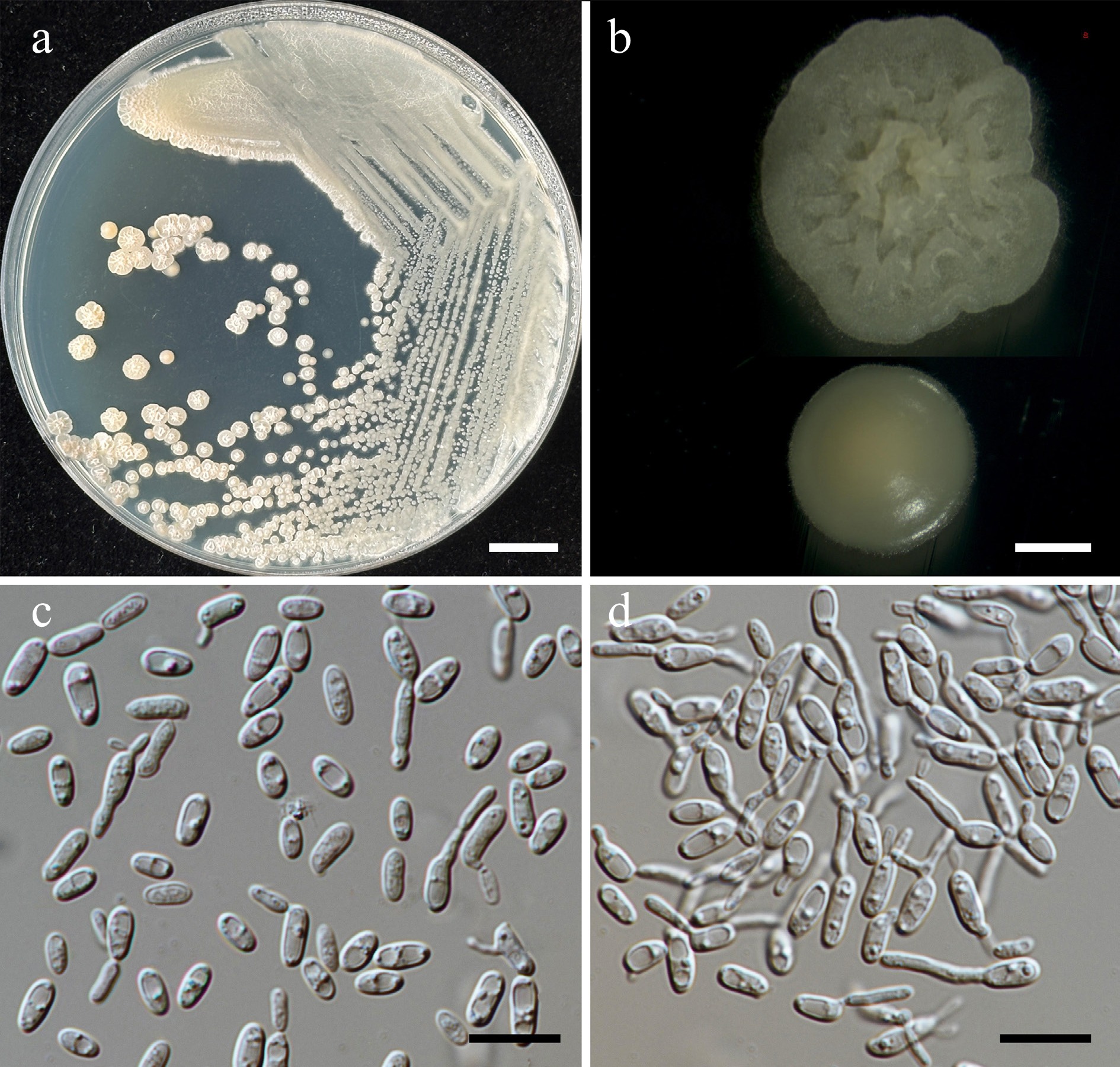
Figure 52.
Morphological characteristics of Pseudozyma saisamorniae (SDBR-CMU564, ex-type). (a) Culture, (b) single colony, (c) cells, and (d) rudimentary pseudohyphae on YMA after 5 d at 25 °C. Scale bars: (a) = 10 mm, (b) = 1 mm, (c), (d) = 10 μm.
-
Phylum: Ascomycota Caval. -Sm. Subphylum: Saccharomycotina O.E. Erikss. & Winka Class: Dipodascomycetes M. Groenew., Hittinger, Opulente & A. Rokas Order: Dipodascales M. Groenew., Hittinger, Opulente & A. Rokas Family: Trichomonascaceae Kurtzman & Robnett 1. Entelexis stigmatis Sipiczki ex Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov., (validation and first record in Thailand) 2. Starmerella etchellsii (Lodder & Kreger-van Rij) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1341 (2018) 3. Starmerella orientalis Alimad., Soudi, F.Y. Bai, S.A. Wang & Q.M. Wang ex Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov., (validation and first record in Thailand) 4. Starmerella thailandica Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 5. Wickerhamiella pollinicola Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Class: Pichiomycetes M. Groenew., Hittinger, Opulente & A. Rokas Order: Serinales M. Groenew., Hittinger, Opulente & A. Rokas Family: Debaryomycetaceae Kurtzman & M. Suzuki 6. Candida tropicalis (Castell.) Berkhout, De Schimmelgesl. Monilia, Oidium, Oospora en Torula, Disset. Ultrecht: 44 (1923) 7. Kodamaea ohmeri (Etchells & T.A. Bell) Y. Yamada, Tom. Suzuki, M. Matsuda & Mikata, Biosc., Biotechn., Biochem. 59: 1174 (1995) 8. Kodamaea restingae (C.A. Rosa, Lachance, Starmer, J.S.F. Barker, J.M. Bowles & Schlag-Edl.) C.Y. Chai & F.L. Hui, MycoKeys 89: 133 (2022) 9. Meyerozyma caribbica (Vaughan-Mart., Kurtzman, S.A. Mey. & E.B. O'Neill) Kurtzman & M. Suzuki, Mycoscience 51: 8 (2010) 10. Priceomyces siamensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Family: Metschnikowiaceae T. Kamieński ex Doweld 11. Metschnikowia cibodasensis Sjamsur., Oetari, C. Nakash., A. Kanti, Saraswati, & K. Ando, J. Microbiol. Biotechnol. 23: 909 (2013), (first record in Thailand) 12. Metschnikowia koreensis S.G. Hong, J. Chun, H.W. Oh & Bae, Int. J. Syst. Evol. Microbiol. 51: 1928 (2001) Class: Saccharomycetes G. Winter Order: Phaffomycetales M. Groenew., Hittinger, Opulente & A. Rokas Family: Phaffomycetaceae Y. Yamada, H. Kawas., Nagats., Mikata & T. Seki 13. Cyberlindnera fabianii (Wick.) Minter, Mycotaxon 110: 474 (2009) Order: Saccharomycodales M. Groenew., Hittinger, Opulente & A. Rokas Family: Saccharomycodaceae Kudryavtsev 14. Hanseniaspora lachancei Čadež, Poot, Raspor & M.T. Sm., Int. J. Syst. Evol. Microbiol. 53: 1679 (2003), (first record in Thailand) Phylum: Basidiomycota R.T. Moore Subphylum: Agaricomycotina Doweld Class: Tremellomycetes Doweld Order: Filobasidiales Jülich Family: Filobasidiaceae L.S. Olive 15. Filobasidium lannaense Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 16. Naganishia albida (Saito) Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 118 (2015) 17. Naganishia diffluens (Zach) Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 119 (2015) 18. Naganishia liquefaciens (Saito & M. Ota) Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 119 (2015) Order: Tremellales Fr. Family: Bulleribasidiaceae Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout 19. Hannaella pagnoccae Landell, L.R. Brandão, A.C. Barbosa, J.P. Ramos, Safar, F.C.O. Gomes, F.M.P. Sousa, P.B. Morais, Broetto, Leoncini, J.R.A. Ribeiro, Fungsin, M. Takash., Nakase, C.F. Lee, Vainstein, Fell, Scorzetti, Vishniac, C.A. Rosa & P. Valente, Int. J. Syst. Evol. Microbiol. 64: 1975 (2013) 20. Hannaella phyllophila Suruss. & Limtong, Int. J. Syst. Evol. Microbiol. 65: 2138 (2014) 21. Vishniacozyma marinae Q.M. Wang, Stud. Mycol.109: 119 (2024) 22. Vishniacozyma pollinicola Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Family: Cryptococcaceae Kütz. ex Castell. & Chalm. 23. Kwoniella bestiolae (Thanh, Hai & Lachance) Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 137 (2015), (first record in Thailand) 24. Kwoniella heveanensis Metin, K. Findley & Heitman, Mycotaxon 116: 227 (2011) 25. Kwoniella limtongiae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 26. Kwoniella saisamorniae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Family: Rhynchogastremaceae Oberw. & B. Metzler 27. Papiliotrema aspenensis (Ferreira-Paim, T.B. Ferreira, Andrade-Silva, D.J. Mora, D.J. Springer, Heitman, F.M. Fonseca, D. Matos, Melhem & Silva-Verg.) Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 96: 135 (2020) 28. Papiliotrema flavescens (Saito) Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 126 (2015) 29. Papiliotrema chiangmaiensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 30. Papiliotrema pollinicola Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 31. Papiliotrema tectonae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Family: Trimorphomycetaceae Xin Zhan Liu, F.Y. Bai, M. Groenew. & Boekhout 32. Saitozyma thailandensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Order: Trichosporonales Boekhout & Fell Family: Trichosporonaceae Nann. 33. Trichosporon asahii Akagi ex Sugita, A. Nishikawa & Shinoda, J. Gen. Appl. Microbiol., Tokyo 40: 405 (1994) Subphylum: Pucciniomycotina R. Bauer, Begerow, J.P. Samp., M. Weiss & Oberw. Class: Agaricostilbomycetes R. Bauer, Begerow, J.P. Samp., M. Weiss & Oberw. Order: Agaricostilbales Oberw. & R. Bauer Family: Chionosphaeraceae Oberw. & Bandoni 34. Boekhoutia pollinicola Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Class: Cystobasidiomycetes R. Bauer, Begerow, J.P. Samp., M. Weiss & Oberw. Order: Cystobasidiales R. Bauer, Begerow, J.P. Samp., M. Weiss & Oberw. Family: Cystobasidiaceae Gäum. 35. Cystobasidium benthicum (Nagah., Hamam., Nakase & Horikoshi) Yurkov, Kachalkin, H.M. Daniel, M. Groenew., Libkind, V. de García, Zalar, Gouliam., Boekhout & Begerow, Antonie van Leeuwenhoek 107: 180 (2014), (new habitat record and first record in Thailand) 36. Cystobasidium keelungense C.F. Chang & S.M. Liu, Arch. Microbiol. 201: 31 (2018), (first record in Thailand) 37. Cystobasidium minutum (Cif. & Redaelli) Yurkov, Kachalkin, H.M. Daniel, M. Groenew., Libkind, V. de García, Zalar, Gouliam., Boekhout & Begerow, Antonie van Leeuwenhoek 107: 180 (2014) 38. Cystobasidium thailandicum Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 39. Halobasidium lannaense Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Class: Cystobasidiomycetes R. Bauer, Begerow, J.P. Samp., M. Weiss & Oberw. Order: Erythrobasidiales R. Bauer, Begerow, J.P. Samp., M. Weiss & Oberw. Family: Erythrobasidiaceae Denchev 40. Erythrobasidium primogenitum Y.P. Tan, Bishop-Hurley & R.G. Shivas, Index of Australian Fungi 1: 1 (2022), (first record in Thailand) Families Incertae sedis Family: Symmetrosporaceae Q.M. Wang, F.Y. Bai, M. Groenew. & Boekhout 41. Symmetrospora chiangmaiensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 42. Symmetrospora hydei Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Class: Microbotryomycetes R. Bauer, Begerow, J.P. Samp., M. Weiss & Oberw. Order: Incertae sedis Families Incertae sedis 43. Curvibasidium chiangmaiense Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Order: Thailandicolales Kodchasee, Senwanna, J. Kumla & N. Suwannar. Family: Thailandicolaceae Kodchasee, Senwanna, J. Kumla & N. Suwannar. 44. Thailandicolales Kodchasee, Senwanna, J. Kumla & N. Suwannar., ord. nov. 45. Thailandicolaceae Kodchasee, Senwanna, J. Kumla & N. Suwannar., fam. nov. 46. Thailandicola Kodchasee, Senwanna, J. Kumla & N. Suwannar., gen. nov. 47. Thailandicola hydei Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 48. Thailandicola limtongiae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Order: Sporidiobolales Doweld Family: Sporidiobolaceae R.T. Moore 49. Rhodosporidiobolus fluvialis (Fell, Kurtzman, Tallman & J.D. Buck) Q.M. Wang, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 181 (2015) 50. Rhodosporidiobolus ruineniae (Holzschu, Tredick & Phaff) Q.M. Wang, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 182 (2015) 51. Rhodotorula paludigena (Fell & Tallman) Q.M. Wang, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 181 (2015) 52. Rhodotorula toruloides (I. Banno) Q.M. Wang, F.Y. Bai, M. Groenew. & Boekhout, Stud. Mycol. 81: 181 (2015) 53. Rhodotorula thailandensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 54. Sporobolomyces thailandensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Subphylum: Ustilaginomycotina Doweld Class: Exobasidiomycetes Begerow, M. Stoll & R. Bauer Order: Exobasidiales Henn. Family: Brachybasidiaceae Gäum. 55. Meira argovae Boekhout, Scorzetti, Gerson & Sztejnb. ex Denchev & T. Denchev, Mycobiota 11: 3 (2021), (first record in Thailand) 56. Meira plantarum Q.M. Wang, Y.Y. Li, M. Groenew. & M.M. Wang, Frontiers Microbiol. 12: 16 (2022), (first record in Thailand) 57. Meira chiangmaiensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 58. Meira limtongiae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 59. Meira pollinicola Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Family: Laurobasidiaceae Pinruan, Sommai, Suetrong, Somrith. & E.B.G. Jones 60. Laurobasidium hachijoense (Y. Otani, Kakish. & Iijima) Kakish., Nagao & Denchev, Phytotaxa 303: 97 (2017) Order: Microstromatales R. Bauer & Oberw. Families Incertae sedis 61. Jaminaea lantanae Q.M. Wang, Y.Y. Li, M. Groenew. & M.M. Wang, Frontiers Microbiol. 12: 22 (2022), (first record in Thailand) 62. Parajaminaea hydei Kodchasee, Senwanna, J. Kumla & N. Suwannarach, sp. nov. 63. Sympodiomycopsis europaea Q.M. Wang, Y.Y. Li, M. Groenew. & F.Y. Bai, Frontiers Microbiol. 12: 22 (2022), (first record in Thailand) 64. Sympodiomycopsis paphiopedili Sugiy., Tokuoka & Komag., in Sugiyama, Tokuoka, Suh, Hirata & Komagata, Antonie van Leeuwenhoek 59: 101 (1991), (first record in Thailand) 65. Sympodiomycopsis hydei Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 66. Sympodiomycopsis limtongiae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 67. Sympodiomycopsis saisamorniae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Class: Ustilaginomycetes Warm. Order: Ustilaginales Bek. Family: Ustilaginaceae Tul. & C. Tul. 68. Anthracocystis heteropogonicola (Mundk. & Thirum.) McTaggart & R.G. Shivas, Persoonia 29: 123 (2012), (first record in Thailand) 69. Moesziomyces antarcticus (Goto, Sugiy. & Iizuka) Q.M. Wang, Begerow, F.Y. Bai & Boekhout, Stud. Mycol. 81: 81 (2015) 70. Moesziomyces bullatus (J. Schröt.) Vánky, Bot. Notiser 130: 133 (1977) 71. Moesziomyces parantarcticus (Sugita, M. Takash., Mekha & Poonwan) Q.M. Wang, Begerow, F.Y. Bai & Boekhout, Stud. Mycol. 81: 81 (2015) 72. Pseudozyma chiangmaiensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 73. Pseudozyma lannaensis Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 74. Pseudozyma limtongiae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 75. Pseudozyma pollinicola Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. 76. Pseudozyma saisamorniae Kodchasee, Senwanna, J. Kumla & N. Suwannar., sp. nov. Table 1.
Content of anthophilous yeasts in this study.
-
Location sites Flower species Chiang Mai Province Doi Saket District, Samran Rat (CMD) Combretum indicum Phrao District, Nam Phrae (CMPH) Boesenbergia rotunda, Cananga odorata, Canna indica, Crinum asiaticum, Globba winitii, Ixora sp., Musa sapientum, Oncidium sp., Phyllanthus pulcher, Syzygium jambos, Zamioculcas zamiifolia, and Zephyranthes minuta Mueang District, Chang Phueak (CMM-CP) Adenium obesum, Angelonia goyazensis, Antigonon leptopus, Argyranthemum frutescens, Bidens pilosa, Capsicum sp., Citrus japonica, Dolichandrone serrulata, Exacum affine, Jasminum sambac, Mecardonia procumbens, Melampodium divaricatum, Millingtonia hortensis, Murraya paniculata, Nerium oleander, Physalis minima, Plumbago auriculata, Plumeria obtusa, Portulaca grandiflora, Rosa sp., and Salvia farinacea Mueang District, Mae Hia (CMM-MH) Allamanda cathartica, Curcuma sessilis, Hamelia patens, Plumeria rubra, Ruellia tuberosa, and Tecoma stans Mueang District, Suthep (CMM-ST) Alstonia scholaris, Bougainvillea hybrid, Carmona retusa, Ixora chinensis, Passiflora caerulea, Senna spectabilis, and Tectona grandis San Kamphaeng District, Ton Pao (CMSP) Lagerstroemia speciosa, Momordica charantia, and Morinda citrifolia Chiang Rai Province Mae Fah Luang District, Mae Fah Luang (CR) Pentas lanceolata and Thunbergia erecta Phayao Province Mueang District (PY) Catharanthus roseus, Cnidoscolus aconitifolius, Dichorisandra thyrsiflora, Hibiscus rosa-sinensis, Jatropha integerrima, Melampodium divaricatum, Nerium oleander, Ocimum tenuiflorum, Pachystachys lutea, Plumeria pudica, and Thryallis glauca Table 2.
Location of sampling sites and flower species used in this study.
-
Species Strain Host Locality GenBank accession number D1/D2 ITS rpb1 tef1-α Ascomycota, Saccharomycotina
Dipodascomycetes, Dipodascales, TrichomonascaceaeStarmerella etchellsii SDBR-CMU641 Passiflora caerulea Thailand, Chiang Mai Province, Mueang District, Suthep PV834436 PV834623 – – Pichiomycetes, Serinales, Debaryomycetaceae Candida tropicalis SDBR-CMU677 Melampodium divaricatum Thailand, Phayao Province, Mueang District PV834442 PV834627 – – Kodamaea ohmeri SDBR-CMU607 Boesenbergia rotunda Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834443 PV834628 – – SDBR-CMU626 Jasminum sambac Thailand, Chiang Mai Province Mueang District, Chang Phueak PV834444 PV834629 – – SDBR-CMU660 Catharanthus roseus Thailand, Phayao Province, Mueang District PV834445 PV834630 – – SDBR-CMU663 Catharanthus roseus Thailand, Phayao Province, Mueang District PV834446 PV834631 – – SDBR-CMU670 Pentas lanceolata Thailand, Phayao Province, Mueang District PV834447 PV834632 – – Kodamaea restingae SDBR-CMU640 Bougainvillea hybrid Thailand, Chiang Mai Province, Mueang District PV834448 PV834633 – – SDBR-CMU672 Crinum asiaticum Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834449 PV834634 – – Meyerozyma caribbica SDBR-CMU644 Pentas lanceolata Thailand, Chiang Rai Province, Mae Fah Luang District, Mae Fah Luang PV834450 PV834635 – – SDBR-CMU676 Melampodium divaricatum Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834451 PV834636 – – Metschnikowiaceae Metschnikowia cibodasensis SDBR-CMU554 Tecoma stans Thailand, Chiang Mai Province, Mueang District, Suthep PV834454 PV834639 – – Metschnikowia koreensis SDBR-CMU553 Plumeria obtusa Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834455 – – SDBR-CMU555 Syzygium jambos Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834456 PV834640 – – SDBR-CMU558 Allamanda cathartica Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834457 – – SDBR-CMU559 Allamanda cathartica Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834458 – – – SDBR-CMU625 Jasminum sambac Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834459 – – – SDBR-CMU643 Pentas lanceolata Thailand, Chiang Rai Province, Mae Fah Luang, Mae Fah Luang PV834460 – – – SDBR-CMU646 Cnidoscolus aconitifolius Thailand, Phayao Province, Mueang District PV834461 – – – SDBR-CMU649 Jatropha integerrima Thailand, Phayao Province, Mueang District PV834462 – – – SDBR-CMU651 Dichorisandra thyrsiflora Thailand, Phayao Province, Mueang District PV834463 – – – SDBR-CMU655 Pentas lanceolata Thailand, Chiang Rai Province, Mae Fah Luang, Mae Fah Luang PV834464 – – – SDBR-CMU657 Nerium oleander Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834465 – – – SDBR-CMU658 Nerium oleander Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834466 – – – SDBR-CMU669 Pentas lanceolata Thailand, Chiang Rai Province, Mae Fah Luang, Mae Fah Luang PV834467 – – – SDBR-CMU671 Bougainvillea hybrid Thailand, Chiang Mai Province, Mueang District, Suthep PV834468 – – – SDBR-CMU673 Ixora chinensis Thailand, Chiang Mai Province, Mueang District, Suthep PV834469 – – – SDBR-CMU674 Ixora chinensis Thailand, Chiang Mai Province, Mueang District, Suthep PV834470 – – – Saccharomycetes, Phaffomycetales, Phaffomycetaceae Cyberlindnera fabianii SDBR-CMU563 Rosa sp. Thailand, Chiang Mai Province, Mueang District, Chang Phueak, PV834471 PV834641 – – Saccharomycodales, Saccharomycodaceae Hanseniaspora lachancei SDBR-CMU560 Antigonon leptopus Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834472 PV834642 – – SDBR-CMU598 Antigonon leptopus Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834473 PV834643 – – Basidiomycota, Agaricomycotina
Tremellomycetes, Filobasidiales, FilobasidiaceaeNaganishia albida SDBR-CMU556 Nerium oleander Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834476 PV834646 – – Naganishia diffluens SDBR-CMU712 Argyranthemum frutescens Thailand, Chiang Mai Province, Mueang District, Chang Phueak, PV834477 PV834647 – – SDBR-CMU716 Argyranthemum frutescens Thailand, Chiang Mai Province, Mueang District, Chang Phueak, PV834478 PV834648 – – Naganishia liquefaciens SDBR-CMU609 Rosa sp. Thailand, Chiang Mai Province, Mueang District, Chang Phueak, PV834479 PV834649 – – Tremellales, Bulleribasidiaceae Hannaella pagnoccae SDBR-CMU576 Curcuma sessilis Thailand, Chiang Mai Province, Mueang District, Mae Hia PV834480 PV834650 – – SDBR-CMU653 Dichorisandra thyrsiflora Thailand, Phayao Province, Mueang District PV834481 PV834651 – – SDBR-CMU683 Plumeria pudica Thailand, Phayao Province, Mueang District PV834482 PV834652 – – SDBR-CMU688 Plumeria pudica Thailand, Phayao Province, Mueang District PV834483 PV834653 – – Hannaella phyllophila SDBR-CMU583 Oncidium sp. Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834484 PV834654 – – Vishniacozyma marinae SDBR-CMU692 Plumeria pudica Thailand, Phayao Province, Mueang District PV834485 PV834655 – – Cryptococcaceae Kwoniella bestiolae SDBR-CMU642 Passiflora caerulea Thailand, Chiang Mai Province, Mueang District, Suthep PV834488 PV834658 – – SDBR-CMU699 Bougainvillea hybrid Thailand, Chiang Mai Province, Mueang District, Suthep PV834489 PV834659 – – Kwoniella heveanensis SDBR-CMU633 Tectona grandis Thailand, Chiang Mai Province, Mueang District, Suthep PV834490 PV834660 – – SDBR-CMU726 Alstonia scholaris Thailand, Chiang Mai Province, Mueang District, Suthep PV834491 PV834661 – – Rhynchogastremaceae Papiliotrema aspenensis SDBR-CMU652 Plumeria pudica Thailand, Phayao Province, Mueang District PV834496 PV834666 – – Papiliotrema flavescens SDBR-CMU668 Pachystachys lutea Thailand, Phayao Province, Mueang District PV834499 PV834669 – –
Trichosporonales, TrichosporonaceaeTrichosporon asahii SDBR-CMU549 Bidens Pilosa Thailand, Chiang Mai Province, Mueang District, Chang Phuea PV834507 PV834677 – – SDBR-CMU656 Nerium oleander Thailand, Phayao Province, Mueang Districtin PV834508 PV834678 – – Pucciniomycotina
Cystobasidiomycetes, Cystobasidiales, CystobasidiaceaeCystobasidium benthicum SDBR-CMU720 Lagerstroemia speciosa Thailand, Chiang Mai Province, San Kamphaeng District, Ton Pao PV834511 PV834681 – – Cystobasidium keelungense SDBR-CMU585 Mecardonia procumbens Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834512 PV834682 – – SDBR-CMU713 Argyranthemum frutescens Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834513 PV834683 – – Cystobasidium minutum SDBR-CMU608 Rosa sp. Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834514 PV834684 – – SDBR-CMU721 Lagerstroemia speciosa Thailand, Chiang Mai Province, San Kamphaeng District, Ton Pao PV834515 PV834685 – – Cystobasidiomycetes, Erythrobasidiales, Erythrobasidiaceae Erythrobasidium primogenitum SDBR-CMU689 Plumeria pudica Thailand, Phayao Province, Mueang District PV834525 PV834695 – –
Microbotryomycetes, Sporidiobolales, SporidiobolaceaeRhodosporidiobolus fluvialis SDBR-CMU562 Rosa sp. Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834532 PV834702 – – SDBR-CMU568 Citrus japonica Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834533 PV834703 – – SDBR-CMU659 Nerium oleander Thailand, Phayao Province, Mueang District PV834534 PV834704 – – Rhodosporidiobolus ruineniae SDBR-CMU593 Zamioculcas zamiifolia Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834535 PV834705 – – SDBR-CMU665 Ocimum tenuiflorum Thailand, Phayao Province, Mueang District PV834536 PV834706 – – SDBR-CMU724 Combretum indicum Thailand, Chiang Mai Province, Doi Saket District, Samran Rat PV834537 PV834707 – – Rhodotorula paludigena SDBR-CMU580 Exacum affine Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834538 PV834708 – – Rhodotorula toruloides SDBR-CMU678 Melampodium divaricatum Thailand, Phayao Province, Mueang District PV834542 PV834712 – –
Ustilaginomycotina
Exobasidiomycetes, Exobasidiales, BrachybasidiaceaeMeira argovae SDBR-CMU709 Morinda citrifolia Thailand, Chiang Mai Province, San Kamphaeng District, Ton Pao PV834546 PV834716 – – SDBR-CMU729 Alstonia scholaris Thailand, Chiang Mai Province Mueang District, Suthep PV834547 PV834717 – – SDBR-CMU730 Alstonia scholaris Thailand, Chiang Mai Province Mueang District, Suthep PV834548 PV834718 – – Meira plantarum SDBR-CMU687 Plumeria pudica Thailand, Phayao Province, Mueang District PV834555 PV834725 – –
Exobasidiales, LaurobasidiaceaeLaurobasidium hachijoense SDBR-CMU615 Syzygium jambos Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834559 PV834729 – – SDBR-CMU616 Syzygium jambos Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834560 PV834730 – – Microstromatales, Microstromatales genera incertae sedis Jaminaea lantanae SDBR-CMU693 Hibiscus rosa-sinensis Thailand, Phayao Province, Mueang District PV834561 PV834731 – – Sympodiomycopsis europaea SDBR-CMU605 Plumeria obtusa Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834564 PV834734 PV941870 PV947475 SDBR-CMU637 Bougainvillea hybrid Thailand, Thailand, Chiang Mai Province, Mueang District, Suthep PV834565 PV834735 – – Sympodiomycopsis paphiopedili SDBR-CMU567 Hamelia patens Thailand, Chiang Mai Province, Mueang District, Mae Hia PV834572 PV834742 – – SDBR-CMU708 Morinda citrifolia Thailand, Chiang Mai Province, San Kamphaeng District, Ton Pao PV834573 PV834743 – – Ustilaginomycetes, Ustilaginales, Ustilaginaceae Anthracocystis heteropogonicola SDBR-CMU551 Bidens pilosa Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834576 PV834746 – – SDBR-CMU589 Antigonon leptopus Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834577 PV834747 – – Moesziomyces antarcticus SDBR-CMU548 Zephyranthes minuta Thailand, Chiang Mai Province, Phrao District, Nam Phrae PV834578 PV834748 – – SDBR-CMU634 Senna spectabilis Thailand, Chiang Mai Province, Mueang District, Suthep PV834579 PV834749 – – SDBR-CMU647 Hibiscus rosa-sinensis Thailand, Phayao Province, Mueang District PV834580 PV834750 – – SDBR-CMU706 Momordica charantia Thailand, Chiang Mai Province, San Kamphaeng District, Ton Pao PV834581 PV834751 – – Moesziomyces bullatus SDBR-CMU571 Citrus japonica Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834582 PV834752 – – SDBR-CMU582 Exacum affine Thailand, Chiang Mai Province, Mueang District, Chang Phueak PV834583 PV834753 – – SDBR-CMU645 Cnidoscolus aconitifolius Thailand, Phayao Province, Mueang District PV834584 PV834754 – – SDBR-CMU675 Melampodium divaricatum Thailand, Phayao Province, Mueang District PV834585 PV834755 – – SDBR-CMU679 Cnidoscolus aconitifolius Thailand, Phayao Province, Mueang District PV834586 PV834756 – – Moesziomyces parantarcticus SDBR-CMU654 Pentas lanceolata Thailand, Phayao Province, Mueang District PV834587 PV834757 – – SDBR-CMU733 Ixora chinensis Thailand, Chiang Mai Province, Mueang District, Suthep PV834588 PV834758 – – SDBR-CMU734 Ixora chinensis Thailand, Chiang Mai Province, Mueang District, Suthep PV834589 PV834759 – – SDBR-CMU732 Ixora chinensis Thailand, Chiang Mai Province, Mueang District, Suthep PV834590 PV834760 – – '−', undetermined sequence. Table 3.
Known yeast species obtained in this study, with their hosts, localities, and GenBank accession numbers.
-
Characteristics 1 2 3 4 Fermentation Glucose + – – – Carbon assimilation D-Ribose + – + v D-Xylose + – w + L-Arabinose w – w v L-Rhamnose – – – + Sucrose + + – + Maltose + + – + α-α-Trehalose + – – + Cellobiose + w – v Salicin – – + + Melibiose – + – v Lactose – – – + Raffinose – – – v Melizitose + – – + Soluble starch – – – + Erythritol + + + – Succinate + – + + Ethanol – + + + Nitrogen assimilation Potassium nitrate – – – nd Growth characteristics Growth at 37 °C – + + + Strains 1: P. siamensis sp. nov, 2: P. melissophilus[54], 3: P. fermenticarens[55], and 4: P. castillae[27]. Table 4.
Phenotypic characteristics differentiating Priceomyces siamensis from closely related Priceomyces species.
-
Characteristics 1 2 3 4 5 Carbon assimilation L-Sorbose – + – + + N-Acetyl glucosamine – + – – – D-Ribose + + – – – D-Xylose + + w w – L-Arabinose + + + w + D-Arabinose + – – – + L-Rhamnose + + w w v Methyl-α-D-glucoside + + w w + Cellobiose + + + w + Salicin + + – w – Melibiose + + + w + Lactose + + + – + Raffinose + + + w + Inulin – + + – + Glycerol w + – – – Ribitol w + – – + D-Glucitol + + – – + Galactitol + + – + + myo-Inositol + + w w + Succinate + + w w + Citrate + + – – + Ethanol w – – w + Nitrogen assimilation Sodium nitrite + + – – + Ethylamine HCl w + + – + Cadaverine – – – + – Growth characteristics Growth at 25 °C + + + + + Growth at 30 °C + – – v + Growth at 35 °C – – – v – Strains 1: F. lannaense sp. nov., 2: F. castaneae[62], 3: F. globosum, 4: F. mali[63], and 5: F. pseudomali[62]. Table 5.
Phenotypic characteristics differentiating Filobasidium lannaense from closely related Filobasidium species.
-
Characteristics 1 2 3 Carbon assimilation D-Ribose w + + D-Arabinose + + – Salicin w – + Glycerol w + – Erythritol – + + D-Glucono-1,5-lactone – + nd D-Glucuronate – w + DL-Lactate + + w Citrate – – w Ethanol – – + Nitrogen assimilation Creatine – – + Growth characteristics Growth at 35 °C + w nd Growth at 37 °C + – + Strains 1: Kw. limtongiae sp. nov., 2: Kw. saisamorniae sp. nov., and 3: Kw. heveanensis[69]. Table 6.
Phenotypic characteristics differentiating Kwoniella limtongiae and Kw. saisamorniae from closely related Kwoniella species.
-
Characteristics 1 2 3 4 Carbon assimilation L-Sorbose + – – – N-Acetyl glucosamine + + + l/w D-Arabinose + w +/w +/w L-Rhamnose + + +/w l/w α-α-Trehalose + + + – Salicin + + – l/w Melibiose + + + +/w Lactose w + + – Inulin – w – +/w Soluble starch – + +/s – Glycerol – – +/w – Erythritol + + – – Ribitol w w + l/w D-Glucitol + + –/+ l/w D-Mannitol + + –/w l/w Galactitol + w –/w + myo-Inositol w + + + DL-Lactate + w +/s – Succinate + + +/s – Citrate + w +/s – Nitrogen assimilation of Ammonium sulfate + – – + Ethylamine HCl + nd – + L-Lysine + – – l/w Growth characteristics Growth at 30 °C + + – + Growth at 35 °C + – nd – Growth at 37 °C + – nd – Growth on 50% Glucose w – – nd Strains 1: Sa. thailandensis sp. nov, 2: Sa. flava[77], 3: Sa. paraflava[78], and 4: Sa. pseudoflava[63]. Table 7.
Phenotypic characteristics differentiating Saitozyma thailandensis from closely related Saitozyma species.
-
Characteristics 1 2 3 4 5 6 Carbon assimilation L-Sorbose + w l/w + – w D-Ribose + + l/w + + – D-Xylose + + + + + – D-Arabinose + + l/w + + – L-Rhamnose + – l/w + v – Maltose + + + + – – Methyl-α-D-glucoside + + l/w + + – Salicin – w w + nd – Melibiose – – – + – + Lactose + + – – + – Raffinose + + l/w + nd – Melizitose + + l/w + + – D-Glucuronate + – + + nd nd D-Galacturonic acid + – + – nd nd DL-Lactate + + l/w + nd – Citrate + – + + nd nd Xylitol + w nd nd nd nd Nitrogen assimilation Potassium nitrate + w – – v + Sodium nitrite + w nd nd nd – Ethylamine HCl + w – – nd – L-Lysine + w – – + + Cadaverine w w – – v – Growth characteristics Growth at 30 °C + – – + v – Growth at 35 °C – – – + – – Strains 1: Sy. chiangmaiensis sp. nov, 2: Sy. hydei sp. nov, 3: Sy. coprosmae, 4: Sy. oryzicola[88], 5: Sy. pseudomarina[87], and 6: Sy. rhododendri[63]. Table 8.
Phenotypic characteristics differentiating Symmetrospora chiangmaiensis and Sy. hydei from closely related Symmetrospora species.
-
Characteristics 1 2 3 4 5 6 7 8 9 Carbon assimilation D-Galactose + + + s + – – – – L-Sorbose + + + + + + + – – D-Ribose – + – w – – – + w D-Xylose – + + s + – + + – L-Arabinose + + – s + + – + + D-Arabinose – + – w s – – + w Sucrose + – – – – + + + + Maltose + – – – – – + + + α-α-Trehalose + – – – – + v – + Methyl-α-D-glucoside – – – – – + + w w Salicin + + + + + + – + + Melibiose – – – – – + – nd – Raffinose – – – + – – – – – Melizitose + – – – nd + + nd + Inulin – – – – nd + + nd + Soluble starch – – – – – + w nd + Glycerol – – – s nd w + – – D-Glucitol + + + + + + – – – Galactitol + + + + + – – – – DL-Lactate – – – s – + + + + Succinate + + + + + + + + + Citrate – + + s – + – – – Ethanol – + + + + + + + + Nitrogen assimilation Potassium nitrate – – + – – + + + + Sodium nitrite – – – – nd + w – w L-Lysine + + + + + – + + + Cadaverine – + – – – + + w + Growth characteristics Growth at 15 °C + nd nd nd nd 17 °C nd nd 17 °C Growth at 25 °C + + + + + nd nd + nd Growth at 30 °C + – – – – – nd – – Growth at 35 °C + – nd – nd nd nd nd nd Growth in 0.01% Cycloheximide – nd + + – nd nd nd nd Strains 1: Cu. chiangmaiense sp. nov, 2: Cu. cygneicollum, 3: Cu. nothofagi, 4: Cu. pallidicollinum[89], 5: Cu. rogersii[90], 6: Baiomyces sejilaensis, 7: Fengyania pteridophytorum, 8: Nakaseozyma junci, and 9: N. lonicerae[81]. Table 9.
Phenotypic characteristics differentiating Curvibasidium chiangmaiense and closely related species.
-
Characteristics/order Thailandicolales Heterogastridiales Kriegeriales Leucosporidiales Leucosporidiales Incertae sedis 1 2 3 4 5 6 7 8 9 10 11 12 13 Carbon assimilation D-Galactose s/w – – + w – + + – – – + – L-Sorbose – – – + w + + s w s – + + N-Acetyl glucosamine + w/+ nd nd nd nd nd nd – nd nd nd nd D-Ribose – – – + + + +/l s – s/w – – + D-Xylose – – – + w + +/l + – + – + + L-Arabinose – – – + w + – + – – – +/– – D-Arabinose – – – v + + +/l – – – – – + L-Rhamnose + – – + w + – – – – – – – Sucrose + + – + + + – + + + + – + Maltose + + – + + + – + + + +/l – + α-α-Trehalose + + – + + + +/l + + + +/l + + Methyl-α-D-glucoside + + nd + + nd – – + + – – + Cellobiose w/+ + v + + + +/l – + + – – + Salicin + v nd + + + – – – + – – + Melibiose – – – – – + – – – – – – – Lactose – – – – – + – – – + – – + Raffinose – + – – – + – – +/l – – – + Melizitose + + – + + + – + + + + – + Inulin – – nd – – + – + + v v – – Soluble starch – – – – – + – – w – – – + Glycerol w – – + + + + + w + + + + Erythritol – – – – nd + – – – – – – – Ribitol – – – + w + +/l + +/l s/w nd – + D-Glucitol – – nd + – + + + – + nd + + D-Mannitol + – + + + + + + + + + + + Galactitol – – nd + – + – – – – nd – – myo-Inositol – – – – – + – – – – nd – – D-Glucono-1,5-lactone + + nd nd + + +/l + nd nd nd +/l + D-Gluconate w – nd + nd + nd + nd – + nd + D-Glucuronate + v nd + nd + nd + – nd nd nd + D-Galacturonic acid s v nd nd nd nd nd nd nd nd nd nd + DL-Lactate s s/w nd + + + – s w – – nd + Succinate + v nd + – + + + + s/w – + + Citrate s – + + – + + + + – – – + Methanol – – nd – – – nd nd – – + – – Ethanol – – nd + w + + + – – nd nd – Xylitol – – nd v w nd nd – nd nd nd nd + Nitrogen assimilation Ammonium sulfate + + nd nd nd nd + nd + + nd nd nd Potassium nitrate – – – + – + – + + + – + + Sodium nitrite – – nd + – + – – + – nd nd + Ethylamine HCl – – nd nd + + + + + + nd nd – L-Lysine + + nd nd w + l + + – nd nd + Cadaverine – – nd nd nd + + + + – nd nd – Growth characteristics Growth at 25 °C + + nd + + + + + + + + + + Growth at 30 °C + + nd – + nd nd + + + + + + Strains 1: Th. hydei sp. nov, 2: Th. Limtongiae sp. nov., 3: Reniforma strues[93], 4: Kriegeria eriophori, 5: Meredithblackwellia eburnean[94], 6: Leucosporidium fellii[95], 7: Leu. Intermedium[96], 8: Leu. Krtinense[97], 9: Bauerozyma artemisiae[81], 10: Microbotryozyma swertiae[63], 11: Yunzhangia auriculariae, 12: Yu. sonckii, and 13: Sampaiozyma vanillic[65]. Table 10.
Phenotypic characteristics differentiating Thailandicola hydei, Th. limtongiae and closely related species in order Heterogastridiales Kriegeriales, Leucosporidiales, and Microbotryales.
-
Characteristics 1 2 3 4 5 Carbon assimilation D-Galactose – s v + w D-Ribose – + v +/s – Sucrose + + + + – Maltose + v + + – α-α-Trehalose + v + + – Methyl-α-D-glucoside – v + + nd Cellobiose – – + + – Salicin – – + + nd Melibiose – – + – – Raffinose + + + + – Melizitose + + + + – Soluble starch – + l + – Glycerol w s – + w Ribitol + – – v nd D-Glucitol + v + – nd D-Mannitol + s + + – D-Gluconate – + + + nd Citrate + – v + – Ethanol – – – + + Nitrogen assimilation Potassium nitrate w + v – + Sodium nitrite w – v – – Ethylamine HCl + + v nd w L-Lysine + + v nd – Growth characteristics Growth at 30 °C + + v + + Growth at 35 °C – – – – + Growth at 37 °C – – – nd w Growth in 0.1% Cycloheximide v – nd – + Strains 1: Sp. thailandensis sp. nov. 2: Sp. patagonicus[99], 3: Sp. salmoneus 4: Sp. shibatanus[88,100], and 5: Sp. sucorum[101]. Table 11.
Phenotypic characteristics differentiating Sporobolomyces thailandensis from closely related Sporobolomyces species.
-
Characteristics 1 2 3 4 5 6 7 8 9 10 Carbon assimilation L-Sorbose – – – – – w/l – – – – D-Arabinose w/– – w + + – +/w + – w L-Rhamnose – – – – – – –/w nd – – Methyl-α-D-glucoside – + – – – – w nd – – Cellobiose + + + + w l + + + + Salicin w/+ + + l l w/l w l l/w w Melibiose + – – +/– + l + + + + Melizitose + + + – – + + + + + Inulin – – – w l l –/w – – + Soluble starch – – – +/l l w/l –/w + – – Glycerol – w – w/l l – –/w – – w Ribitol w + w +/l +/l – +/w – – w D-Glucitol + + + + + + –/w + + w Galactitol – – – +/l l – + – v – myo-Inositol w/– – – – – – –/w – – – D-Glucono-1,5-lactone + + – + l nd + – nd nd D-Gluconate – w – – – nd w – nd nd DL-Lactate – + w w/l l l v w – – Succinate + + w + + l +/w + + w/l Citrate w/– + – + l w/l w – – – Ethanol – – – l l + w + – w Xylitol w/– + – +/d +/d nd +/w + nd nd Nitrogen assimilation Potassium nitrate w/– + + + – l + + + + Sodium nitrite – + + + – l + + + + L-Lysine v w w – + l + nd l/w + Cadaverine – – – + + + + nd – + Growth characteristics Growth at 25 °C + + + + + nd + + + + Growth at 30 °C + + + + + + + nd nd nd Growth at 35 °C – – – – + nd nd nd nd nd Growth at 37 °C – – – – + nd – – nd nd Growth in 0.01% cycloheximide – – – – + nd – nd nd nd Growth in 0.1% cycloheximide – – – – + nd – nd nd nd Growth on 50% glucose + + + – – + + nd – – Strians 1: Mei. chiangmaiensis sp. nov., 2: Mei. limtongiae sp. nov, 3: Mei. pollinicola sp. nov., 4: Mei. argovae, 5: Mei. geulakonigii[105], 6: Mei. marina[81], 7: Mei. miltonrushii[103], 8: Mei. nashicola[106], 9: Mei. plantarum, and 10: Mei. pileae[107]. Table 12.
Phenotypic characteristics differentiating Meira chiangmaiensis sp. nov., Mei. pollinicola sp. nov., and Mei. limtongiae sp. nov. from closely related Meira species.
-
Characteristics 1 2 3 4 5 6 7 Carbon assimilation D-Glucose + + + + + + w D-Galactose w + – + +/s + w L-Sorbose + s/+ + + + + – D-Arabinose – – – – +/s + – α-α-Trehalose + v + w + + + Cellobiose – – – – +/s + – Salicin – – – – w w – Melibiose w s – + v + – Lactose – v – + +/s + – Inulin – – – – w/– – – Soluble starch – – – + w/– – – Erythritol w w/+ w + + + + Ribitol - v – – v + – D-Glucitol w v – +/w + + w D-Mannitol + + w + + + + Galactitol – – – – w – – myo-Inositol – – – w w/– + – D-Gluconate v – – nd s/w nd + Succinate + v – nd w + + Citrate – – – nd w + + Ethanol – – – + s + + Xylitol – w/+ – nd v nd – Nitrogen assimilation Potassium nitrate + + + v w + + Sodium nitrite + + + – v + – Ethylamine HCl – – w – w + – L-Lysine + + w + w – – Cadaverine – – – + nd – – Growth characteristics Growth at 25 °C + + + + + + + Growth at 30 °C + + + – + + + Growth in 0.01% Cycloheximide – – – nd + nd + Growth on 50% Glucose + + + – nd + + Growth on 60% Glucose + + + nd nd nd – Growth on 10% NaCl + 5% glucose w + – nd nd nd + Strian 1: Sym. hydei sp. nov, 2: Sym. limtongiae sp. nov, 3: Sym. saisamorniae sp. nov, 4: Sym. europaea[92], 5: Sym. kandeliae[110], 6: Sym. paphiopedili[111], and 7: Sym. yantaiensis[112]. Table 13.
Phenotypic characteristics differentiating Sym. hydei, Sym. limtongiae, and Sym. saisamorniae from closely related Sympodiomycopsis species.
-
Taxa Ps. chiangmaiensis Ps. lannaensis Ps. limtongiae Ps. pollinicola Ps. saisamorniae Ps. hubeiensis Ps. chiangmaiensis – 1.20% (9 nt) 3.28% (25 nt) 1.07% (8 nt) 1.98% (15 nt) 2.73% (19 nt) Ps. lannaensis 1.20% (9 nt) – 3.27% (25 nt) 1.07% (8 nt) 1.19% (19 nt) 1.87% (13 nt) Ps. limtongiae 3.28% (25 nt) 3.13% (25 nt) – 2.62% (20 nt) 2.48% (19 nt) 2.97% (21 nt) Ps. pollinicola 1.07% (8 nt) 1.07% (8 nt) 2.62% (20 nt) – 1.72% (13 nt) 1.87% (13 nt) Ps. saisamorniae 1.98% (15 nt) 2.51% (19 nt) 2.48% (19 nt) 1.72% (13 nt) – 2.29% (16 nt) Ps. hubeiensis 2.73% (19 nt) 1.87% (13 nt) 3.03% (21 nt) 1.87% (13 nt) 2.29% (16 nt) – Each value indicates that nucleotide divergence (%) between pairs of species, the number of nucleotide substitutions (nt) between the indicated species pairs. Table 14.
Percentage of nucleotide divergence in the ITS region among the species most closely related to Pseudozyma hubeiensis.
-
Characteristics 1 2 3 4 5 6 Carbon assimilation L-Sorbose + + + + + – N-Acetyl glucosamine + + + w + + L-Arabinose + w + + + + D-Arabinose + w + + + + Cellobiose + w + w w + Salicin w w + w – – Soluble starch – – – – – +/w Glycerol + w + + – + Erythritol + w + w – + Ribitol + w + – w + D-Glucitol + + + + w + D-Mannitol + + + + w + myo-Inositol + + + + + – D-Gluconate + – + w – nd D-Galacturonic acid + + w w + nd DL-Lactate w + + w + – Succinate w + + + + + Citrate + + + + + – Nitrogen assimilation Sodium nitrite + + + w + + Ethylamine HCl + + + w + + Growth characteristics Growth at 35 °C – w – – – nd Growth at 37 °C – w – – – + Growth on 50% Glucose + + + + w – Strains 1: Ps. chiangmaiensis sp. nov., 2: Ps. lannaensis, 3: Ps. limtongiae, 4: Ps. pollinicola, 5: Ps. saisamorniae, and 6. Ps. hubeiensis[65]. Table 15.
Phenotypic characteristics differentiating Pseudozyma chiangmaiensis, Ps. lannaensis, Ps. pollinicola, Ps. limtongiae, and Ps. saisamorniae from closely related Pseudozyma species.
Figures
(51)
Tables
(15)