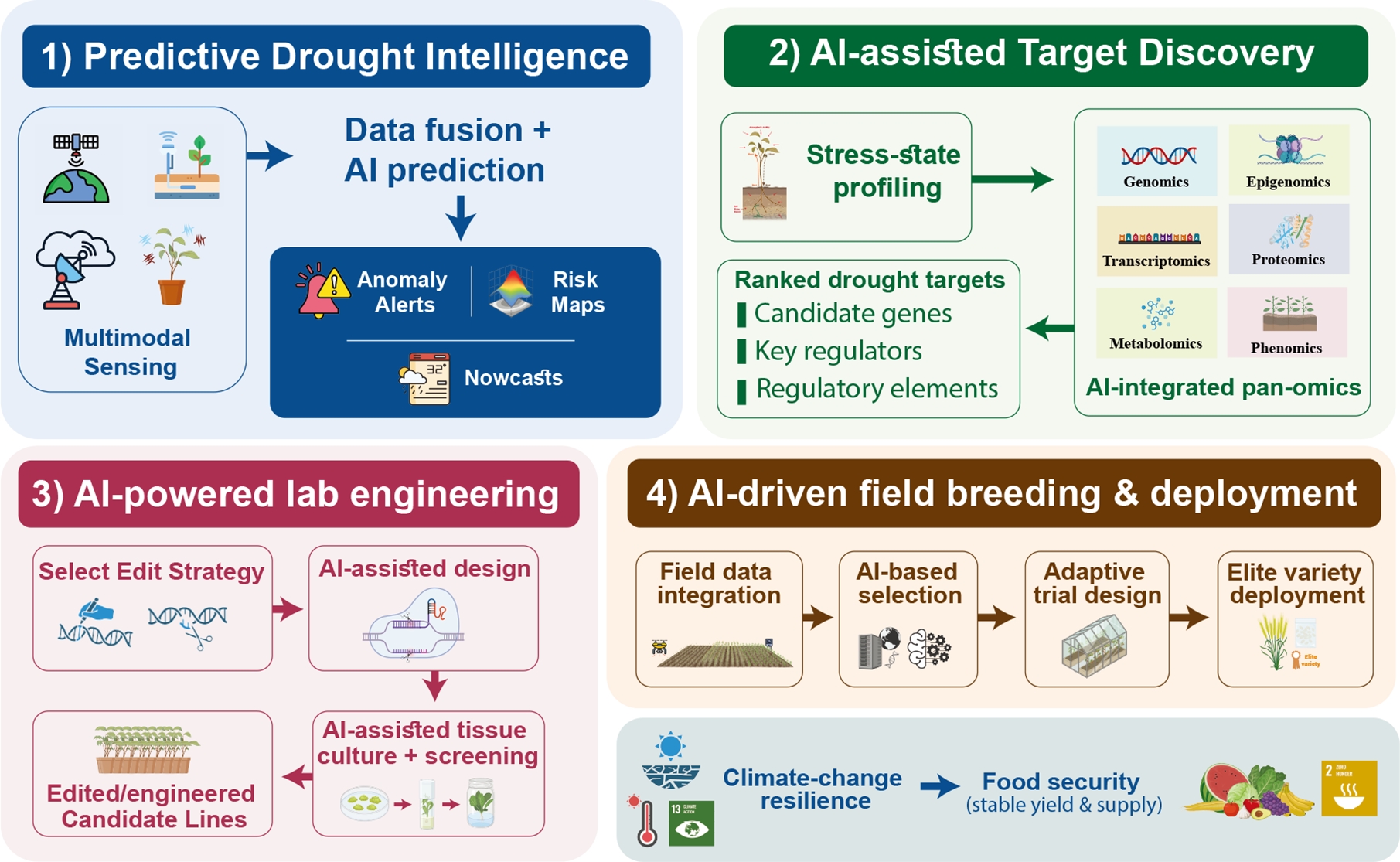
-
The intensifying impacts of climate change are manifesting through more frequent and severe droughts, heatwaves, and erratic rainfall patterns that threaten global food security. These abiotic stresses reduce crop yields and quality, with projections suggesting that climate change could decrease global crop yields by 3%–12% by mid-century and up to 25% by century's end[1]. Drought, already the most critical abiotic stress, is expected to become even more pervasive: over 40% of global land could face year-round droughts under future climate scenarios, and yield failures due to drought may be 4.5 times higher by 2030 and 25 times higher by 2050[1]. These climate-driven stressors impact billions of dollars in agricultural productivity and threaten essential food and crops[2]. Accordingly, there is an urgent need for early-warning systems and AI-enabled breeding pipelines that mitigate losses and accelerate resilient variety development.
Meeting this challenge requires integrating artificial intelligence (AI) across a 'pixels-to-genes-to-fields' pipeline that links early sensing, causal biology, laboratory engineering, and field-scale validation. Modern sensor networks, high-throughput phenotyping, and remote sensing now provide unprecedented volumes of multi-modal field data. AI platforms aggregate soil moisture, canopy health, meteorological streams, and imagery to generate calibrated risk maps and actionable indices for drought management[3]. Convolutional neural networks trained on UAVs and satellite imagery automatically extract subtle drought-responsive features that would be difficult to detect manually, while graph-based models can integrate genomic and transcriptomic information to support QTL refinement and candidate-gene prioritization[4]. Reinforcement-learning algorithms optimize irrigation scheduling based on real-time feedback, reducing water use and maximizing yields, whereas digital-twin simulations replicate field conditions to refine trial allocation and stress application. By integrating multi-source data with advanced AI models, field-scale breeding strategies can accelerate the identification and selection of drought-adaptive lines.
Progress in predictive technologies for early drought detection complements these field analytics. Whereas traditional monitoring relied on sparse ground measurements, modern approaches fuse satellite, UAVs, and ground-based sensors to capture pre-symptomatic indicators of water stress. Hyperspectral and thermal imaging detect changes in chlorophyll content and canopy temperature before visual symptoms appear[4], while bioacoustic sensors measure xylem cavitation events as plants undergo water deficit. AI models combine these diverse data streams—ranging from optical and radar imagery to bioacoustic signals—and employ deep learning and attention mechanisms to detect subtle anomalies indicative of incipient drought[5]. Self-supervised cross-attention frameworks trained on Sentinel-1/2 unify radar and optical inputs for robust regional classification. Quantum-inspired forecasting approaches have been proposed as exploratory methods for representing nonlinear drought dynamics; however, comparative advantage over strong classical baselines and agronomic field validation remain limited, so they should be treated as research frontiers rather than operational tools at present[6]. Together, these advances promise real-time drought-risk forecasts and earlier operational decision points (Fig. 1). Self-supervised multimodal pretraining on large Earth-observation archives has now been demonstrated to improve transfer and few-shot performance across multiple downstream tasks (including agriculture-relevant categories), supporting its use as a robustness strategy when labeled drought datasets are limited[7].
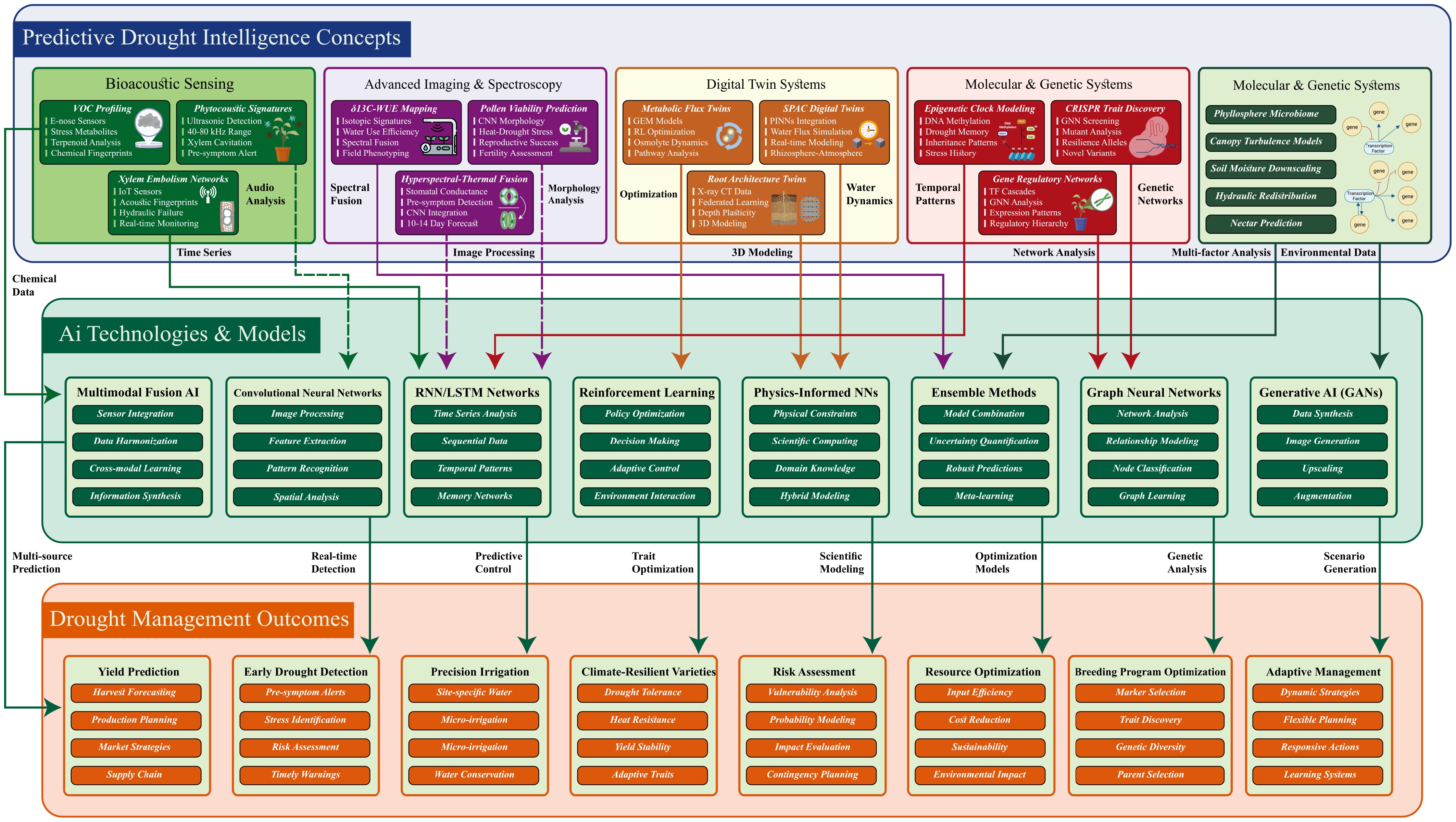
Figure 1.
AI-driven predictive drought-intelligence ecosystem for crops. The schematic organizes the system into three levels: Predictive concepts (top; what is sensed/modeled), AI technologies and models (middle; how information is analyzed), and drought-management outcomes (bottom; decision outputs). Color-coded connectors illustrate data flow between specific inputs, analytical approaches, and actionable results.
At the mechanistic level, plants exhibit a cascade of responses to drought stress, from early perception to systemic signalling and regulatory adaptation. Osmosensors like OSCA channels and mechanosensitive MSL channels perceive changes in water potential and membrane tension, initiating Ca2+ influx and signalling cascades[8,9]. Drought triggers rapid reactive oxygen species (ROS) production, which acts as a secondary messenger during root hydrotropism and stress acclimation (Fig. 2)[10]. Systemic signals include the root-derived peptide CLE25, which travels via xylem to the shoot and stimulates abscisic acid (ABA) biosynthesis, thereby promoting stomatal closure[11−13]. The interaction of ABA with auxin, jasmonic acid (JA), and other hormones finely tunes root growth and stomatal responses[14]. At the chromatin level, plants establish stress memory through dynamic histone modifications (e.g., H3K4me3) and DNA methylation[15,16]; non-coding RNAs recruit chromatin modifiers and modulate gene expression[17]. Proteomic and metabolomic adjustments enhance osmo-protection, with enzymes like Δ1-pyrroline-5-carboxylate synthase driving proline biosynthesis and antioxidants mitigating oxidative damage. The rhizosphere microbiome is also reshaped by root exudates, selectively recruiting beneficial microbes that enhance drought resilience. By pairing these biological insights with AI-driven multi-omics integration, the pipeline can identify regulatory hubs for breeding or editing (Fig. 2).
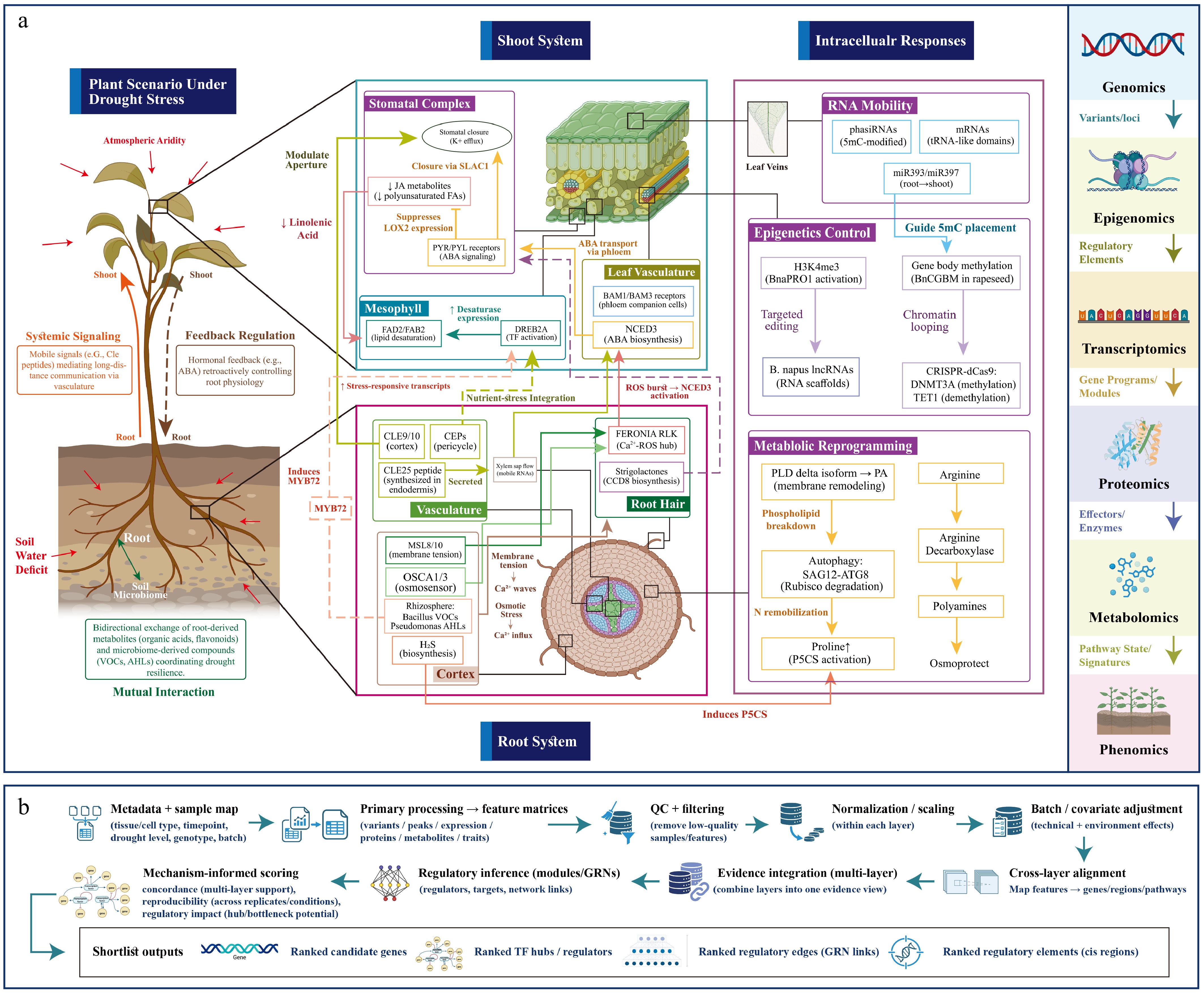
Figure 2.
Integrated molecular and systemic drought responses in crops. (a) Multi-scale drought response mechanisms. The schematic illustrates organ-level signaling and intracellular regulation under drought stress. Roots (left) show early osmosensing (OSCA1/3, MSL8/10), systemic signaling (CLE25, CEP peptides), and rhizosphere interactions (microbial VOCs/AHLs). Shoots (center) detail stomatal regulation (ABA, SLAC1), leaf vasculature signaling (BAM1/BAM3, NCED3), and metabolic reprogramming (lipid desaturation, osmoprotectants). Intracellular layers (right) include RNA mobility, epigenetic control (H3K4me3, targeted editing), and autophagy. The right margin summarizes mechanism-linked omics evidence layers: genomics, epigenomics, transcriptomics, proteomics, metabolomics, and phenomics. (b) AI-guided workflow from omics data to ranked candidates. This panel outlines the analysis pipeline: metadata and sample mapping → primary processing and feature matrices → QC, normalization, and batch correction → cross-layer alignment and evidence integration → regulatory network inference → mechanism-informed candidate scoring (based on multi-layer concordance, reproducibility, and regulatory impact). Outputs are shortlisted ranked candidates: genes, TF hubs, regulatory edges (GRN links), and cis-regulatory elements.
Laboratory breeding and synthetic biology provide precision control over these targets. Convolutional neural networks automate the segmentation and classification of callus, shoot, and root tissues, even under challenging imaging conditions; instance segmentation models count individual structures such as cotyledons with high precision[18]. Generative adversarial networks augment limited training datasets by creating realistic synthetic images, thereby improving model performance and reducing annotation requirements[19]. AI-guided CRISPR/Cas systems leverage deep learning and gradient boosting to predict gRNA specificity and off-target risks; rule-set algorithms combining sequence features and epigenetic context refine gRNA selection (Fig. 3)[20]. Generative models have even designed novel Cas variants and guide sequences with improved activity[20], while deep neural networks like Protein Message Passing Neural Network (ProteinMPNN) accelerate the design of protein sequences for both editing enzymes and stress-response proteins[21]. Synthetic biology frameworks use machine learning to optimize codon usage and design stress-responsive promoters; transformer-based models and CNN/SVM classifiers predict promoter strength and activity (Fig. 3)[22]. These lab innovations generate edited/engineered lines with traceable mechanisms and predicted performance, ready for field-scale validation.
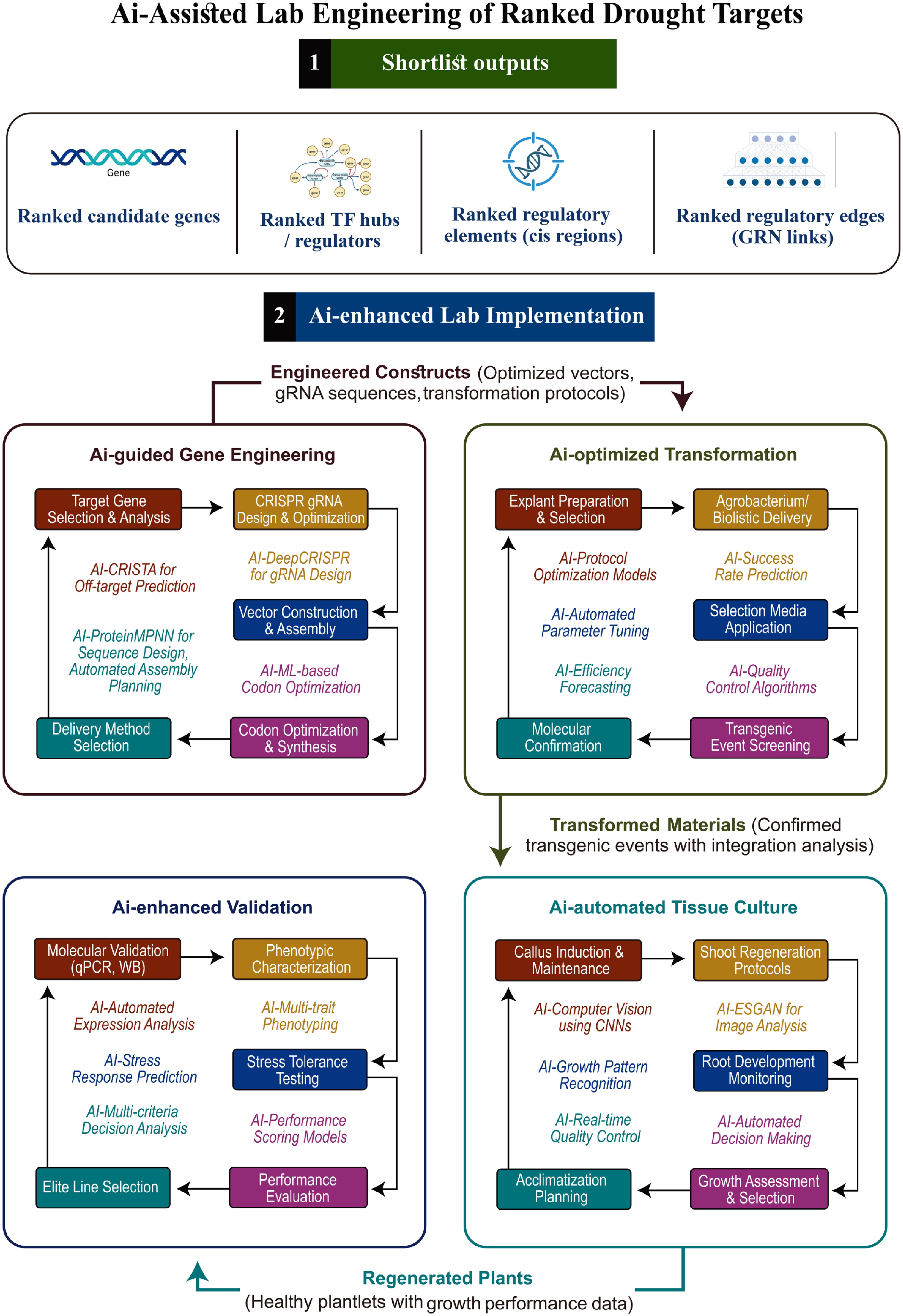
Figure 3.
AI-assisted lab engineering of ranked drought targets. Ranked candidate genes, regulators, and regulatory elements are engineered into constructs. AI-guided gene engineering optimizes CRISPR gRNA design (A-CRISTA, DeepCRISPR), vector assembly (ProteinMPNN), codon optimization, and delivery methods. AI-optimized transformation uses predictive models to guide explant preparation, delivery (Agrobacterium/biolistic), selection, and molecular confirmation. Confirmed transgenic events proceed through AI-automated tissue culture (callus induction, regeneration, rooting) and AI-enhanced validation (molecular, phenotypic, and stress-tolerance testing). The pipeline culminates in regenerated plants with validated growth performance.
Finally, AI-enabled field-scale breeding closes the loop. Multisource data integration, deep learning, GNNs, reinforcement learning, and digital twins improve trial design, rank genotypes across heterogeneous environments, and guide resource-efficient management[23,24]. Continuous-learning platforms retrain on new seasons to refine predictions and recommendations, accelerating advancement of resilient varieties (Fig. 4).
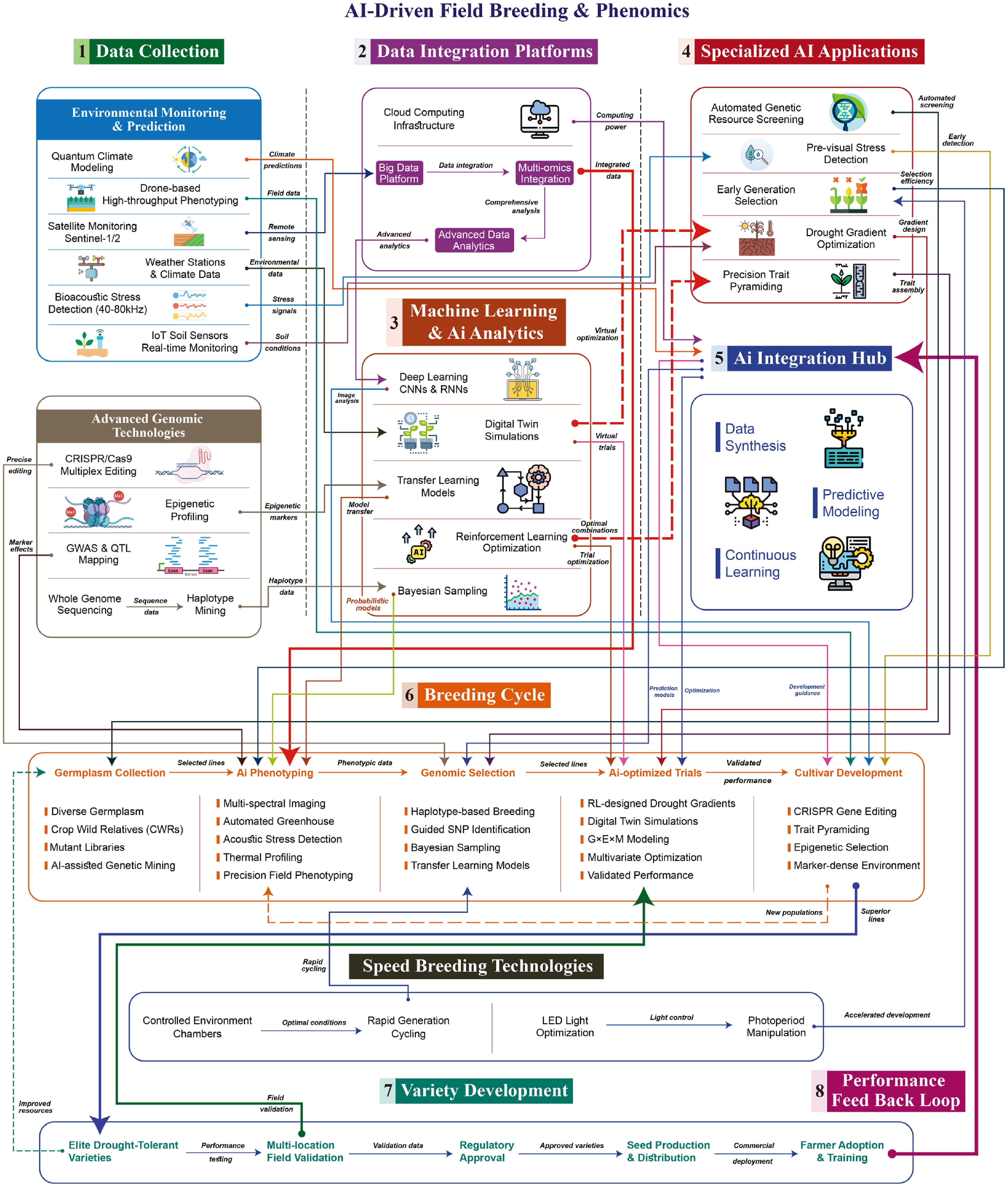
Figure 4.
AI-driven field breeding and phenomics pipeline for drought-resilient crops. The closed-loop workflow integrates field data and genomics to accelerate cultivar development. (1) Data collection: Environmental monitoring (satellites/UAVs, IoT sensors, plant bioacoustics) and genomic inputs (sequencing, GWAS, CRISPR). (2) Data integration: Cloud-based platforms harmonize multi-source data. (3) AI analytics: Deep learning, transfer learning, Bayesian sampling, reinforcement learning, and digital-twin simulations. (4) AI applications: Automated genetic screening, pre-visual stress detection, early-generation selection, drought-gradient optimization, and precision trait pyramiding. (5) AI hub: Synthesizes data, runs predictive models, and enables continuous learning. (6) Breeding cycle: Germplasm collection → AI phenotyping → genomic selection → AI-optimized trials → cultivar development (CRISPR, trait pyramiding). (7) Variety development: Speed breeding accelerates generation turnover; elite lines undergo multi-location validation, regulatory approval, and seed distribution. (8) Feedback loop: Field performance data retrain models and refine selection strategies, closing the iterative cycle.
This review synthesizes a connected pipeline comprising four integrated pillars: (i) predictive technologies for early drought detection and risk mapping (Fig. 1); (ii) mechanistic decoding, from stress perception and systemic signaling to downstream regulatory adaptation (Fig. 2); (iii) AI-enabled laboratory engineering, spanning automated tissue culture, genome editing, and synthetic expression design (Fig. 3); and (iv) AI-driven field-scale breeding, integrating multi-modal data, trial optimization, and predictive modelling (Fig. 4). Together, these components create a closed-loop workflow that links early field signals to causal targets, translates those targets into engineered lines, and evaluates performance across variable environments via continuous-learning platforms. The emphasis throughout is on actionable methods, interoperable workflows, and key decision points designed to shorten breeding cycles and increase the probability that drought-resilience observed in controlled settings translates into durable, field-ready gains under accelerating climate volatility.
-
Early drought monitoring has become critically important for sustainable agriculture and natural resource management, especially in the face of increasingly erratic climate patterns. The traditional models for drought detection often relied on sparse ground-based measurements and empirical thresholds, which were limited in spatial resolution and real-time responsiveness. The convergence of advanced sensors, AI, and novel computational paradigms provides an unprecedented opportunity to identify subtle indicators of water stress well before visible symptoms—i.e., via pre-symptomatic risk sensing[25]. Recent advances have dramatically transformed this field by incorporating robust multimodal data streams and advanced machine learning algorithms. In particular, integrating multimodal data sources ranging from satellite imagery and UAV imaging to emerging plant bioacoustics enables a comprehensive assessment of both large-scale and localized stress patterns. AI models fuse these streams to surface patterns and anomalies that single modalities miss[5,6]. Furthermore, quantum-inspired approaches have been proposed for drought forecasting, but comparative operational advantage over classical models remains unproven[26].
From concept to coverage, satellite missions provide the regional backbone while UAVs resolve the plot-level physiology. Satellite-based remote sensing plays a crucial role in early drought detection by providing large-scale monitoring of soil moisture, vegetation health, and land surface conditions. Missions like Sentinel-1 and Sentinel-2 offer moderate to high spatial and temporal resolution data, including Synthetic Aperture Radar (SAR) imagery that can penetrate clouds and yield consistent soil moisture information under adverse weather conditions. This enables the detection of early drought stress signals over extensive regions, supporting timely regional drought assessments[5]. UAVs equipped with hyperspectral and thermal sensors enable high-resolution, plot-level crop monitoring. Hyperspectral imaging captures detailed spectral reflectance sensitive to biochemical traits such as water content and chlorophyll, while thermal imaging detects canopy temperature differences that indicate water stress before visible symptoms arise. These capabilities facilitate precision phenotyping and localized management. When coupled with deep learning, these datasets yield more accurate field-scale predictions by automatically extracting discriminative features (Fig. 1)[4,27].
At the plant scale, new signals and model classes extend sensitivity. Plant bioacoustics, an emerging sensing modality, detects ultrasonic emissions generated by xylem cavitation inside plants under water stress. In controlled experiments, ultrasonic emissions from drought-stressed plants were shown to be airborne and classifiable, with machine-learning models distinguishing dehydration level and injury states in greenhouse settings, supporting feasibility while also indicating that large-scale field validation remains the next critical step[28]. Although still experimental, combining bioacoustics with spectral and thermal measurements and analyzing them using AI shows promise for non-invasive, cellular-level, real-time drought stress detection, complementing existing remote sensing approaches[29]. Advanced AI algorithms—especially convolutional neural networks (CNNs) and attention-based transformer architectures—are increasingly applied to hyperspectral and thermal datasets for pixel-level classification and anomaly detection. These models capture complex nonlinear relationships between spectral features and physiological states while reducing manual feature engineering[30,31]. Related work in tropical plant imaging likewise shows that deep models can automate feature extraction effectively, although performance may still be constrained by multi-scale targets and leaf overlap in complex scenes[32]. Quantum-inspired forecasting frameworks leverage superposition/entanglement metaphors to handle soil–plant–atmosphere nonlinearity, enabling hybrid AI–quantum models with improved short-term nowcasting and uncertainty representation[26].
Heterogeneous streams become decisions via data fusion plus time-series modelling. To transform diverse environmental sensor streams into actionable early warning signals, data fusion techniques integrate continuous inputs from soil moisture probes, meteorological stations, satellite and UAV-based imagery, and novel bioacoustic sensors. Using advanced assimilation algorithms like Bayesian inference and Kalman filtering helps reconcile differences in resolution and temporal frequency, smoothing gaps, and ensuring stable drought-risk indices. Graph-based machine learning adds spatial coherence by modeling interactions among monitoring sites[33]. These composite indices, such as vegetation stress scores and soil moisture deficits, enable real-time adaptive management of water resources and irrigation, empowering stakeholders to mitigate drought impacts[5]. Modern models can learn time patterns in drought risk. LSTM and Temporal Fusion Transformers (TFTs) are two common choices. They handle delayed effects and changing conditions over time. In soil-moisture prediction, LSTM models have reported strong accuracy (R ≈ 0.95; R2 ≈ 0.85)[34]. Transformer-based models have also shown better performance than LSTMs in some hydrological sequence tasks[35]. LSTM autoencoders provide early anomaly flags in soil-moisture series, and hybrid deep learning–dynamic models extend root-zone soil-moisture forecasts to ~4 weeks, outperforming state-of-the-art dynamic models for flash drought forecasting[36]. These methods capture both seasonal trends and abrupt deviations, providing valuable inputs for short- and long-term drought forecasting critical for proactive resource management (Fig. 1)[34].
When labels are scarce, representation learning and graphs close the gap. Self-supervised and unsupervised learning approaches help address data scarcity. The high cost of labelling environmental datasets has spurred the adoption of self-supervised and unsupervised learning methods. These approaches enable models to learn from the intrinsic structure in unlabelled data, effectively extracting complex patterns that correlate with early stress indicators (Fig. 1)[37]. For example, self-supervised pre-training on large satellite repositories enhances sensitivity to subtle spectral/thermal changes associated with drought stress[6]. This paradigm shift improves predictive accuracy and reduces reliance on labeled drought events, improving transfer across regions and seasons. Graph neural networks (GNNs) fuse heterogeneous sensors by encoding spatial, temporal, and functional relations as edges and efficiently modeling complex interactions underlying drought phenomena[33]. Such integration enables the capture of spatial correlations and propagation effects that are critical for accurate risk prediction. Because edge relations can evolve, GNNs naturally represent dynamic systems, ensuring that fused data reflect current conditions and yield actionable indices for risk alerts and irrigation optimization (Fig. 1)[33].
The outputs of this section are uncertainty-aware drought nowcasts, risk maps, and anomaly alerts derived from multimodal sensing. These outputs define where, when, and which genotypes should be sampled for downstream mechanistic profiling, prioritizing informative timepoints/tissues and enabling targeted pan-omics study design rather than untargeted screening. In the next section, we describe how drought perception/signaling modules provide the mechanistic scaffold for interpreting these stress-linked signatures and converting them into ranked candidate regulators and loci.
-
Drought stress severely constrains plant growth and productivity. It triggers interconnected physiological and molecular responses that promote survival and adaptation. Understanding these mechanisms requires an integrative approach that spans from initial perception at the cellular level to systemic regulation involving multi-layered signaling networks and regulatory feedback. In the context of the early-warning pipelines described above, these mechanisms explain why genotypes diverge once stress is detected and which nodes are most promising for intervention. Advances in technology, particularly in AI and multi-omics analyses, are now enabling unprecedented insights into these processes and translating mechanism-linked signatures into ranked candidate genes and regulatory elements for downstream validation and crop improvement (Fig. 2)[25,38].
Early drought perception and signaling
-
Root cells perceive the initial impact of drought via an intricate set of molecular sensors that include osmosensors and mechanosensitive channels. For example, hyperosmolarity-sensing proteins of the OSCA family act as Ca2+-permeable channels; OSCA1 was the first plant osmotic-stress sensor identified, and OSCA1 expression is upregulated in response to salt and drought, triggering Ca2+ influx and ABA-mediated signalling (Fig. 2)[8]. Mechanosensitive channels such as MSL8 and MSL10 (encoded by MSL8 and MSL10) serve as 'osmotic safety valves,' opening upon membrane tension to prevent cell bursting and thereby modulating turgor under sudden water loss[9]. Concurrently, drought-induced osmotic imbalance triggers rapid reactive oxygen species (ROS) production; NADPH oxidases generate superoxide radicals that are converted to hydrogen peroxide, which acts as a secondary messenger modulating hydrotropic root bending and stress responses[10]. Together, these sensors convert water-deficit cues into fast ionic and redox signatures that form the first layer of drought perception. Because these early ionic and redox signatures are initiated by discrete sensor and signalling gene families, they also define constrained candidate sets that can be systematically prioritized using AI-enabled regulatory and network inference frameworks described later in this section[39].
This constraint is useful computationally: it allows downstream AI pipelines to operate on mechanism-informed candidate spaces rather than genome-wide searches. In practice, drought experiments that pair time-resolved molecular readouts (e.g., transcript abundance and chromatin accessibility) with physiological measurements can be integrated into regulatory models to rank the most plausible causal regulators and cis-elements for targeted validation[39,40].
Drought signals perceived in the roots are rapidly transmitted to shoots through systemic signaling pathways that integrate peptide signals and phytohormones. The root-derived peptide CLE25 is induced by water deficit and transported via xylem to shoots, where it stimulates ABA biosynthesis and promotes stomatal closure (Fig. 2)[11,13]. Classical hormones interact in a spatiotemporally regulated manner: ABA accumulates to induce stomatal closure, auxins promote lateral root development, cytokinins delay senescence, and jasmonic acid (JA) influences root architecture and stress signalling[14]. JA biosynthesis genes such as LOX2 are upregulated under stress; mutants defective in LOX2 exhibit heightened stress sensitivity, whereas exogenous methyl jasmonate enhances stress tolerance[41]. Advanced AI frameworks can assimilate multi-sensor time-series data (thermal, fluorescence, and spectral) with RNNs to reconstruct the likely sequence and intensity of these hormone and peptide signalling events under fluctuating drought.
In the shoot, drought triggers guard-cell responses and leaf adjustments that minimize water loss. ABA binds to PYR/PYL/RCAR receptors, which in turn inhibit PP2C phosphatases. This releases SNF1-related protein kinase (SnRK2) activity from inhibition, allowing it to phosphorylate downstream targets such as the SLAC1 anion channel, ultimately inducing stomatal closure[42]. JA and lipoxygenase LOX2 act synergistically to modulate stomatal behaviour[41]. AI-assisted thermal mapping with CNNs quantifies canopy temperature changes to infer stomatal conductance[4], while digital phenomics platforms integrate RGB, infrared, and hyperspectral imaging to deliver high-resolution assessments of leaf and stomatal responses. At the transcriptional level, drought induces extensive gene-network reprogramming; transcription factors such as DREB, NAC, bZIP, MYB, and AP2/ERF orchestrate expression of osmoprotective and antioxidant genes. Graph neural network (GNN) models infer gene regulatory networks by encoding gene interactions as edges and using attention mechanisms to capture directionality; cross-attention GNNs have outperformed traditional correlation and Bayesian methods in predicting regulatory relationships[43]. These models help prioritise master regulatory nodes that can be targeted by genome editing or breeding, linking perception/signalling modules to downstream engineering strategies. Complementary to network inference, deep learning models that decode plant cis-regulatory sequences can nominate promoter/UTR variants and regulatory elements likely to modulate drought-induced transcriptional reprogramming, enabling target selection beyond coding regions[38].
Regulatory adaptation and resilience
-
The drought-stress response in crops is regulated at multiple molecular levels—epigenetic, proteomic/metabolomic and microbiome interactions—coordinated through feedback loops. Epigenetic memory is established through histone modifications and DNA methylation. Short-term drought 'priming' involves accumulation of H3K4me3 at stress-responsive loci, demethylation of H3K4me3 by H3K4 demethylases such as SsJMJ11 attenuates drought tolerance[15]. Long-term memory is mediated by DNA methylation; CHH methylation levels are particularly sensitive to drought, and drought-tolerant plants maintain more stable methylation patterns, enabling primed responses upon re-exposure[16]. Long non-coding RNAs (lncRNAs) regulate chromatin states by recruiting Polycomb Repressive Complex 2 (PRC2) to deposit H3K27me3 or by sequestering miRNAs; some drought-responsive lncRNAs exhibit memory-related expression and produce miRNAs that modulate hormone signalling[17]. AI models employing convolutional neural networks and autoencoders integrate single-cell chromatin accessibility and methylome data to identify cell-type-specific regulatory elements and candidate epigenetic markers for epigenetic breeding. These layers together define a tunable 'memory module' that conditions how plants react to repeated drought cycles.
At the proteome/metabolome level, drought stress prompts dynamic shifts that support survival. Plants upregulate antioxidant enzymes (superoxide dismutase, catalase, peroxidases) and induce Late Embryogenesis Abundant (LEA) proteins and dehydrins, which act as molecular chaperones stabilizing proteins and membranes[44]. Drought-tolerant cultivars show elevated levels of late embryogenesis abundant (LEA) proteins, dehydrins, heat shock proteins (HSPs), and enzymes linked to antioxidant defence and osmotic adjustment[44]. Proteomic studies reveal increased abundance of heat shock proteins, ROS scavenging proteins, and enzymes involved in proline biosynthesis[44]. The rate-limiting enzyme Δ1-pyrroline-5-carboxylate synthase (P5CS) mediates proline biosynthesis. During drought, P5CS expression increases dramatically, and silencing P5CS reduces proline accumulation and drought tolerance, whereas overexpression enhances proline levels, reduces ROS, and improves drought resistance[45]. Metabolite accumulation (e.g., proline, glycine-betaine, soluble sugars) contributes to osmotic balance and ROS scavenging[45]. Machine-learning algorithms (Random Forests, Support Vector Machines, and CNNs) have successfully integrated proteomic and metabolomic datasets to identify drought-tolerance biomarkers and predict trait performance[23]. This buffering system at protein and metabolite levels translates upstream signals into concrete physiological resilience traits.
Plant roots shape the rhizosphere microbiome through the secretion of exudates that selectively recruit beneficial microbial taxa. Drought conditions enhance the secretion of organic acids like citric and malic acid, attracting halotolerant bacteria, fungi, and Actinobacteria; drought stress also enriches Bacillus and Pseudomonas species that help retain soil moisture and promote plant survival. Stress-resistant groups such as Actinobacteria preferentially utilize phenolic compounds in exudates[46]. Graph-based AI methods and interpretable machine learning identify marker taxa within soil microbiomes; a Random Forest classifier trained on 16S rRNA profiles achieved 92.3% accuracy in distinguishing drought-stressed soils and highlighted specific genera associated with drought resilience[47]. These approaches inform the strategic deployment of microbial consortia to enhance plant drought adaptation. In this way, plant genetics and soil microbial community structure are treated as a coupled system rather than independent levers.
Finally, whole-plant homeostasis under drought is orchestrated through feedback loops involving ABA signalling, root–shoot communication, and chromatin modulators. Predictive modeling frameworks that integrate multi-season phenomics (digital phenotyping platforms, UAVs, and ground-based sensors) with multi-omics data use ensemble and deep learning models to simulate feedback-driven responses[23]. Multi-modal models combining genomics, metabolomics, and spectral imagery achieve high predictive accuracy (R2 ≈ 0.78 – 0.88) for yield and stress tolerance[23]. These simulations identify genotypes with optimal stress memory and resilience, enabling breeders to select cultivars tailored to specific climatic niches[23]. Thus, perception, signalling, epigenetic memory, proteome/metabolome buffering, and microbiome interactions form an integrated network that not only explains observed drought responses but also yields concrete targets for epigenetic breeding and downstream AI-guided genome engineering and field-scale selection. AI-guided target prioritization approaches are outlined below, including deep learning of plant cis-regulatory code[38], integrative regulatory-network inference in plants[39,48], and neural models leveraging multiome data and transfer learning[40,49]. This provides a direct mechanistic bridge from drought-response biology to the downstream laboratory engineering strategies presented in the next section.
AI-guided prioritization of candidate genes and regulatory elements
-
The mechanistic modules described above ultimately converge on transcriptional reprogramming, yet the regulatory logic is frequently encoded in non-coding sequences that shape when, where, and how strongly a drought-responsive gene is expressed. Sequence-to-expression deep learning models can now learn this cis-regulatory code directly from plant gene flanking regions and predict gene expression profiles at scale. In Arabidopsis thaliana, Solanum lycopersicum, Sorghum bicolor, and Zea mays, models trained on flanking sequences achieved > 80% accuracy[38] and enabled predictive feature selection, highlighting major contributions of UTR regions to gene expression levels. The same framework demonstrated cross-species predictive performance and was applied across fourteen tomato genomes to link genetic variation with gene expression differences, including genotype-specific expression of key functional gene groups involving the drought-resistant wild relative Solanum pennellii[38]. For drought stress biology, these models provide an in silico route to rank promoter/UTR variants and candidate regulatory motifs for key response genes (e.g., ABA signaling components, aquaporins, ROS-scavenging enzymes, and TF hubs) prior to wet-lab validation. More broadly, sequence-based machine-learning classifiers have also been used to nominate agronomically relevant gene classes at the proteome scale, as shown by flowering-time gene prediction across 81 plant species[50].
Mechanism-to-target translation also benefits from AI/ML approaches that integrate heterogeneous regulatory evidence into unified gene regulatory networks (GRNs) and then prioritize upstream drivers. In Arabidopsis thaliana, a supervised-learning strategy combined complementary regulatory input networks (DNA motifs, open chromatin, TF binding, and expression-based interactions) to construct an integrated GRN spanning 1,491 TFs, 31,393 target genes, and ~1.7 million interactions[39]. For TFs predicted to regulate ROS stress, 75% were confirmed to function in ROS and/or physiological stress responses, including 13 previously unlinked regulators that were experimentally validated using ROS-specific phenotypic assays of gain- or loss-of-function lines[39]. Because drought responses include rapid ROS bursts coupled to ABA and Ca2+ signaling, integrative GRN frameworks provide a practical route to shortlist master regulators and network bottlenecks while generating testable TF–target hypotheses for follow-up via chromatin profiling, reporter assays, and mutant analysis[39].
Crop-scale target nomination requires regulatory networks that accommodate large genomes and trait mapping resources. In wheat, an integrative regulatory network (wGRN) was built by combining gene expression, sequence motif information, TF binding, chromatin accessibility, and evolutionarily conserved regulation. The resulting network contained ~7.2 million interactions spanning 5,947 TFs and 127,439 target genes and was systematically verified using known regulatory relationships, functional information, and experiments[48]. The wGRN was used to assign genes to 3,891 biological pathways and to prioritize candidate genes associated with complex traits in GWAS, and it also supported the interpretation of a spike temporal transcriptome dataset to identify regulators that improved phenotypic trait prediction using machine learning[48]. For drought stress biology, the same logic can be operationalized by intersecting drought-associated loci (e.g., yield stability, root architecture, canopy temperature, or osmotic adjustment QTL/GWAS signals) with drought-responsive modules and upstream regulators predicted by crop GRNs, thereby reducing broad genomic intervals to a ranked shortlist of regulators and targets for downstream genome editing or marker-assisted introgression[48].
Finally, drought responses are cell-type specific, and AI methods that leverage paired chromatin accessibility and gene expression at single-cell resolution can recover regulatory programs that are otherwise obscured in bulk tissue. LINGER infers GRNs from single-cell multiome data by integrating external bulk datasets and TF motif priors, and it reported a four- to sevenfold relative increase in accuracy compared with existing approaches; it can also estimate TF activity using only gene expression once a reference multiome-derived GRN is learned, enabling identification of driver regulators in case–control designs[40]. In parallel, transformer foundation models such as scGPT have been pretrained on > 33 million cells and can be adapted via transfer learning for downstream tasks including multi-omic integration, perturbation response prediction, and gene network inference[49]. Together, these frameworks support a scalable route for drought studies, resolve relevant cell populations (e.g., guard cells and drought-responsive root tissues), infer cell-resolved GRNs from multiome references, and prioritize influential TFs and regulatory elements controlling drought-response trajectories for targeted validation and engineering[40,49].
Overall, mechanism-informed pan-omics integration and regulatory inference convert drought-response biology into concrete engineering inputs: ranked candidate genes, prioritized TF hubs/regulators, high-confidence GRN edges, and candidate cis-regulatory regions. These shortlists define plausible intervention points and become direct inputs to the AI-assisted genome editing and expression-engineering workflows described next.
-
The laboratory stage consumes the ranked outputs generated through AI-guided prioritization of candidate genes and regulatory elements, including candidate genes, TF hubs/regulators, GRN edges, and cis-regulatory regions, and converts them into executable designs. In practice, this means (i) selecting edit strategies (knockout, allele replacement, multiplex edits, or regulatory tuning), (ii) generating gRNA sets with predicted activity/specificity and designing regulatory parts (promoters/UTRs or synthetic elements) when non-coding control is the goal, and (iii) regenerating and screening edited/engineered plants through automated tissue-culture imaging pipelines.
AI increasingly supports multiple laboratory steps such as automated tissue culture, precision phenotyping, genome editing, and synthetic expression engineering, compressing the design–build–test cycle. To maintain narrative flow with the pipeline, this section focuses on how laboratory AI converts mechanism-derived targets into edited/engineered candidate lines that can be evaluated under field variability.
Automated tissue culture
-
Computer vision has become indispensable for automating in vitro developmental staging in crop tissue culture. Convolutional neural networks (CNNs) learn hierarchical features from raw image data, enabling them to distinguish morphological traits—such as callus texture, shoot emergence, and root formation—even under conditions of high humidity and non-uniform backgrounds. Recent studies demonstrate that CNNs can automatically extract complex spatial and structural information and provide state-of-the-art performance in classification, object detection and segmentation[18]. Instance segmentation models delineate individual structures (e.g., cotyledons) and count them accurately, outperforming threshold-based methods and remaining robust to noise and reflections[18]. When combined with Red–Green–Blue (RGB), hyperspectral or depth imaging, CNNs generate quantitative phenotypic metrics that inform culture optimisation and growth kinetics.
Generative adversarial networks (GANs) complement CNNs by synthesising realistic tissue images. A CycleGAN-based framework generates synthetic phase-contrast images containing morphological details that mimic real tissues, thereby augmenting training datasets and improving segmentation accuracy when annotated images are scarce[19]. This synthetic augmentation enables CNN classifiers to maintain high accuracy despite limited manual annotation, making high-throughput phenotypic screening more scalable. Integrated imaging platforms that stream RGB/depth into CNN classifiers provide continuous, non-invasive monitoring and feedback for subculture timing, reducing operator bias. Overall, the integration of CNNs and GANs streamlines high-throughput tissue culture by reducing annotation requirements, improving detection accuracy, and accelerating regeneration cycles (Table 1)[18,19]. This produces more, cleaner regenerants for downstream editing and testing, tightening the loop to the field.
Table 1. AI-driven field phenotyping and lab-to-field genomic approaches in crops.
No. Task Algorithm Data/sensor Crop/system Key outcome (reported) Scale Ref. 1 Yield mapping Random Forest; functional linear regression Sentinel-2 time series Canola (B. napus) Yield error 12%–16% Producer field datasets [51] 2 Early crop mapping (seasonal windows) InceptionTime (DL); Random Forest Sentinel-1 SAR time series Rapeseed F1 ≈ 95% (same year full season); early season F1 77%–89% Dept.-scale regions (FR) [52] 3 Flowering phenotyping → yield proxy Thresholded NDYI; linear models UAV multispectral Canola Flower count R2 up to 0.95; yield R2 up to 0.42 Five site years [53] 4 Yield prediction in breeding pipeline XGBoost; Random Forest UAV remote sensing + breeding data Peanut Best models R2 up to 0.93 Breeding program [54] 5 Root zone soil moisture estimation RF, SVM, BPNN, ELM Plot hyperspectral (incl. red-edge) Rapeseed RF model: R2 0.944 (0–20 cm), RMSE 0.005 Field plots [55] 6 Genomic prediction (trait prediction) XGBoost; Random Forest vs DL Genome-wide markers Soybean ML > DL in 13/14 prediction tasks Multi trait dataset (n ≈ 1,110) [56] 7 Web deployable DL for GP Deep neural nets (SoyDNGP) Genotypes + historical phenotypes Soybean End-to-end DL framework (GP service) Germplasm scale [57] 8 Compact GP under G × E RF (50 SNP subset) vs full models Multi env. breeding data Soybean Small RF ≈ full models' ability Multi env. scenarios [58] 9 Drought gene discovery Multi locus GWAS + RNA seq Field trials + omics Sunflower Candidate drought tolerance loci identified Panel n = 226 [59] 10 Image based tissue metrics Deep CNN device for leaf area Non destructive leaf area Rapeseed DL device quantified leaf area in situ Device/controlled [60] 11 Callus developmental roadmap Time series RNA seq + network mining Callus induction time course Soybean Key callus regulators mapped for culture optimization Lab time series [61] 12 Functional genomics atlas Review + case syntheses Multi omic targets in stress Rapeseed Stress tolerance targets summarized for editing _ [62] 13 PAM-less CRISPR toolkit SpRY (engineered Cas9); broad target design and scoring gRNA design; PAM expansion Soybean Greatly expanded editable sites; validated in planta _ [63] 14 Early drought impact detection & nowcasting Random Forest; decomposition of physiological vs structural signals Satellite SIF + optical/thermal archives Multi-crop landscapes Physiological changes explained 60%–97% of functional response; anomalies detectable ~1 month before drought peak in drier regions Global [64] 15 Sub-seasonal soil-moisture drought forecasting RISE-UNet deep learning (physics-aware) Reanalysis + in-situ soil moisture Multi-crop regions Skillful forecasts at 1 to 4 weeks lead times for soil-moisture drought Global hindcasts [36] 16 Field digital twins for management and quality Digital-twin ML framework Multisensor orchard data (weather, soil, imagery) Citrus (mandarin) Intra-orchard analysis explained more variation in quality than inter-orchard; demonstrated operational twin Commercial orchards [65] 17 Wheat disease phenotyping from UAV imagery ML pipeline on spectral features UAV multispectral/hyperspectral Wheat Demonstrated aerial disease detection and mapping (leaf rust/other) Field plots [66] 18 Sub-canopy crop–soil discrimination Spectral unmixing (linear, bilinear, sparse) Drone hyperspectral + ground HSI Vegetables (field setting) 99%–100% crop–soil discrimination depending on height and endmember source Field trial [67] 19 Global UAV trait dataset (N status and yield) Meta-analysis dataset enabling ML UAV RGB/multispectral/hyperspectral (compiled) 11 major crops 11,189 observations across 41 studies; VI–trait relationships by stage 13 countries [68] 20 Regional wheat yield prediction 1D-CNN vs ML baselines Weather, soil, phenology Wheat CNN outperformed traditional models for county-level yield 271 counties (Germany) [69] 21 Genomics-informed prebreeding Genomic prediction + phenotyping Genome-wide markers + multi-site trials Wheat Prebreeding framework unlocked diversity; advanced progenies out-yielded contemporaries across sites Multi-environment trials [70] 22 Root system adaptation and drought Dynamic genetics + MRI phenotyping MRI of roots + GWAS Maize Allelic variation (e.g., ZmHb77) linked to hydraulic conductance and root architecture under water deficit Seedling to field links [71] 23 Genomic selection platform (production use) AutoGP (integrated ML/DL stack) Genotypes + breeding phenotypes (web platform) Maize End-to-end, breeder-facing GS platform integrating modern ML/DL models; emphasizes reproducibility and deployment Program scale [72] 24 GP with enviromics (G × E) ML models with engineered environment features Markers + climate/soil covariates Multiple crops (breeding datasets) Shows that adding environmental features to ML-based GP efficiently captures G × E Multi-environment [73] 25 Methods guidance for AI-enabled GP Review of statistical ML + software Genomic + phenomic pipelines General crops Practical blueprint for democratizing ML-driven GP; factors that improve accuracy and gains _ [74] 26 Structure-aware editing & complex design AlphaFold 3 (diffusion architecture) Sequence → 3D complexes (protein–DNA/RNA, ligands, ions) General (incl. plant targets) Predicts joint structures and interactions with substantially improved accuracy vs prior methods; enables editor & TF design decisions Foundation model [75] 27 Protein engineering for stress pathways/editors ProteinMPNN (graph NN for sequence design) Target backbones (incl. designed editors, chaperones) General High sequence recovery (~52%) and robust experimental validation; practical for designing enzymes/regulators used in crops Multi-system [21] 28 CRISPR guide design (RNA targeting) DeepCas13/ML models Large CRISPR-Cas13d screens General Deep learning predicts on-target activity and characterizes off-target viability effects for Cas13d Large-scale screens [76] 29 Generative gRNA design BADGERS (model-directed Cas13a guide generation) Population-scale genomes General Model-generated guides improved nucleic-acid detection sensitivity and variant discrimination vs conventional designs Multi-virus datasets [77] 30 Cell-specific GRN inference (target nomination) scMultiomeGRN (cross-modal attention + GNN) Single-cell multiome (RNA + ATAC) General DL framework outperforms prior GRN inference; supports identifying cell-type-specific regulators for editing Multi-dataset [78] 31 Microscopy automation for lab phenotyping LVACR (deep-learning 3D segmentation/aggregation) Light-sheet & X-ray imaging Plant tissues Fully automated 3D cell reconstruction; enables atlas-level quantitation for regeneration and culture optimization Multi-tissue atlas [79] 32 Organelle/structure segmentation OrgSegNet/Plantorganelle Hunter (DL segmentation) Confocal/light microscopy Plant cells State-of-the-art organelle segmentation across species; supports quantitative lab pipelines Cross-species [80] 33 Cell-type-agnostic regulatory prediction Multi-modal Transformer (masked-accessibility pre-training) DNA sequence + chromatin accessibility General Learns regulatory representations to predict expression and nominate causal motifs/links—useful for promoter/circuit design Multi-modal [81] AI-guided genome editing
-
AI-guided genome editing uses machine-learning models to forecast guide RNA (gRNA) specificity, on-target efficiency and off-target risk for CRISPR/Cas pipelines. Deep-learning models such as CNNs are widely used to scan genomic sequences and predict gRNA activity; for example, DeepCRISPR, a deep learning platform for predicting guide RNA (gRNA) on-target efficiency, and off-target profiles[20]. Hybrid models combining CNNs with attention mechanisms or recurrent layers (e.g., CRISP-RCNN) further improve predictive power. Gradient-boosting approaches also contribute: the Rule Set 3 model incorporates Light Gradient Boosting Machine (LightGBM) to integrate trans-activating CRISPR RNA (tracrRNA) identity and sequence context, improving on-target predictions (Fig. 3)[20]. Traditional algorithms remain relevant—BoostMEC, a LightGBM-based model for predicting guide RNA editing efficiency, outperformed several deep-learning models while providing interpretable feature insights. Comparative analyses show that random forest, support vector machine (SVM), gradient-boosted regression trees, and other conventional classifiers can still compete with deep learning for gRNA design when trained on high-quality datasets[82].
AI models also support multiplex genome editing by predicting editing outcomes across multiple targets and balancing editing efficiency with off-target risk. Integrative frameworks that couple sequence features with chromatin accessibility and expression context guide gRNA selection for polygenic drought traits. AI-driven protein-structure prediction enhances rational editing: the AlphaFold series (AlphaFold2 and the generative AlphaFold3) achieves near-experimental accuracy in modelling protein structures. Structural insights enable the design of high-fidelity Cas variants and optimized base editors. Recent studies used AlphaFold2 to predict Cas13 structures and then applied iterative domain cropping to generate miniaturised variants that retained activity[20]. Generative models such as Evo, trained on tens of thousands of microbial genomes, have produced entirely novel CRISPR–Cas complexes and gRNAs[20]. Deep-learning frameworks like ProteinMPNN further accelerate protein engineering: this message-passing neural network designs amino-acid sequences for given backbones and achieves higher sequence recovery (≈ 52%) than Rosetta and AlphaFold redesigns, demonstrating broad applicability to monomers, oligomers, and binding proteins[21]. Together, sequence-to-function predictors and structure-aware generators reduce wet-lab search space, delivering edited lines with clearer hypotheses to test in the field.
Synthetic biology and gene expression engineering
-
AI-driven approaches underpin modern synthetic biology pipelines for drought tolerance. Codon optimisation and in silico gene synthesis rely on machine-learning models that analyse genomic and transcriptomic data to predict codon usage patterns, maximizing translation efficiency and stability under stress. Generative models extend this by designing synthetic promoters and regulatory elements: transformer-based architectures and large language models trained on plant promoter datasets can predict cis-regulatory elements and generate novel promoter sequences tailored for stress-inducible expression. Machine-learning frameworks have been used to screen promoter libraries and identify high-performance promoters with cell-state specificity[83]. CNN- and SVM-based promoter models include Depicter (deep learning) and iProEP (SVM). Together, they predict promoter presence and classify promoter strength (Fig. 3). Hybrid models that combine CNNs, long short-term memory (LSTM) layers, and graph embeddings (e.g., DeePromClass) further enhance promoter identification[22]. Such tools facilitate de novo construction of synthetic promoters by recombining cis-regulatory elements in silico, enabling precise control of gene expression under drought.
Predictive modelling of protein sequences and regulatory pathways accelerates engineering of osmoprotectants, signalling proteins, and metabolic enzymes. Deep-learning tools like ProteinMPNN design amino-acid sequences for target backbones, achieving high sequence recovery and validating designs via X-ray crystallography and cryogenic electron microscopy (Fig. 3)[21]. Generative models such as Evo produce novel CRISPR–Cas variants and gRNAs with improved activity[20]. These advances, coupled with high-throughput phenotyping platforms for rapid feedback, enable iterative design-build-test cycles that optimise synthetic circuits for drought tolerance while balancing stress resistance and yield (Fig. 3). Collectively, AI-powered codon optimization, promoter design, and protein engineering provide a modular toolkit to program drought responses and generate edited or engineered candidate lines with traceable mechanisms and predicted performance; the next imperative is to demonstrate that these lines perform under real field variability. Accordingly, the pipeline transitions to AI-enabled field-scale breeding—data integration, trial optimization, and phenotyping—to validate lab candidates, rank them across environments, and close the loop back to selection.
-
Field-scale breeding is the critical stress test for lab-derived candidates under real environmental heterogeneity. In this stage, edited/engineered lines and conventionally selected genotypes are evaluated across sites and seasons using envirotyping covariates and multi-environment trials (MET), so that performance can be ranked for deployment and selection decisions can be updated under realistic weather and management variability[84]. Digital twins and reinforcement learning (RL) can then use these field data to optimize trial allocation, irrigation scheduling, and designed stress gradients, and the resulting performance signals feed back into model retraining and next-cycle target selection (Fig. 4)[24,84].
While many AI drought-phenomics workflows are first validated in temperate row crops, the same pipeline is highly relevant to tropical and subtropical systems where drought commonly co-occurs with heat, high humidity, and heterogeneous management. A concrete subtropical perennial example is an agricultural digital twin for mandarins that aggregated multi-source island-scale data and showed that intra-orchard (micro-environmental) resolution explained fruit-quality variation substantially better than inter-orchard summaries, highlighting why fine-grained envirotyping and management covariates are essential for decision support in orchards[65]. Complementing this, a recent synthesis of cassava (a drought-resilient tropical staple) emphasizes the accelerating shift from conventional selection toward genomic selection and gene editing for stress-resistance improvement, an ideal fit for a 'lab-to-field' pipeline that couples target discovery to field validation[85]. In tomato, where heat tolerance is a major breeding priority, tropical tomato represents a logical next target for extending this framework to tropical horticultural systems under compound climate stress[86]. Likewise, rice accessions adapted to combined heat and drought stress represent a relevant extension, given the recognized importance of simultaneous heat and drought stress in rice improvement[87]. Finally, for tropical cereals, recent G × E-aware genomic prediction work that explicitly fuses genotype with temporally resolved environmental variables (demonstrated across maize, rice, and wheat) provides a practical template for embedding envirotyping into selection decisions rather than treating environment as a nuisance factor[88].
Lab-to-field generalization: genotype-by-environment (G × E) domain shift
-
A central reason drought-tolerance mechanisms validated in controlled conditions may underperform in farmers' fields is genotype-by-environment (G × E), where drought timing and intensity, soil constraints, and management can reshape effect sizes and re-rank genotypes. Field-focused modeling provides practical ways to address this gap. Transformer-based genotype–environment frameworks (e.g., GEFormer) explicitly combine genotype with time-resolved environmental variables and are evaluated under realistic breeding scenarios that include untested environments, directly targeting lab-to-field generalization[88]. Likewise, MET studies show that incorporating engineered environmental features into machine-learning genomic prediction can indirectly model G × E and improve yield prediction across environments relative to classical baselines[73]. Within the proposed pipeline, this implies (i) designing training/validation splits that hold out environments or years, (ii) encoding environments via interpretable envirotyping covariates (weather/soil/management) rather than site labels alone, and (iii) using digital-twin style integration like demonstrated in a subtropical perennial system to make micro-environmental heterogeneity actionable rather than hidden noise[65].
AI in data integration and field trial optimization
-
AI has emerged as a transformative tool in field-scale breeding by enabling efficient data integration, real-time sensing, and adaptive trial optimization. Field platforms aggregate multi-source data streams by integrating climate variables (e.g., temperature and precipitation), soil characteristics (e.g., water retention), and remote-sensing indices obtained from UAVs, satellite, and ground-based sensors, thereby quantifying drought exposure and plant water status for informed breeding decisions[3,24]. Deep-learning models, particularly convolutional neural networks, are trained on UAV-based field imagery to automatically extract subtle drought-responsive phenotypic features that would be challenging to detect manually. By analyzing spectral, textural, and structural differences in plant images, these models enable precise phenotype mapping and assist in genotype selection through accurate identification of stress markers[4]. In parallel, modern AI frameworks continuously fuse environmental, physiological, and phenotypic data streams from sensor networks and high-throughput observations[4], supporting efficient tracking of drought-tolerant individuals across large plots and enabling rapid intervention when necessary (Table 1).
To merge and interpret large, heterogeneous datasets from field experiments, network-aware approaches can be applied where appropriate. For example, graph-based models can integrate genomic and transcriptomic information to support QTL refinement and candidate-gene prioritization[89]. A concrete validation of network-guided prioritization is provided by an integrative regulatory-network study in Arabidopsis that combined motif, chromatin, and expression evidence and experimentally confirmed stress-relevant regulators, validating 13 previously unlinked ROS/stress transcription factors via phenotypic assays and illustrating how network outputs can translate into testable intervention candidates[39]. Supervised and transfer-learning approaches can further fine-tune pretrained multimodal models, while multi-omics integration captures nonlinear genotype–phenotype relationships that can shorten breeding cycles[90]. Workflows that combine GWAS with machine learning can extract the most predictive features. These features help rank drought-adaptive lines more accurately than using raw markers alone. For example, deep-learning phenotyping paired with GWAS has identified stress markers and significant SNPs in soybean[91]. This highlights the value of AI-assisted candidate selection (Table 1; Fig. 4).
Adaptive trial design and efficient resource allocation also benefit significantly from AI-driven approaches (Fig. 4). Reinforcement learning algorithms dynamically optimize irrigation scheduling by continuously adjusting water application based on real-time plant and environmental feedback. In particular, deep reinforcement learning frameworks that integrate high-dimensional sensor inputs have demonstrated superior performance, achieving higher resource-use efficiency and profits than conventional rule-based irrigation strategies in simulated production systems[92]. In parallel, AI-powered digital twins create virtual representations of field conditions; these digital twins simulate irrigation and fertilization scenarios, enabling researchers to optimize trial allocation and control stress application across plots while reducing resource waste[24]. Moreover, reinforcement-learning mechanisms and digital-twin simulations cycle lines through designed drought gradients, promoting optimal expression of drought-resistance traits during critical developmental stages (Table 1).
Collectively, these data-integration and trial-design capabilities turn complex, heterogeneous signals into actionable field decisions, accelerating the path from candidate generation to robust, climate-ready varieties[3].
AI for predictive modeling, phenotyping, and validation
-
AI-driven breeding platforms integrate diverse multi-trait data to enable predictive modeling for drought-resilient breeding. These systems combine genomic, phenotypic, multi-environment trial (MET), and environmental data using machine-learning models, such as random forests, support vector machines, and deep neural networks, to capture nonlinear genotype-by-environment interactions. By assimilating high-throughput genomic sequencing, sensor-derived phenotypes, and climate variables (Table 1; Fig. 4), such platforms prioritize complex genotype assemblies that exhibit drought adaptation across heterogeneous sites[23,93]. To reduce deployment failures, evaluation should prioritize leave-one-environment/year-out validation and report both accuracy and calibration under 'untested environment' conditions (Table 2). Recent MET-based evidence indicates that incorporating engineered environmental covariates can efficiently approximate G × E structure within ML genomic prediction frameworks[73].
Table 2. AI model evidence maturity/readiness.
No. Component in drought-intelligence pipeline Evidence maturity What is still missing for field reliability Example evidence 1 CNN/vision models on UAV/field imagery Multiple field phenotyping studies + growing open datasets Standardized splits (site/year-held-out), label harmonization, calibration reporting [7] 2 Bioacoustic drought sensing Controlled chamber/greenhouse feasibility with ML classification Multi-site field trials, sensor standardization, confound controls (wind/insects/machinery) [28] 3 Network/GRN inference to prioritize targets Experimental validation exists in model plants Translation rate to crops, causal verification under drought field conditions [39] 4 G × E-aware genomic prediction (envirotyping + ML/transformers) Tested in realistic breeding scenarios incl. untested environments Consensus on evaluation protocols + integration with breeder decision rules [88] 5 ML combining genetic + environmental features in MET Demonstrated gains over classical baselines in MET settings Portability across regions, transparent envirotyping standards [73] 6 Digital twins for crop systems Working agricultural DT demonstration in a subtropical perennial crop Wider drought-focused validation, integration with intervention policies [65] 7 Multisensor benchmarks (EO) Benchmark datasets exist in adjacent domains Plant-stress specific benchmarks spanning sensors + physiology ground truth [94] In predictive modeling for trait pyramiding and drought management, AI models synthesize multi-omics data plus environmental inputs to simulate crop performance under variable drought stress scenarios. By constructing predictive frameworks, these models generate drought gradients in multi-location trials and dynamically integrate real-time data (soil moisture, temperature, precipitation) to maximize heritability estimates. For example, combining genomic and transcriptomic markers with ML classifiers (random forests, SVMs, XGBoost) achieved prediction accuracies up to 0.95, outperforming single-omics models[90]. Multi-omics integration that incorporates genomic, DNA methylation, and gene-expression data has increased prediction accuracy for complex traits (0.72–0.92) relative to genomic-only models (0.55–0.86)[90].
Automated AI systems leverage continuous-learning architectures by routinely retraining on new phenotypic and environmental datasets. These closed-loop feedback mechanisms update predictive models season after season, ensuring enhanced drought predictions and refined recommendations tailored to evolving environmental conditions. Simultaneously, AI-powered integration hubs consolidate data from diverse sources, ranging from high-throughput phenotyping and remote sensing to genomic and environmental monitoring, and deploy decision-support tools for real-time breeding decisions (Table 1). Such centralized data hubs enable faster response times and higher precision in managing breeding schemes under climatic uncertainty[23].
High-throughput phenotyping is advanced by UAV-based, multimodal imaging. These systems, when combined with convolutional neural networks (CNNs), enable rapid, high-resolution image analysis for early stress detection in field conditions. UAV-based remote sensing platforms acquire spectral indices (e.g., vegetation indices, canopy temperature, chlorophyll content) with high spatio-temporal resolution, providing non-destructive assessments of plant water status and facilitating water-stress detection and crop health monitoring[27]. Integrating hyperspectral/multispectral imagery with GWAS has identified chromosomal regions related to drought and heat stress, demonstrating the value of high-throughput phenotyping in identifying adaptive traits[95]. Transfer learning and few-shot learning reduce the need for manual labels. Many pipelines can fine-tune pre-trained CNNs using only 100–200 images per class and still keep strong accuracy[31]. For detection tasks, models such as YOLOv5 can speed up plant image analysis. To capture change over time, some pipelines use RNN variants for temporal patterns. For relationships across organs, pixels, or genotypes, GNN variants can add structured context[31].
Genomic innovation is further bolstered by integrating field phenotyping with genomic data to predict performance in CRISPR-edited or transgenic lines. AI-driven approaches merge sequencing data and phenotypic outcomes to construct gene regulatory networks; for example, supervised learning integrating gene expression and chromatin accessibility data has been used to identify drought-responsive transcription factors with high precision (area under the precision–recall curve, AUC – PR ≈ 0.81)[96]. Machine-learning ranking systems can score candidate cultivars for drought response and yield stability. For example, LASSO and random-forest feature selection on transcriptomic data can nominate key markers. Using those markers, an SVM classifier achieved 0.95 accuracy (AUC 0.9865) for resistant vs susceptible lines[97].
At the field-decision level, advanced AI models such as CNNs and Random Forests process multisource data—satellite imagery, sensor-network outputs, and climate models—to generate accurate soil-moisture and drought-stress maps. For example, a comparative study of soil-moisture inversion models found that Random Forest significantly outperformed support vector machines, back-propagation neural networks, and convolutional neural networks, achieving only 9.52% error relative to observed data[98]. Similarly, multi-sensor approaches using radar, optical satellite data, and digital elevation models to train feed-forward artificial neural networks achieved high correlation (R ≈ 0.80) and low Root Mean Square Error (RMSE ≈ 0.04 m3·m−3) for estimating surface soil moisture[99]. The blending of multimodal data supports near-real-time drought-risk forecasting, which is critical for adapting breeding and field-management strategies on the fly. Ensemble learning, GNNs, and reinforcement learning further optimize trait selection under complex genotype-by-environment interactions by iteratively refining selection criteria[90,99].
By explicitly modeling G × E, standardizing evaluation under untested environments, and closing the loop between field performance and model retraining, AI-enabled field-scale breeding increases the likelihood that lab-derived candidates translate into stable yield and resilience under real-world variability. In the context of intensifying drought risk, this lab-to-field integration supports faster deployment of drought-resilient cultivars, helping stabilize production and contributing to longer-term food security goals through improved yield reliability and more efficient resource use[23].
-
Developing drought-resilient crops through AI and multi-omics technologies brings significant promise, yet several challenges must be addressed to realize its full potential (Table 2). The integration of multi-source field data remains a major obstacle. AI models trained on one dataset often struggle to generalize to others because biological and technical variations create context-specific patterns[100]. High-throughput sensing generates noisy, sparse, and imbalanced datasets, making it difficult to distinguish genuine biological signals from background noise[101]. Overfitting is a persistent risk in deep learning, and the 'black-box' nature of many models hampers interpretability and adoption[102]. Future research must standardize sensor protocols, develop self-supervised and transfer-learning methods to improve cross-environment robustness, and incorporate explainable AI to provide biologically meaningful insights. Quantum-inspired algorithms and reinforcement learning may enhance dynamic trial optimization, while digital twins that fuse remote sensing with ground truth could support adaptive field management.
A major barrier to reproducible progress is the lack of benchmark-ready, multi-modal drought datasets with agreed evaluation protocols (Table 2). Recent work in adjacent plant phenomics and EO demonstrates the value of curated, multi-site compilations and benchmark experiments, but drought-intelligence still lacks standardized splits and reporting that directly quantify generalization (e.g., leave-one-location/year-out and 'untested environment' evaluation)[68,88]. We recommend that future community benchmarks report: (i) task performance (e.g., RMSE/MAE for continuous traits; AUROC/F1 for detection), (ii) calibration of probabilistic outputs (reliability curves/expected calibration error), and (iii) lead-time utility for early warning (how many days/weeks before visible symptoms). Benchmarking should also encode sensor/platform variability (satellite–UAVs–in situ), and leverage existing multi-sensor benchmark concepts where appropriate, while prioritizing stress-ground-truth anchoring from physiology and yield[68,94].
Beyond benchmark construction, a critical next step will be the development of multimodal foundation models capable of learning shared representations between temporally resolved field-sensing data and mechanistic omics layers. Recent reviews in plant bio-genomics suggest that genomic language models may broaden the scope of plant sequence analysis and regulatory inference, though their effective application in plant science remains in its early stages[103]. At the plant scale, emerging signals and model architectures are expanding analytical sensitivity. For instance, recent single-cell foundation models have demonstrated that pretrained transformers can capture transferable biological structure at scale, supporting tasks such as multi-omic integration, perturbation-response prediction, and gene-network inference. Similarly, multiome-based regulatory frameworks leverage paired chromatin accessibility and gene-expression data to recover cell-type-specific regulatory programs and project these signals into expression-only datasets[40, 49]. Multimodal regulatory transformers further suggest that cross-modal attention can learn generalizable mappings across heterogeneous molecular inputs, even in previously unobserved cellular states[81]. Although the direct integration of temporal satellite or UAV trajectories with single-cell multiome data remains largely prospective in crop drought biology, such models hold transformative potential. They could ultimately align field-scale stress dynamics with cell-resolved regulatory states, improve transferability across crops and environments, and reduce dependence on extensively annotated field datasets through pretraining, weak supervision, and few-shot adaptation[40, 49].
While advances in hyperspectral, thermal, and radar imaging have significantly improved early drought detection, key technological and operational hurdles persist. These include the need for precise sensor calibration, seamless multi-scale data fusion, and the development of cost-effective, widely accessible monitoring platforms. Emerging tools such as bioacoustic sensing and plant wearables remain largely experimental, with their practical adoption constrained by field confounds, a lack of calibration standards, and insufficient multi-site validation (Table 2). Promising but nascent approaches like quantum-enabled forecasting demand considerable computational resources, with most work to date being proof-of-concept and lacking standardized field benchmarking[25]. Future directions involve integrating remote sensing data with omics information, deploying self-supervised models for real-time anomaly detection, and developing cross-platform frameworks that unify satellites, UAVs, and ground-based sensors. Addressing combined stressors—where plants experience drought along with heat, salinity, or disease—will be critical to mirror real-world conditions.
Understanding plant mechanisms is challenging because drought perception and signalling networks—encompassing osmosensors, mechanosensitive channels, reactive oxygen species (ROS) waves, peptides, hormones, epigenetic modifications, and microbiome interactions—are exceptionally complex and present significant analytical and experimental hurdles (Fig. 2). Multi-omics datasets are high-dimensional and often noisy; integrating them demands sophisticated feature selection and information-extraction methods[104]. Many regulatory interactions remain poorly characterised, particularly those involving long non-coding RNAs, transgenerational epigenetic memory, and cross-talk between drought and other stresses. Future studies should leverage graph neural networks and integrative modelling to unravel causal networks, expand single-cell and spatial omics analyses to resolve tissue-specific responses, and explore how root–microbiome–soil interactions shape drought resilience.
Laboratory breeding and genome editing, despite notable progress in automated tissue culture, still face limitations. Image-based phenotyping requires large, well-annotated datasets, and its transferability across species and tissue types remains constrained. In genome editing, the accurate prediction of on-target efficiency and off-target risk continues to be a major challenge, as models trained in one organism often fail to generalize to others[100]. Ethical and regulatory considerations pose significant challenges, requiring transparent reporting and governance frameworks[105]. Future work should expand open, curated datasets of tissue images and gRNA outcomes, develop multimodal models that integrate sequence, chromatin, and structural information, and couple protein-structure prediction with gRNA design to improve specificity. Generative models for synthetic promoters and regulatory elements must be validated experimentally, and guidelines for responsible use of AI-driven editing should evolve in parallel with technological advances (Fig. 3).
Integrative and societal perspectives underscore that, across all domains, ensuring data privacy, fair access, and equitable benefit sharing is essential. Collaboration between plant scientists, data scientists, ethicists, policymakers, and growers will be vital to translate AI innovations into field-ready solutions[100]. Scaling these approaches will require robust computational infrastructure, training programs for interdisciplinary researchers, and open-source platforms that democratize AI tools. By addressing these challenges and pursuing these future directions, the integration of AI, multi-omics, and advanced breeding technologies can sustainably accelerate the development of drought-resilient crops and contribute to global food security in a changing climate.
-
Drought-resilient crop breeding becomes more feasible when pipelines are redesigned around timely, information-rich signals and closed-loop learning. Multimodal sensing, spanning satellites, UAVs, in situ probes, and bioacoustics, combined with foundation-style, self-supervised models, can shift detection from lagging, symptom-based assessments to proactive, probabilistic nowcasts with explicit uncertainty, enabling earlier irrigation decisions and real-time selection indices. These early warnings are most powerful when anchored to mechanisms: osmo- and mechanosensory Ca2+ influx, ROS dynamics, peptide and hormone circuits, and stomatal ion-channel cascades provide a causal scaffold against which AI can learn. Mechanism-informed multi-omics integration across the epigenome, transcriptome, proteome, metabolome, and microbiome, together with graph neural networks and causal inference, translates complex drought biology into actionable shortlists of candidate genes, TF hubs/regulators, regulatory edges, and cis-regulatory elements for editing or selection. Field breeding accelerates when phenomics and mechanistic priors drive continuous-learning optimization: data-fusion frameworks standardize environmental covariates, genotype-environment models capture nonlinear responses, and reinforcement learning with digital twins reallocates trials and resources to maximize information yield while hybrid GWAS-ML pipelines reduce overfitting. Laboratory workflows now keep pace through CNN- and GAN-enabled imaging for tissue culture staging, machine-learning refinement of gRNA efficiency and specificity, and multiplex CRISPR designs guided by AlphaFold, ProteinMPNN, and generative promoters to compress the design-build-test cycle and stabilize expression under drought. Together, these advances define a practical 'pixels-to-genes-to-fields' roadmap: pretraining multisensor encoders, coupling them to graph-based multi-omics with uncertainty quantification, integrating digital twins for resource-aware optimization, and connecting directly to automated genome editing. With explainability, calibration, and equitable governance embedded, this framework can deliver climate-ready crop varieties and a generalizable template for resilient agriculture.
This work was supported by the National Natural Science Foundation of China (32271913), the National Tropical Agriculture Science and Technology Innovation Project for the Chinese Academy of Tropical Agricultural Sciences (CATAS202617), the Project of State Key Laboratory of Tropical Crop Breeding (NKLTCBZRJJ6).
-
The authors confirm contribution to the paper as follows: literature review, draft manuscript preparation: Mushtaq MA; figure preparation, manuscript writing enhancement: Mushtaq MA, Tian X; manuscript review and editing: Dai C, Ma C, Zhang J. All authors reviewed the results and approved the final version of the manuscript.
-
The datasets generated during and/or analyzed in the current study are available from the corresponding author on reasonable request.
-
The authors declare that they have no competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
-
Received 17 November 2025; Accepted 2 March 2026; Published online 30 April 2026
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press on behalf of Hainan University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Mushtaq MA, Tian X, Dai C, Zhang J, Ma C. 2026. AI-aided early sensing, decoding, and lab-to-field breeding for drought-resilient crops. Tropical Plants 5: e012 doi: 10.48130/tp-0026-0011
AI-aided early sensing, decoding, and lab-to-field breeding for drought-resilient crops
- Received: 17 November 2025
- Revised: 11 February 2026
- Accepted: 02 March 2026
- Published online: 30 April 2026
Abstract: Drought stands as the foremost abiotic constraint on global crop productivity. With climate change increasing the frequency and severity of drought events, a paradigm shift toward faster, more predictive, and mechanistically informed breeding is urgently required. This review synthesizes current advances to propose a connected 'pixels-to-genes-to-fields' framework, integrating early drought phenotyping, causal gene discovery, AI-assisted laboratory engineering, and field-scale validation. We first examine how multimodal monitoring platforms, from satellites and UAVs to in-field sensors, coupled with advanced AI models, enable early stress detection and predictive risk mapping. We then distill the complex mechanistic pathways of drought response, spanning perception (e.g., OSCA, MSL), signaling (ROS, CLE-ABA), stomatal regulation, and epigenetic memory, into structured biological priors. These priors, we argue, are crucial for guiding graph-based AI in identifying high-confidence genetic intervention points. At the field scale, we survey strategies where AI integrates genotype, environment, and phenomics data to model genotype-by-environment interactions and optimize trials via digital twins. At the laboratory scale, we summarize the role of AI in accelerating the design-build-test cycle through precision CRISPR design, synthetic expression engineering, and automated phenotyping. Finally, we highlight critical translational challenges, emphasizing the need for standardized data sharing, explainable AI, and responsible governance to bridge these innovations into the development of scalable, drought-resilient crop varieties.