-
Rice (Oryza sativa L.) is an important staple grain cultivated globally. It serves as a primary host to numerous pathogenic microorganisms. Sheath blight (ShB) is one of the three major diseases of rice, caused by the infection of Rhizoctonia solani Kühn, leading to a substantial reduction in crop yields[1]. Rice ShB exhibits a broad distribution, frequent occurrence, high harm, and a soil-borne nature. Consequently, it has become a global threat to the safety and productivity of rice cultivation[2].
Since the Green Revolution, dwarf and semi-dwarf crop varieties exhibit low sensitivity to nitrogen fertilizers, resulting in the widespread application of nitrogen-based fertilizers[3]. Nitrogen is the most critical mineral nutrient for plants. In agricultural production, the consistent utilization of chemical fertilizers, particularly nitrogen-based fertilizers, has significantly contributed to a substantial increase in grain yields. But it has also resulted in adverse effects on the ecosystem[4,5]. The excessive application of nitrogen fertilizers further leads to air pollution, eutrophication of surface water, and soil acidification, causing serious environmental damage. The molecular mechanism underlying the relationship between nitrogen application and ShB resistance is still largely unknown.
Ammonium is the main form of nitrogen present within rice fields and is also the preferred source of nitrogen absorption by rice. Ammonium transporter proteins (AMT/MEP/Rh protein superfamily) represent the main pathway through which plant roots uptake NH4+. Notably, these proteins exhibit high homology among a wide array of organisms, including plants, bacteria, fungi, and humans[6].
In the early stage, AMT1;1 OX and AMT1;1 RNAi construct were generated using Dongjin (DJ) cultivar as the genetic background. Quantitative reverse transcriptional PCR (RT-qPCR) analysis revealed an upregulation of AMT1;1 in AMT1;1 OX and a downregulation in AMT1;1 RNAi plants (Fig. 1a). Field testing showed that the growth of AMT1;1 OX resembled that of the wildtype (WT), whereas AMT1;1 RNAi plants displayed substantial chlorosis and a reduced number of tillers (Fig. 1b, c). Furthermore, AMT1;1 RNAi showed pale green leaf colors with slender leaf blades and slower rate of leaf developmental (Fig. 1d). It has been previously reported that chlorophyll levels positively regulate the resistance of rice to ShB[7]. Upon inoculation of R. solani AG1-IA, it was observed that AMT1;1 OX plants displayed low susceptibility, while AMT1;1 RNAi plants exhibited high susceptibility to ShB (Fig.1e, f).
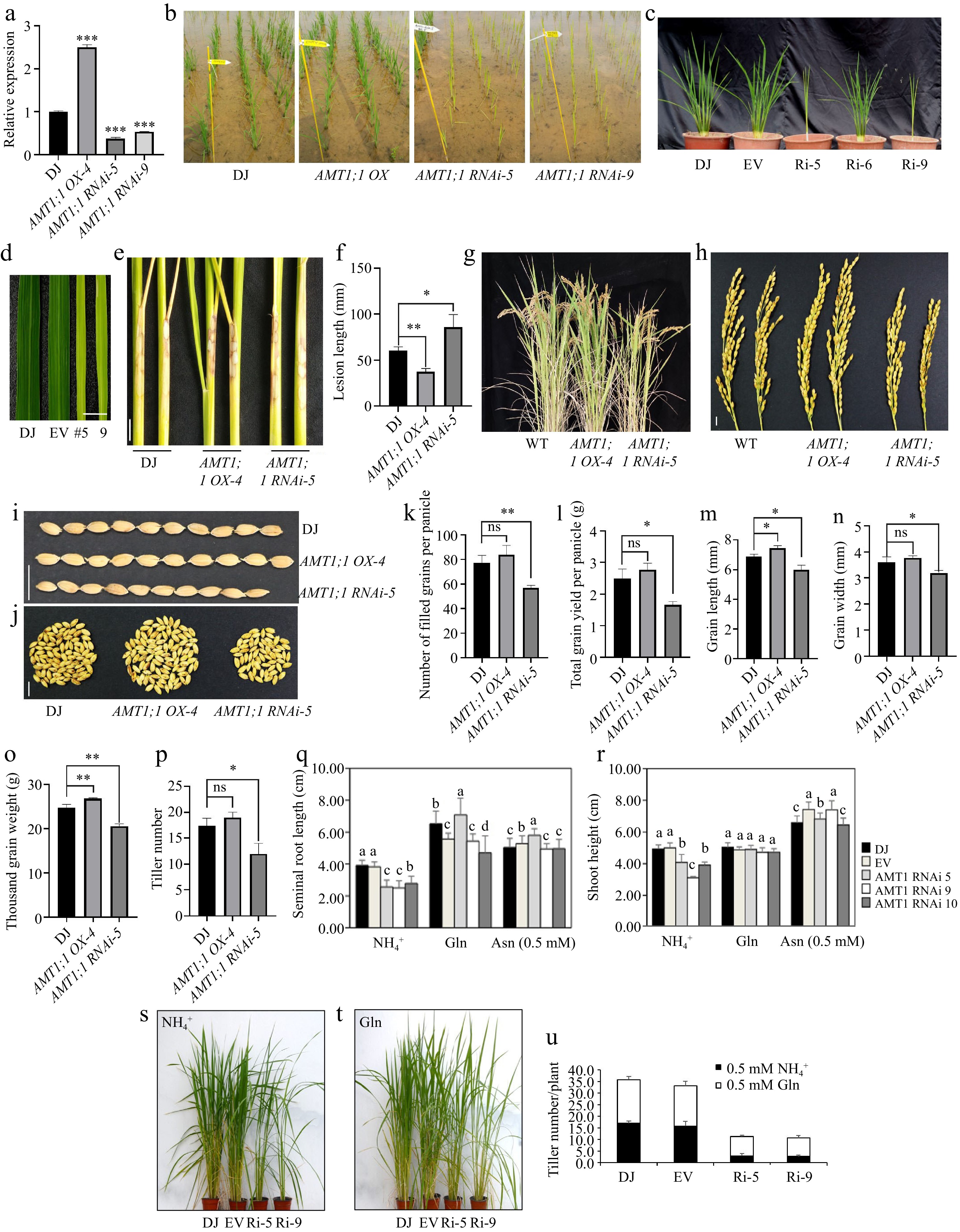
Figure 1.
Characterization of AMT1; 1 OX and AMT1;1 RNAi plant morphology and AMT1;1 OX was less susceptible to ShB and without yield losses.
(a) The expression level of AMT1;1 in WT (DJ), AMT1;1 OX and AMT1;1 RNAi. Sample mRNA levels were normalized to those of Ubiquitin mRNA. Error bars means ± SE. Significant differences are indicated by asterisks. (b) DJ, AMT1;1 OX and AMT1;1 RNAi field grown plant morphology. (c) Characterization of AMT1;1 RNAi mutant, field phenotype of knockdown mutant with its corresponding wild type of 48 d after transplanting to GNU GMO paddy field. EV: Empty Vector. (d) AMT1;1 RNAi shows a thin leaf with a pale green color. Scale bar = 1 cm. (e) AMT1; 1 OX and AMT1;1 RNAi were inoculated with R. solani AG1-IA for 7 d. Scale bar = 2 cm. (f) Measurement of lesion lengths from the sheaths shown in (e). Data means ± standard error (SE). (g) Picture of 3-month-old WT, AMT1;1 OX and AMT1;1 RNAi. (h) Panicle morphology of WT, AMT1;1 OX and AMT1;1 RNAi. Scale bar = 1 cm. (i) Phenotypes of the WT, AMT1;1 OX and AMT1;1 RNAi grains. Scale bar = 1 cm. (j) Yield per panicle of the WT, AMT1;1 OX and AMT1;1 RNAi. Scale bar = 1 cm. (k) Number of filled grains per panicle. (l) Total grain yield per panicle. (m) Grain length. (n) Grain width. (o) Thousand-grain weight. (p) Tiller number of the WT, AMT1;1 OX and AMT1;1 RNAi. (q) Seminal root length. (r) Shoot height. Uniformly germinated seeds grown hydroponically in ¼ nutrient solution containing 0.5 mM NH4+, 0.5 mM Glutamine (Gln), and 0.5 mM Asparagine (Asn) for 14 d in (q) and (r). Significant differences among more than two groups at p < 0.05 are marked with different letters. Uniformly germinated seeds grown in nitrogen-free soil with ¼ nutrient solution containing (s) 0.5 mM NH4+, (t) 0.5 mM Gln for 75 d after germination. (u) Tiller number for (s) and (t). The experiments were repeated at least three times. Significant differences are indicated by asterisks (* p < 0.05, ** p < 0.01, *** p < 0.001). ns: no significant differences.Next, various agronomic traits, including grain morphology, the number of filled grains per panicle, total grain yield per panicle, grain length, grain width, thousand-grain weight, and tiller number, were evaluated for both AMT1;1 OX and AMT1;1 RNAi plants. The results showed that there was no significant difference in most agronomic traits between the WT and AMT1;1 OX. The grain length and thousand-grain weight in AMT1;1 OX were significantly higher than those in WT (Fig. 1g−p). In contrast, the yield of AMT1;1 RNAi was significantly reduced compared to that of WT. These results suggested that AMT1;1 OX exhibited a lower susceptibility to ShB and did not incur any yield losses.
Nitrogen is the most important and abundant element in the growth and development of plants, exerting a pivotal role in various plant life processes. Plants possess the ability to directly assimilate both organic nitrogen sources, such as amino acids and small molecule peptides, and inorganic nitrogen forms, including ammonium nitrogen and nitrate nitrogen from the soil[8]. The growth and development of plants require a large amount of nitrogen sources. Amino acids are the main modes of nitrogen transport in most plants, serving as precursors for protein synthesis, participating in nitrogen metabolism, affecting cell structure, and enhancing plant resistance to biotic and abiotic stresses. Most plants absorb inorganic nitrogen from the soil, but in certain environmental conditions, the direct uptake and utilization of amino acids by plants also play a crucial role[8]. WT (DJ), plants transformed with Empty vector (EV), and those with AMT1;1 RNAi were treated with 0.5 mM NH4+, 0.5 mM Glutamine (Gln), and 0.5 mM Asparagine (Asn) for 14 d. Subsequently, the seminal root length and shoot height were measured. Following the NH4+ there was no significant difference in the seminal root length and shoot height between DJ and EV plants, whereas AMT1;1 RNAi-5 plants exhibited significantly shorter seminal root lengths and shoot heights compared to those of WT and EV. With the application of 0.5 mM Gln, the seminal root length of AMT1;1 RNAi-5 plants was similar to that of WT and EV, and there was no significant difference in shoot height between AMT1;1 RNAi plants and WT. Similarly, the seminal root length and shoot height of AMT1;1 RNAi plants treated with 0.5 mM Asn displayed no significant differences compared to those of wild type (WT) and were sometimes even longer than those of WT (Fig 1q, r). AMT1;1 RNAi plants were treated with 0.5 mM NH4+ and 0.5 mM Gln. The results showed that the tiller number of AMT1;1 RNAi treated with 0.5 mM NH4+ was significantly lower than that of WT and EV. Conversely, the tiller number of AMT1;1 RNAi increased significantly in plants treated with 0.5 mM Gln (Fig. 1s−u). These results suggested that amino acids play a direct role in supporting the growth and development in the AMT1;1 RNAi line, and to some extent, amino acids can partially salvage the NH4+-induced phenotype.
The relationship between plant resistance and growth is typically characterized by mutual antagonism, where the activation of the immune system and the improvement of disease resistance often lead to delay of plant growth and potential yield reduction. Nevertheless, if plants suppress their immune system activation to support rapid growth, they will become susceptible to various pathogens, which can lead to plant death or severe yield losses. Therefore, exploration and identification of functional genes that can achieve both high yield and robust disease resistance have emerged as a prominent focus in the field of molecular improvement breeding in this modern era. Nitrogen, as an essential macromolecular element for normal plant growth and development, plays an important role in the interaction between plants and pathogens. The excessive use of nitrogen fertilizers has introduced intricate effects on this interaction between plants and pathogens, influencing both the growth of pathogens as well as the initiation and activation of plant defense mechanisms[9,10]. The present research uncovered that AMT1;1 OX improves resistance to ShB without compromising yield, which may be explained by the reason that AMT1;1 OX can promote root development and nutrient absorption in rice, ultimately improving photosynthesis efficiency, and thereby increase yield. However, it's important to note that overexpressing AMT1;1 could potentially result in poor growth or other adverse effects. Therefore, using AMT1;1 to improve the agronomic traits and resistance of rice demands a thorough consideration of various factors, including environmental factors and the regulation of other genes, and conducting comprehensive optimization to achieve better disease resistance and yield formation. These findings provide a significant theoretical and practical basis for the future development of rice breeding and crop production. Comprehensive optimization aimed at achieving both enhanced disease resistance and increase yield formation.
HTML
-
The authors confirm contribution to the paper as follows: validation: Li D, Xuan Y; conceptualization, draft manuscript preparation: Li D; funding acquisition, supervision, resource: Xuan Y; manuscript review & editing: Kumar V, Xuan Y. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article.
This work was supported by grants from the National Natural Science Foundation of China (31571985 and 31330063). We extend our gratitude to Prof. Chuang Wang from Huazhong Agricultural University for providing the pt8-2 deletion mutant seeds. Additionally, we are grateful to Prof. Chengcai Chu from the Chinese Academy of Sciences for providing the mutant nrt1.1b seeds.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press on behalf of Yunnan Agricultural University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
| Li D, Kumar V, Xuan Y. 2024. Ammonium transporter 1 promotes resistance to sheath blight in rice without yield loss. Agrobiodiversity 1(1): 13−15 doi: 10.48130/abd-0024-0002 |












