-
Drought is one of the major abiotic stresses that limits plant growth, development and yield worldwide. Climate change and soil aridification are predicted to reduce soil moisture content and affect crop growth significantly[1]. Therefore, attaining a better understanding of the molecular mechanisms that underlie plant's drought response is needed to develop drought mitigation strategies.
Tomato (Solanum lycopersicum) is a valuable fruit and vegetable crop with profound economic and nutritional values. International trade in tomato products has increased over the years, making tomato production an essential component of global agriculture[2,3]. Drought stress severely affects tomato yield and this has negative impacts for both the tomato industry and agricultural economy. Currently, ABA (abscisic acid) is known to play a crucial role in regulating plant's drought response through acting on a number of protein kinases, transcription factors (TFs), various hormonal signaling pathways, and membrane/channel proteins. In this review, we will highlight the most recent information of how these components contribute to the molecular mechanisms underlying tomato's responses to drought.
-
In the presence of low water availability, osmotic stress can occur. Within seconds of osmotic shock, plants increase their cellular concentration of free calcium ions ( [Ca2+]cyt )[4−6]. It is reported that plants modulate [Ca2+]cyt via OSCA1 (REDUCED HYPEROSMOLALITY-INDUCED CA2+ INCREASE 1)[7,8]. Plants can also utilize AQPs (aquaporins) to regulate the transport of water, small neutral solutes, and CO2 between plasma membranes[9−12] . In tomato, plasma membrane intrinsic proteins SlPIP2.1, SlPIP2.5 and SlPIP2.7 were found as candidate AQPs for contributing root water uptake since they can increase water content and maintain osmotic balance[13].
Plants can mediate intracellular signals and gene expression in response to drought via two different ABA signaling pathways: ABA-dependent and ABA-independent pathways[4,14]. ABA-dependent pathways are more common in plant responses to drought stress. Osmotic stress is also known to induce ABA synthesis[15]. As an endogenous messenger for biotic and abiotic stresses, ABA acts as a long-distance water stress signal for monitoring early signs of drought[16]. In fact, ABA plays a crucial role in plant response to drought because it triggers protective mechanisms such as regulating stomatal closure and gene expression to prevent further water loss. The molecular mechanisms underlying the responses of plants to drought involve a series of actions. Multiple processes starting from elaborate sensing, signaling, transcription, transcript processing, translation to post-translational protein modifications all contribute to plant's drought response. The ABA signaling pathway is by far the most well recognized stress response mechanisms utilized by plants to regulate abiotic stress.
The ABA-dependent signaling pathway is composed of ABA receptors PYR/PYL/RCAR, co-receptors PP2Cs, SnRK2 kinase and TF ABI5/ABFs[17]. Under normal conditions, PP2C binds to SnRK2, thereby inhibiting the ABA signaling pathway. When plant roots detect water deficiency, drought signal is transmitted to the stem, leading to the accumulation of ABA throughout the plant[18]. The ABA receptor PYR/PYL/RCAR recognizes and binds to ABA, which causes the release of the phosphorylation site of SnRK2 from PP2C. This in turn allows SnRK2 to phosphorylate downstream proteins, such as the cis-acting element, ABA-responsive element (ABRE), and ABRE-binding protein/ABRE-binding factors (AREB/ABF) TFs[19−21].
In the ABA-independent pathway, a cis-element, dehydration-responsive element/C-repeat (DRE/CRT) and DRE-/CRT-binding protein 2 (DREB2) TFs contribute pivotal roles in response to drought and osmotic stress[22]. DREB2 proteins are AP2/ERF family members in plants. Arabidopsis DREB2A and DREB2B were found to be highly inducible by drought, high salinity, and heat stress[23,24]. In fact, it is the key TF for osmotic stress response in the ABA-independent pathway. Under normal conditions, DREB2A is repressed by GROWTH-REGULATING FACTOR7 (GRF7) to maintain plant growth. DREB2A-INTERACTING PROTEINs (DRIP1 and DRIP2), which are ubiquitin E3 ligases, target DREB2A to 26S proteasome for proteolysis under normal conditions. DREB2A protein is known to be instable in plants and once it is activated, it can induce a myriad of genes that encode proteins responsible for stress response and tolerance. Some of these genes encode the LEA (late embryogenesis abundant) proteins, which are thought to protect enzymes and lipids from drought-induced dehydration[25].
Since GRF7 was discovered to repress DREB2A, ABA-inducible genes, and osmotic stress-inducible genes, crosstalk was suggested to occur between ABA-independent and dependent signaling pathways. Furthermore, AREB/ABFs in the ABA-dependent pathway have been shown to interact with DREBs to regulate drought stress response, highlighting both ABA-independent and dependent pathways are involved in mediating plant's response to drought[26,27]. Most of what we know about how plants respond to drought via the ABA-independent pathway and crosstalk of the two signaling pathways came from Arabidopsis studies. This information remains to be explored in tomato, and crosstalk is suspected to occur between the two signaling pathways in tomato.
-
Protein kinases are a class of enzymes that catalyzes protein phosphorylation. Protein kinases that are involved in drought stress response include SNF1-related protein kinase 2 (SnRK2), mitogen-activated protein kinase (MAPK), and histidine kinase (HK). SnRK2.6, also known as OST1, is a SnRK2 family member that has been reported to play a major role in plant's response to drought[28,29]. SnRK2.6 can phosphorylate several ion channel proteins of SLAC1 (SLOW ANION CHANNEL ASSOCIATED 1), QUAC1 (the R-/QUAC-type anion channel), and KAT1 (POTASSIUM CHANNEL IN ARABIDOPSIS THALIANA 1) under drought stress to positively regulate ABA/drought-induced stomatal closure[30]. In addition, SnRK2.6 can phosphorylate several TFs including ABF2 (ABSCISIC ACID RESPONSIVE ELEMENT BINDING FACTOR 2) and ABI5 (The PP2C ABA INSENSITIVE 5) to regulate drought responses at the transcriptional level[19,28]. In tomato, SnRK2.6 can modulate the balance between plant growth and development under drought. In fact, tomato SnRK2.6 can enhance the protein stability of NAC (NAM (No Apical Meristem), ATAF1 (Arabidopsis thaliana Activation Factor 1), and CUC2 (Cup-shaped Cotyledon)-type TF SlVOZ1 (VASCULAR PLANT ONE-ZINC-FINGER) through phosphorylation, thereby regulating the expression of flowering gene SFT (SINGLE FLOWER TRUSS) and promoting drought-induced flowering[31]. Arabidopsis SnRK2s can phosphorylate sucrose transporters SWEET11 and 12 to enhance their oligomerization and sucrose transport activity under drought conditions, which help to increase the root's sucrose content and plant root-to-shoot ratio[32].
The MAP kinase cascade also contributes to plant's drought response through the phosphorylation of MAPKKK, which can be activated by lipid membrane receptors or extracellular stimuli. Activated MAPKKK promotes the phosphorylation of MAPKK, which subsequently activates MPK through dual phosphorylation[33]. The MAPK cascade is a highly conserved signal transduction pathway in plants. In tomato and multiple plant species, the transcription, protein accumulation, and kinase activity of MAPK signaling components are regulated by drought and ABA signals[34]. In Arabidopsis, the E3 ubiquitin ligases RGLG1 (RING DOMAIN LIGASE 1) and RGLG2 (RING DOMAIN LIGASE 2) ubiquitinate MAPKKK18 at lysine residues K32 and K154, thus promoting MAPKKK18 degradation. The absence of these two ubiquitin ligases stabilizes MAPKKK18 and enhances the drought tolerance of MAPKKK18-OE lines[35]. Water deficit or ABA treatment rapidly up-regulate MAPKKK levels in plants and inhibit the expression of RGLG1 and RGLG2, leading to the accumulation of MAPKK18, thereby increasing plant resistance to drought stress[36]. SlMAPK1 overexpression enhanced the relative expression of a series of genes associated with antioxidant enzyme activities, such as SlAPX (ascorbate peroxidase), SlCAT (catalase), SlPOD (peroxidase), and SlGST (glutathione S-transferase zeta class-like isoform X1). These changes led to lower ROS accumulation in SlMAPK1-OE lines than in the WT plants. Tomato plants overexpressing SlMAPK1 showed less H2O2 and ROS accumulation and exhibited higher antioxidant enzyme activity under drought stress[37,38].
HKs serve diverse functions in plant development and responses to environmental stress. They are particularly known as osmotic stress sensors in plants. In Arabidopsis, AHK2, AHK3, and AHK4 negatively regulate ABA sensitivity and drought response, while AHK1 positively regulates drought response through the activation of ABA-responsive gene expression, ultimately enhancing plant drought tolerance[39,40]. In tomato, SlHK2 has been shown to specifically respond to drought. Inhibition of the cytokinin signaling pathway in SlHK2-RNAi plants resulted in increased tolerance to drought and high-temperature stress[41].
-
Drought stress induces an excessive amount of reactive oxygen species (ROS), which lead to severe cellular damage, leaf wilting and declined photosynthetic efficiency in plants[42]. Despite the negative effects that ROS can cause to plants, increasing evidence is suggesting that ROS accumulation is an adaptive response of plants under stress situations. If they are balanced properly, ROS can act as stress-responsive substance and plants can circumvent some of the serious defects caused by ROS such as programmed cell death. The hormone of ABA is associated with ROS because ABA can engage with the plant's non-enzymatic scavenging system to eliminate ROS. ABA can also integrate with Ca2+ (calcium) signaling to transmit stress signals to intracellular sensor proteins and initiate downstream stress responses. Since Ca2+ is a versatile second messenger, ABA can further trigger Ca2+ signals and act on Ca2+ channels to induce stomatal pore closure in response to drought.
ABA can further help plants survive drought through acting on the PYR/PYL/RCAR receptors, which are responsible for inducing downstream TFs such as AREB/ABFs via a cascade of kinase reactions[18,43−46]. Thus far, PYL9 has been reported to participate the most in drought resistance. ABRE is a conserved cis-acting regulatory element in the promoter of stress-induced genes and AREB/ABF proteins can bind to the ABRE element in the promoter region of stress responsive genes and regulate gene expression under stress[47−49]. In plants, overexpression of the AREB1/ABF2, AREB2/ABF4, and ABF3 genes can enhance plant's drought resistance[4,43,50−53].
In addition to ABA, phytohormones such as brassinosteroids (BRs), jasmonic acid (JA), and ethylene (ETH) have also been reported to be involved in plant's response to drought. In particular, BRs have been documented to positively regulate drought resistance in tomato[54]. Exogenous BRs can improve drought tolerance through increasing leaf water potential and water absorption capacity. BRs can also reduce transpiration rate and maintain metabolic responses. Moreover, BRs increase the activities of POD, CAT, ABA and carotenoids to eliminate ROS, stabilizing cell membrane and alleviate stress damage[55,56]. In Arabidopsis, BR regulates early ABA signaling mainly by affecting the phosphorylation of SnRK2.2, SnRK2.3, and SnRK2.6[51,57]. However, ABA accumulates in plants and antagonizes BR regulators under unfavorable conditions such as drought[28,58,59]. In tomato, high level of SlBRI1 negatively influences tomato's drought tolerance. High expression of SlBRI1 leads to increased BR signaling, which not only augments H2O2 content but also reduces stomatal closure, antioxidant enzyme activity, and the expression of drought stress-related genes[60].
Like BR, JA also coordinates with ABA to synergistically activate drought stress responses to regulate stomatal closure, extracellular Ca2+ influx, H2O2, and NO signaling. The crosstalk between JA and ABA signaling pathways can induce the transcriptional activation of the basic-helix-loop-helix (bHLH) TF MYC2 (MYELOCYTOMATOSIS2). MYC2 has been reported to serve as a master regulator of the JA signaling pathway in Arabidopsis. MYC2 in tomato was shown to negatively regulate PP2C1 of the ABA signaling and RR26, a type-B response regulator in the cytokinin pathway that was shown to decrease drought tolerance in tomato[60,61]. ETH is another hormone that links with ABA, JA, BR, auxin, GA (gibberellic acid), SA (salicylic acid) and CK (cytokinin) to help plants mediate the negative effects of abiotic stress on plant growth and development. ETH has the ability to induce defense and survival mechanisms in response to abiotic stress such as drought, thereby influencing plant growth and structure maintenance under drought conditions[62,63]. The involvement of ETH receptors in drought stress response are complex. ERF (ethylene response factors) are AP2/ERF (APETALA2/ethylene-responsive factor) superfamily proteins and they are the largest family of TFs reported to participate in various abiotic stresses including drought. In tomato, SlERF5 gene can be induced by various abiotic stresses. Transgenic tomatoes overexpressing ERF5 showed resistance to drought and salt stresses[64].
It is evident that phytohormones play essential roles in plant's tolerance to drought. Their intricate network of crosstalk leads to transcriptional reprogramming in plants and phytohormone crosstalk in drought stress response will be briefly discussed next. Drought stress inhibits both salicylic acid (SA) and GA biosynthesis to help plants conserve energy for growth. Lower amount of circulating SA in plants leads to higher ABA production, thus increasing the chances for ABA to synergize with other phytohormones to induce drought stress response. Like ABA, phytohormones of JA, BR, and auxins are also increased under drought conditions. While BR signaling can suppress JA production, it helps plants to respond to drought via promoting ABA and auxin biosynthesis. ABA and auxin can then coordinately regulate stomatal aperture. ABA also cooperates with ETH and SA to inhibit CK production, thereby mediating plant's drought tolerance mechanism[4,65].
-
TFs are proteins that bind to specific DNA motifs in the promoter regions of target genes to regulate their transcription. TFs possess multifunctional roles in signal transduction networks ranging from stress signal sensing to expressing stress-responsive genes. Often times, stress mechanisms such as drought are complex and multiple TFs are involved to mediate numerous pathways simultaneously in plants. Thus far, more than 80 TF families have been isolated from a wide range of crops[66]. TFs that have been reported to regulate plant drought resistance include the NAC (NAM, ATAF1/2, CUC1/2), MYB (v-myb avian myeloblastosis viral oncogene homolog), WRKY (WRKY-type TF), AP2/ERF, ZFP (zinc-finger protein), and bZIP (basic region/leucine zipper). We will next provide a brief description of their roles and functions in drought stress response.
NAC
-
To date, a number of NACs has been shown to regulate plant's drought response. In Arabidopsis, ANAC096 encodes a key NAC TF in the ABA-independent signaling pathway and it has been shown to interact directly with ABF2 and ABF4 to regulate gene expression under drought stress[67]. In transgenic tomato plants, the silent expression of SlNAC4 was found to reduce their water and chlorophyll content, while accelerated their desiccation under drought stress[68]. In SlNAC6-RNAi seedlings subjected to drought, the ABA content and expression level of the genes in the ABA signaling pathway were reduced. These seedlings also exhibited signs of lower tolerance to PEG-stimulated osmotic stress, and increased water loss and oxidative damage[69]. Additionally, JUNGBRUNNE1 (JUB1) in the NAC family was identified as a regulator of tomato drought tolerance. Silencing SlJUB1 caused an excessive accumulation of H2O2 in tomato plants, leading to oxidative damage and more water loss under water-limiting conditions, thereby negatively affecting tomato's drought tolerance and expression of drought-responsive genes SlDREB1, SlDREB2, and SlDELLA[42]. In tomato, JUB1 was demonstrated to directly activate the expression of DREB2A, and inhibit intracellular H2O2 accumulation by directly activating DELLA for protein accumulation[70]. The reduction of H2O2 levels in tomato was dependent on the activation of SlDREB1, SlDREB2 and SlDELLA by SlJUB1[42].
MYB
-
As one of the largest TF families in plants, MYB plays an important role in plant's responses to various biotic and abiotic stresses. The MYB TF family is divided into four subclasses by the number of N-terminal structures: 1R-MYB/MYB-related, R2R3-MYB, 3R-MYB/R1R2R3-MYB, and 4R-MYB-containing, which consists of 1, 2, 3, and 4 repeat sub-structures, respectively. Among them, the R2R3-MYB proteins are the most widely distributed class of MYB TFs in plants. They have been reported to participate the most in abiotic stress regulation. These proteins can regulate secondary metabolite biosynthesis, hormone signaling, and stress responses[71]. In tomato, the overexpression of SlMYB49 was shown to enhance the plant's drought tolerance via eliminating ROS, inhibiting cell membrane damage, and suppressing cell death[72]. Silencing SlMYB50 in tomato could significantly reduce the levels of O2 and H2O2 accumulation as well as increase the activation of genes related to chloroplast development, flavonoid and ABA biosynthesis to cope with osmotic stress[73]. Evidently, flavonoid content regulated by SlMYB55 can help with tomato's drought resistance[74].
AP2/ERF
-
The AP2/ERF TFs have the ability to mediate biological and physiological roles such as stress responsive mechanisms, hormone signal transduction, plant morphogenesis, and metabolite regulation. They are classified into four subfamilies: DREB, AP2, ERF, RAV (Related to ABI3/VP), and Soloists[75]. DREBs and ERFs are reported mostly in the involvement of stress responsive mechanisms. Originally discovered in tobacco, ERFs could bind to the ETH response element GCC box and affect ETH signal transduction[76]. The roles of ERF TFs have been expanded to stress protective mechanism and responses in other plants. In Arabidopsis, AtERF84 overexpression can enhance the plant's ROS scavenging ability and improve drought and salt stress tolerance[77] while the overexpression of SlERF5 in tomato can trigger similar stress protective mechanisms[64]. DREB TFs are highly inducible by abiotic stresses. DREB1 TF was shown to function under low-temperature stress while DREB2 functions in response to dehydration[78]. A majority of DREBs in tomato seedlings was found upregulated under drought stress, indicating DREBs can serve as potential targets for developing drought resistant crops[79,80].
WRKY
-
The WRKY gene family is a class of plant-specific TFs. Various WRKY family members have been identified from horticultural plants and they were mainly found to be involved in plant responses to biotic and abiotic stresses[81,82]. Tomato WRKY81 can inhibit the transcription of NR (nitrate reductase) and NR expression in tomato leaves. NO accumulation was inhibited in guard cells due to NR suppression, thereby attenuating stomatal closure and drought tolerance[83].
ZFPs
-
Zinc finger proteins (ZFPs) have finger-like structure that can bind to Zn2+. They are characterized into nine subfamilies by their conserved Cys and His motifs. The subfamily of C2H2-type ZFPs are investigated for their roles and functions in abiotic stress responses. SLZF57-silenced tomato plants accumulated more ROS, and the ability of plants to scavenge free radicals and hydrogen peroxide was reduced[84]. ZAT12 is a stress responsive C2H2-type ZFP reported to mediate several stress-activated genes through ROS signaling[85]. In one study, ZAT12 was transferred from Chinese cabbage to tomato and BcZAT12 overexpression in transformed tomato showed stronger membrane integrity, higher water retention and lower H2O2 levels than control plants[85,86]. ZFP proteins can help plants such as tomato to mitigate drought stress and tomato ZFPs such as SlZAT12 should be investigated further as it could be a target for improving tomato's drought tolerance.
-
Many bZIP TFs serve various essential roles in the ABA signaling pathway and abiotic stress response. They also regulate genes related to seed maturation, light, pathogen defense, and flower development. In tomato, SlbZIP1 was found to regulate salt and drought stress tolerance. Silencing of SlbZIP1 decreased the expression of genes involved in ABA biosynthesis and signal transduction. The transcription levels of various abiotic and biotic stress-related genes were downregulated in SlbZIP1-RNAi, suggesting SlbZIP1 can mediate abiotic stress tolerance[87]. In contrast, SlbZIP38 was reported to have opposite functions of SlbZIP1. SlbZIP38 inhibits tomato's drought and salt resistance through regulating the ABA signaling pathway[88].
-
In response to drought, plants can activate their response mechanisms ranging from mediating morphological/structural changes, osmosis regulatory components, and hormone synthesis. These processes are complex and many elements are required to drive drought stress response in plants. The predominant phytohormone involved in the drought stress response is ABA. ABA accumulates during water deficient conditions[16]. Increasing evidence suggests extensive crosstalk presents among ABA and other phytohormones that can regulate plant's drought tolerance. The synergism and functional interaction between ABA and other phytohormones during drought lead to transcriptional reprogramming and downstream responses that help plants cope with stress (Fig. 1). Thus far, we observed that the molecular mechanism underlying tomato's drought response shares some similarities with Arabidopsis and other plants. However, there are other potential routes that tomato can take to trigger drought response and enhance its drought tolerance.
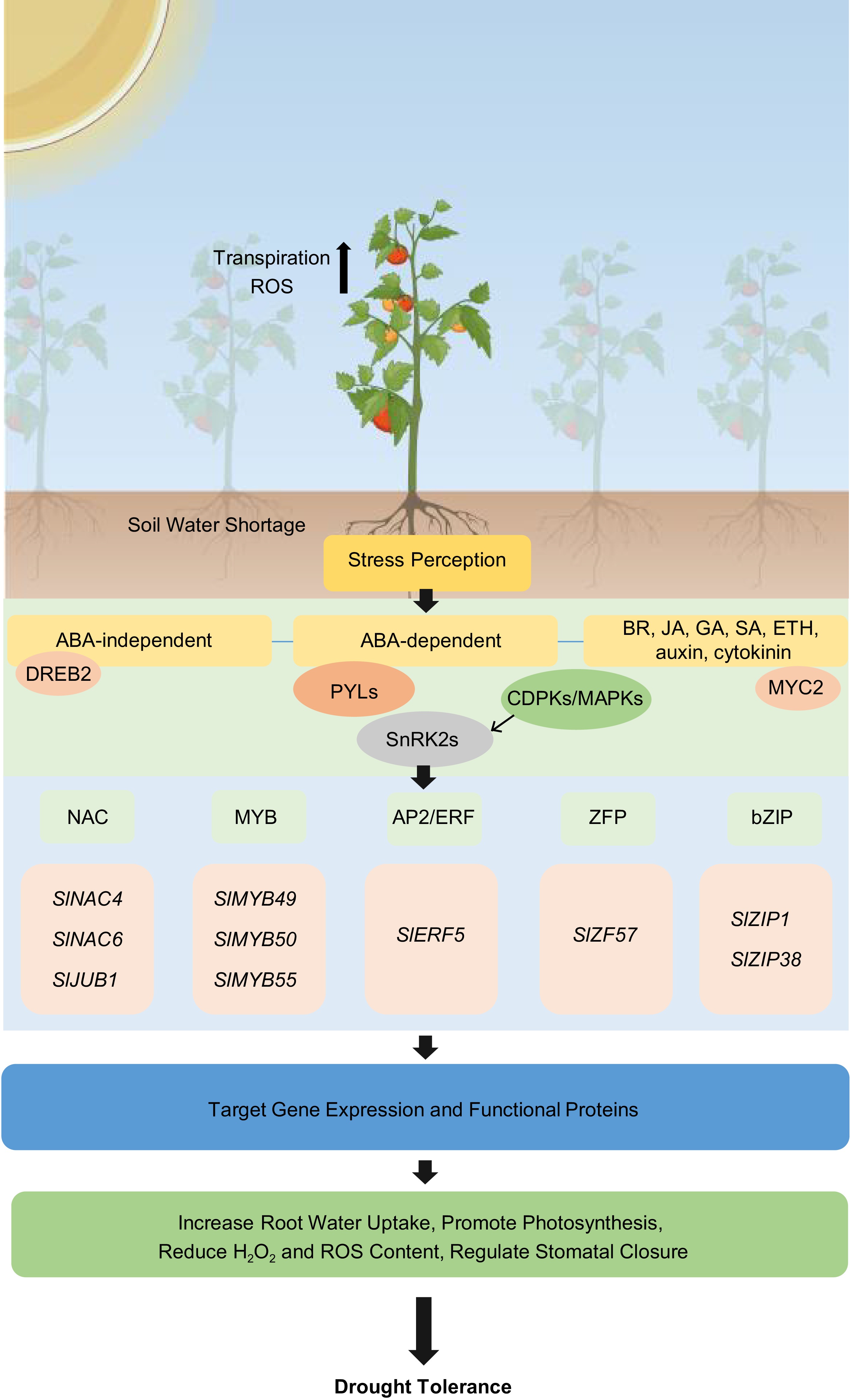
Figure 1.
Drought stress signals, signal transduction pathways including ABA-independent, ABA-dependent, and phytohormone crosstalk play a major role in regulating drought stress response mechanisms in tomato. These pathways involve the modulation of protein kinases, transcription factors, ion channels, aquaporins, and oxidative species to help tomato plants survive under drought conditions.
In Arabidopsis, COST1 (constitutively stressed 1) was found to negatively regulate drought resistance through affecting the autophagy pathway. Similar to SnRK2.6/OST1, COST1 can balance between growth and stress responses in plants, revealing a potential target for improving tomato's drought resistance[89,90]. In addition to COST, other gene members of Arabidopsis such as CLE peptides can be investigated for their functions in tomato's drought tolerance. Arabidopsis CLEs have been found to play a role between water deficiency and ABA synthesis[18,91]. In response to dehydration stress, the expression of AtCLE25 was found in vascular tissues and augmented in roots to form polypeptides. AtCLE25 gene can further induce AtNCED3 (NINE-CIS-EPOXYCAROTENOID DIOXYGENASE 3), an enzyme needed for ABA biosynthesis during drought[18].
The functions of CDPK (calcium-dependent protein kinases) in tomato's drought tolerance should be explored in-depth. CDPK is a crucial calcium sensor that conveys calcium signals into physiological changes. Once ABA synthesis is activated in response to drought, CDPK can phosphorylate various substrates including AQPs, ion channels, metabolic enzymes and TFs to maintain intracellular water balance[6,92]. Whole genome sequence analysis and mRNA expression data from a variety of potatoes revealed that StCDPK21/22 and StCDPK3 can regulate MDA (malondialdehyde) and proline content under drought stress[93,94]. However, the exact role of CDPKs in tomatoes has not been investigated.
Moreover, studies have shown that miRNAs, lncRNAs (long non-coding RNAs) and phytohormones are integrated to regulate environmental stress in plants such as tomato[95−99]. This is still a relatively new area and requires more studies for validation. Plant's drought resistance is complex and regulated by multiple genes, application of '-omics' approaches such as transcriptomics, proteomics, epigenomics, interactomics, metabolomics and phenomics remain essential for a thorough understanding of the regulatory and signaling networks involved in drought stress response in tomato[100].
-
The authors confirm contribution to the paper as follows: study conception and design: Zhu Y, Chong L; data collection: Xie G, Xu R; draft manuscript preparation: Xie G, Xu R, Chong L, Zhu Y. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article.
This work was supported by the National Natural Science Foundation of China (NSFC 32350610245, 32150410345, 32070307 and 32270308), and Scientific and Technological Innovation Talents in Colleges and Universities in Henan, China (23HASTIT036).
-
The authors declare that they have no conflict of interest. Yingfang Zhu is the Editorial Board member of Vegetable Research who was blinded from reviewing or making decisions on the manuscript. The article was subject to the journal's standard procedures, with peer-review handled independently of this Editorial Board member and the research groups.
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Xie G, Xu R, Chong L, Zhu Y. 2024. Understanding drought stress response mechanisms in tomato. Vegetable Research 4: e001 doi: 10.48130/vegres-0023-0033
Understanding drought stress response mechanisms in tomato
- Received: 20 October 2023
- Revised: 08 December 2023
- Accepted: 10 December 2023
- Published online: 09 January 2024
Abstract: Tomato (Solanum lycopersicum) is significantly affected by drought. An excessive amount of reactive oxidative species (ROS) is generated in tomato under drought stress, causing devastating effects on tomato growth and yield. Drought stress response and adaptation in plants are complex and their regulatory gene networks are being investigated to improve our understanding. The signaling molecule, ABA (abscisic acid), is one of the key players in drought stress response. Plants synthesize and accumulate ABA because ABA can act on membrane transporters, protein kinases, transcription factors, and other hormonal signaling pathways to elicit appropriate stress response. In this review, we will discuss our current knowledge of how tomato plants perform drought stress response. Moreover, we will offer future directions that can extend our understanding of tomato's adaptation and tolerance to drought stress.
-
Key words:
- Tomato /
- Drought /
- Protein kinase /
- Transcription factor /
- Phytohormone /
- Abiotic stress












