-
Modern agriculture's dependence on chemical herbicides has substantially increased crop yields, but at the cost of persistent soil contamination, which undermines microbial diversity, nutrient cycling, and agroecosystem resilience[1,2]. Commonly used herbicides, such as atrazine, metolachlor, and fomesafen are widely applied in maize, soybean, and cereal production systems and are consistently ranked among the most persistent contaminants, with detection frequencies exceeding 70% in agricultural soils worldwide and reported half-lives ranging from 60 to 200 d[3,4]. Owing to their physicochemical properties, including strong sorption, long environmental persistence and high mobility, these compounds often persist in soil and readily leach into groundwater[5,6], affecting soil function. For example, soil nitrogen cycling is inhibited by atrazine, which reduces nitrification and denitrification by nearly 40%, leading to increased N2O emissions[7]; arbuscular mycorrhizal fungi were suppressed by metolachlor, impairing plant phosphorus uptake and symbiotic interactions[8,9]; and ligninolytic enzyme function was disrupted by fomesafen, inhibiting carbon decomposition and further favoring pathogens over decomposers[4,10]. Together, this cascading connection between herbicides leads to increased agroecosystem yield variability (up to a 15%–20% reduction in contaminated fields) and increasing climate sensitivity. Conventional mitigation methods, such as phytoremediation or activated carbon amendment, provide minimal to no relief, as upfront high costs and additional negative environmental effects outweigh the gains of using the latter method[11,12].
Over the past two decades, single-strain bioaugmentation (e.g., Pseudomonas sp., strain ADP for atrazine or various Arthrobacter and Sphingobium spp. for s-triazine and chloroacetanilide herbicides) has achieved 50%–80% degradation in 14–28 d under laboratory conditions, yet field performance typically drops below 20% due to narrow substrate range, toxic intermediate accumulation and poor resilience to abiotic stresses and microbial competition[13]. Due to these limitations, the attention of various scientists has shifted from natural microbial communities to synthetic microbial communities. The scientists at various labs are putting their efforts into developing consortia that divide metabolic labor across complementary strains while maintaining ecological stability.
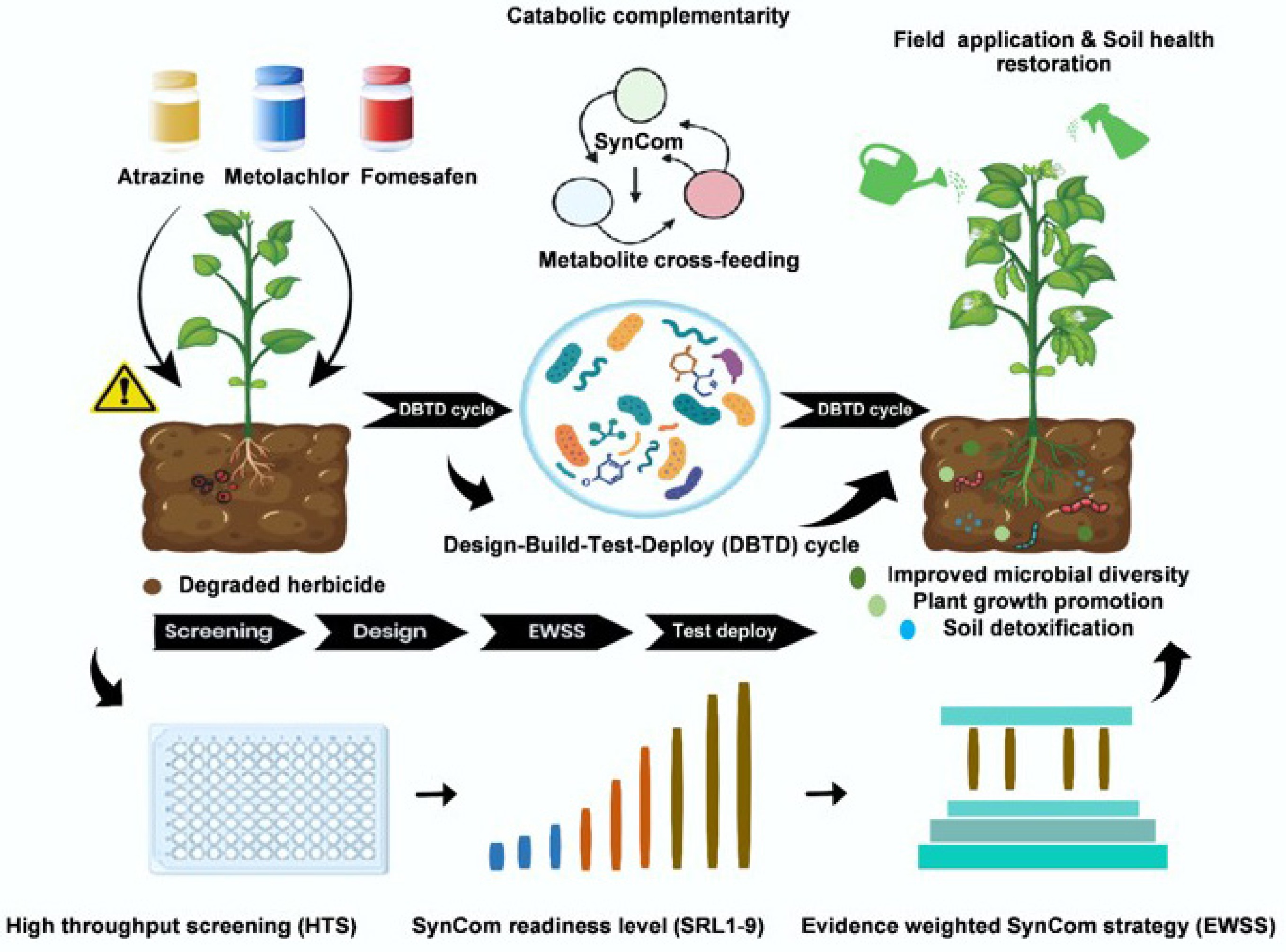
The aforementioned microbial pathways can now be divided at the sub-pathway level. These tools and algorithms, combined with community-level design tools, build the foundation for a Design–Build–Test–Deploy (DBTD) paradigm in SynCom engineering[13,14]. However, there still exist great challenges in taking SynComs optimized in the laboratory to the field that may contain variable abiotic characteristics, stochastic ecological developments, and biosafety issues[15]. To bridge this knowledge gap, we propose two frameworks: (1) The EWSS, which links herbicide chemical structure, known degradation pathways, and metabolic bottlenecks to systematically identify required catabolic and supportive traits (e.g., specific degradative enzymes, intermediate detoxification capacity, and stress resistance). Traits are weighted according to the strength of empirical evidence from literature and experimental data, enabling prioritization of microbial strains or consortia members during assembly. (2) The SynCom Readiness Level (SRL1–9) framework is an adapted maturity assessment analogous to technology readiness levels: SRL1–3 covers conceptual design and initial strain screening; SRL4–5 involves laboratory validation of degradation performance; SRL6 confirms stable function in non-sterile soil microcosms; and SRL7–9 addresses field deployment, long-term persistence, and regulatory considerations. Together, EWSS and SRL support an iterative DBTD cycle, aiming to increase the predictability and reliability of SynCom application for herbicide bioremediation. For example, atrazine degradation time 90% decreased from 41.20 ± 1.95 d (natural attenuation) to 15.48 ± 0.79 d (bioaugmentation), i.e., ~2.7× faster in soil mesocosms[16]. Similar soil studies report rapid atrazine removal after inoculation (e.g., 95.9% removal within 7 d at 5 mg kg–1) and near-complete removal (> 99% by 8 d) in soil microcosm experiments[17]. Given the high mobility of atrazine observed in soil columns (a large fraction recovered in leachate), faster in-soil dissipation is expected to reduce leaching potential, though the magnitude of groundwater-risk reduction must be quantified with site-specific fate/transport modeling and field validation[18]. Herein, we synthesize 2019–2025 advances in SynCom mechanisms and engineering, applying EWSS and SRL to dissect case studies, forecast barriers and chart a roadmap for resilient, herbicide-free farming systems.
-
The complete mineralization of certain herbicides in soils requires multi-step catabolic pathways that are unlikely to be encoded within a single organism due to structural differences between compounds (Fig. 1) (e.g., s-triazine ring in atrazine, chloroacetanilide in metolachlor, and diphenyl ether with nitro/trifluoromethyl groups in fomesafen)[19,20].
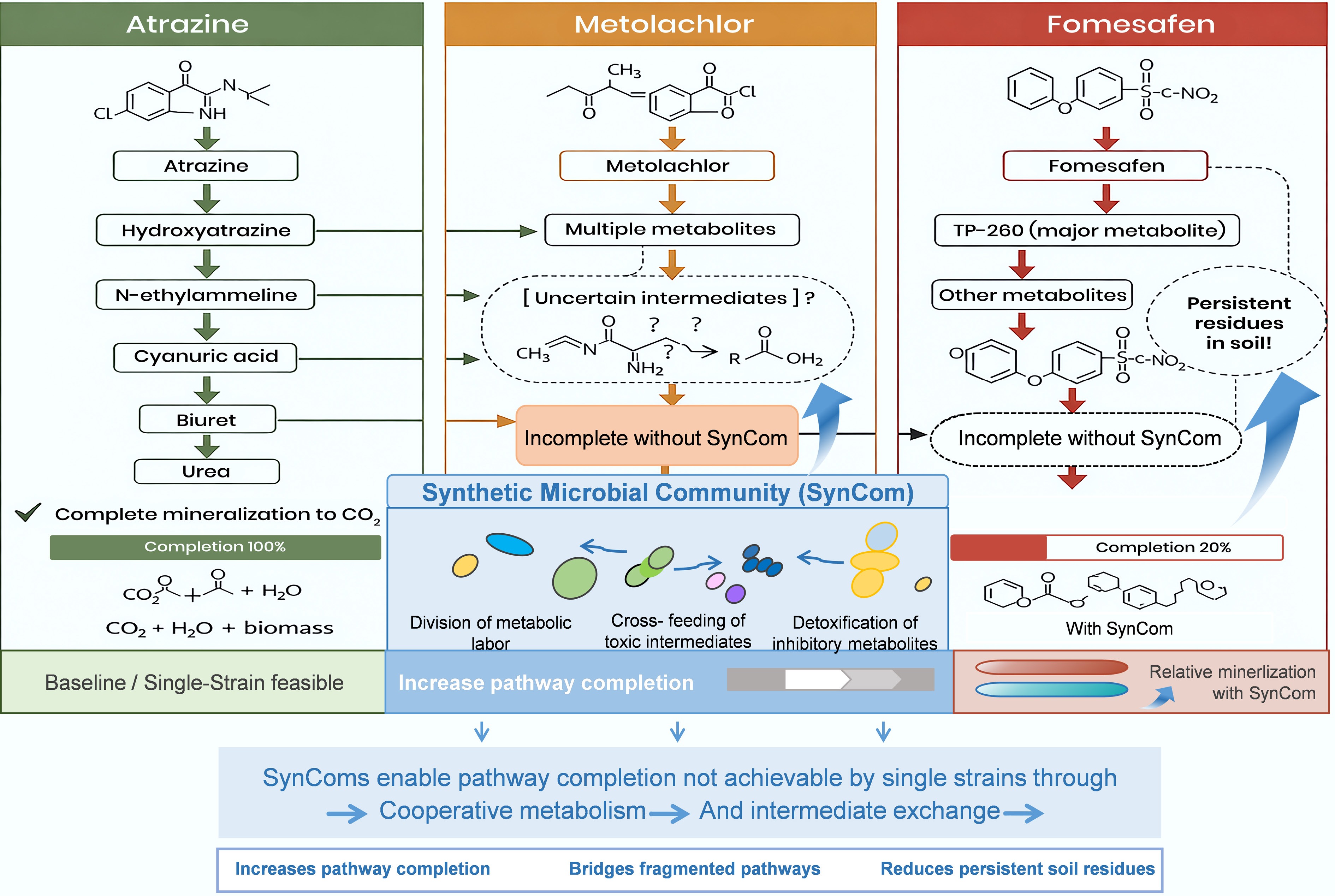
Figure 1.
Synthetic microbial communities enhance the completeness and robustness of soil herbicide degradation.
There may be a capability for natural attenuation by indigenous microbiota; however, the catabolic pathway may be incomplete, intermediates may accumulate and inhibit further degradation, or the population may not be well-suited to win (or survive sufficiently over many passages) in that specific soil and all its environmental variability[21]. To circumvent these disadvantages, SynComs are able to build multi-strain consortia rationally designed with organismal catabolic capabilities of encoding entire catabolism modules, while even 'stabilizing' or 'binding' themselves and reaching a broad niche overlap for occupancy through processes analogous to what indeed happens in natural microbiomes: functional redundancy, division of labor, metabolic cross-feeding, and niche partitioning[22,23].
SynCom allows for resilient and effective herbicide mineralization in more varied field conditions than simpler approaches such as single-strain bioaugmentation[24]. The effectiveness of SynCom comes from interleaved mechanisms.
Catabolic complementarities (division of labor)
-
Different consortium members perform discrete pathway steps, completing cascades unattainable by monocultures. For atrazine, upper-pathway genes (e.g., trzN and atzA for dichlorination, atzB and atzC for dealkylation) predominate in Arthrobacter or Pseudomonas spp., yielding cyanuric acid, while lower-pathway ring cleavage (atzD–atzF and trzD) requires complementary taxa; no single isolate typically mineralizes fully[25,26]. Metolachlor degradation involves glutathione S-transferase conjugation (yielding polar metabolites) followed by dechlorination/oxidation, with enzymes (e.g., amidases, cytochrome P450s) distributed across genera[27,28]. Fomesafen requires nitro reduction (e.g., nitro reductases in Bacillus spp.) and subsequent amide hydrolysis or oxidative cleavage (e.g., cytochrome P450 in fungi like Fusarium)[29] (Table 1).
This distribution of catabolic functions across community members prevents metabolic bottlenecks at any single step, reduces toxic intermediate buildup and enhances robustness through functional redundancy and complementary capabilities.
Metabolic cross-feeding
-
Microbial communities in nutrient-limited soils often achieve herbicide degradation through metabolic cross-feeding, where members exchange pathway intermediates and thereby create interdependence that both detoxifies inhibitory compounds and sustains metabolic flux. For atrazine, this process is well supported by genetic, biochemical, and cultivation-based evidence: upper pathway strains (e.g., carrying atzA, atzB, and atzC genes) convert atrazine and release cyanuric acid (or intermediates like hydroxyatrazine), which downstream members mineralize as a C/N source, reducing inhibition and enabling continued turnover[30,31]. Similar cooperative catabolism has been demonstrated in enrichment cultures, where labour division improves overall degradation rates and coexistence[23,28]. In contrast, metolachlor transformation products such as ethane sulfonic acid (MESA) and oxanilic acid (MOXA) accumulate in soils and groundwater and while these metabolites could theoretically serve as substrates for complementary microbial guilds, direct evidence of cross-feeding (e.g., via isotope tracing or defined consortia) remains limited[32]. Similarly, for fomesafen, nitro reduction products have been proposed as potential substrates that could support interkingdom (bacterial–fungal) cross-feeding, but this remains hypothetical without specific metabolite tracking or enzymatic data in mixed cultures[4,23]. Through such exchanges, cross-feeding can accelerate mineralization and prevent toxic intermediate accumulation by allowing downstream community members to utilize partially processed compounds effectively closing catabolic loops. Cooperative metabolism may also promote long-term persistence of degrading communities by distributing metabolic costs, enhancing overall fitness, and providing resilience to fluctuating soil conditions and episodic herbicide inputs[33,34].
Bacterial–bacterial cross-feeding is well documented in herbicide degrading consortia (e.g., interspecies transfer of atrazine intermediates such as hydroxyatrazine and cyanuric acid); however, interkingdom interactions, for example, between bacteria and fungi or protists, remain comparatively underexplored in herbicide degradation contexts. Some evidence for general microbial ecology suggests fungi may aid bacterial dispersal via hyphal networks (fungal highways) and/or provide complementary enzymatic capacity (e.g., laccases or other oxidoreductases), but specific metabolic exchanges, metabolite transfer, or enzymatic mechanisms driving interkingdom cross-feeding in herbicide systems are poorly defined and lack direct empirical support (e.g., no targeted isotope tracing or perturbation experiments in the cited works)[35]. Future work should explicitly test these interactions (e.g., via multi-omics, isotope tracing and compartmentalized co-cultures or defined SynComs) to determine their contribution to SynCom performance and overall bioremediation efficiency[30].
Recent advances using stable isotope probing (SIP) and single-cell imaging techniques provide direct evidence for metabolite exchange and cross-feeding within microbial communities. For example, quantitative SIP (qSIP) combined with co-occurrence network analysis has demonstrated direct carbon transfer between fungi and bacteria in soil systems, confirming active interkingdom metabolic coupling under natural conditions[36]. In parallel, NanoSIMS-based single-cell imaging approaches have enabled spatially resolved visualization of isotope incorporation, revealing that microbial cells can assimilate contaminant-derived carbon at the individual-cell level and thereby directly participate in community-level metabolic fluxes[37]. Methodological advances integrating Raman micro spectroscopy with NanoSIMS further confirms the robustness of isotope incorporation measurements at single-cell resolution, strengthening the quantitative interpretation of cross-feeding interactions in complex microbiomes[38]. Collectively, these approaches provide direct mechanistic validation of cross-feeding as a key driver of enhanced mineralization efficiency in synthetic microbial communities.
Ecological stress buffering and niche stabilization
-
Non-catabolic 'helper' strains can mitigate abiotic stressors (e.g., pH shifts, moisture fluctuations, oxygen gradients and oxidative stress from herbicides) through the production of extracellular polymeric substances (EPS) for desiccation/herbicide protection, antioxidant enzymes (e.g., catalase, superoxide dismutase), biofilm formation for adhesion and resource retention and nutrient mobilization (e.g., siderophores, organic acids)[39,40]. Cooperative traits like quorum sensing may further enhance population stability in heterogeneous microhabitats, where single degraders often fail[41]. However, while these mechanisms are supported in general microbial ecology and biofilm studies, their specific roles in herbicide-focused SynComs (e.g., direct buffering of atrazine or metolachlor induced stress) are largely extrapolated and would benefit from targeted experiments in relevant systems.
Integration into the DBTD framework
-
These mechanisms inform iterative SynCom engineering: design selects strains for complementary catabolism and helper functions; Build optimizes ratios/promiscuity; Test evaluates flux, detoxification, and resilience in soil microcosms; and Learn (via multi-omics/modeling) refines predictions for field robustness[35].
Table 1. SynCom-mediated herbicide degradation across experimental scales with SRL classification and quantitative judgment criteria
Herbicide Consortium Scale Key performance Field relevance SRL Judgment basis (quantitative criteria) Ref. Atrazine Paenarthrobacter
sp. AT-5 + native microbiomeSoil microcosm 95.9% removed of 5mg kg–1 atrazine with 7 d Controlled soil mesocosm validation 6 Reproducible mesocosm performance across soil histories [33] Atrazine Enriched 3 strain consortium Soil mesocosm 90% degradation in 6 d in pre-exposed soil; 15 d in soil without prior atrazine history Controlled soil mesocosm validation 6 Near-complete removal; performance reproducible across replicated systems [42] Atrazine Arthrobacter sp.
strain AAC22Soil column/
mesocosmNearly 70% removal after 2 d and > 99% after 8 d; 70.2% leachate recovery in column study Demonstrated relevance to leaching and soil remediation 6–7 Effective bioaugmentation under soil and flow conditions [18] Metolachlor Penicillium oxalicum MET-F-1 + native microbiota Field plot 88.6% degradation of
50 mg L–1 within 384 h
under optimal laboratory conditions; 88.3% degradation on day 7 in
field plotLower field residues vs control 7 Quantitative lab and field [43] Metolachlor Native microbial community Field soil cores Faster dissipation
(non-sterile soil)Confirms biodegradation in situ 7 Demonstrated biodegradation in non-sterile soil; control comparison available [44] Fomesafen Bacillus sp. FE-1 Soil microcosm > 82.9% degradation within 14 h in liquid medium at 0.5, 1 and 10 mg L–1 enhanced degradation observed in soil Soil applicability indicated; field data unavailable 5–6 Strong liquid-phase degradation with supporting soil evidence [29] SRL classification is based on system complexity, reproducibility, and ecological realism. SRL6 indicates stable performance in non-sterile soil microcosms (≥ 14–28 d), while SRL7 includes mesocosm or field validation. -
The outlined restrictions in previous sections, such as incomplete degradation by single strains, accumulation of toxic intermediates, poor colonization in the soil and variable field performance highlight a clear gap between the theoretical potential of SynComs and their real-world efficacy in contaminated agroecosystems. Bridging this gap requires a design-to-deployment DBTD pipeline integrating mechanistic biochemistry (e.g., niche complementarity and community stability) and synthetic biology (e.g., rational assembly and genetic engineering). We highlight opportunities for advancement and introduce two translational frameworks, EWSS and SRL, as practical tools to guide the DBTD process toward robust field-ready technologies capable of accelerating herbicide dissipation while preserving soil health.
High-throughput isolation and multi-omics integration
-
Advances in high-throughput isolation and phenotyping enable rapid screening for degradation tolerance and compatibility[45]. Combined with metagenomics (uncultured degraders), transcriptomics (pathways), proteomics (bottlenecks) and metabolomics (flux), this approach enables the design of coherent consortia. Datasets that classify primary degraders, sinks, and helpers are necessary, which is especially crucial for the herbicides that require degradation pathways involving bacterial–fungal interaction (e.g., fomesafen).
A concrete example of multi-omics guided refinement is provided by recent work on atrazine-degrading consortia[46]. Metagenomic and meta-transcriptomic analyses of herbicide-contaminated soils identified keystone degraders (e.g., Arthrobacter spp.) and revealed co-occurrence networks with cyanuric acid-utilizing strains[27]. These data informed genome-scale metabolic modeling to predict optimal strain ratios that balance intermediate production and consumption[47], while co-occurrence networks guided inclusion of supportive taxa. Iterative testing in such studies has demonstrated improved atrazine mineralization in soil microcosms, illustrating how multi-omics directly refines SynCom design[17,40].
Genome-scale metabolic modeling for rational community design
-
Genome-scale metabolic models (GEMs) provide predictive capability by simulating metabolic fluxes, identifying gaps in individual strains, suggesting optimal community partners, predicting the toxicity and accumulation timing of intermediates and determining beneficial cross-feeding exchanges and member ratios[17,40]. Extensions such as community flux balance analysis and evolutionary algorithms explore assembly potentials in 'good' environments, useful for atrazine atzA–atzF or fomesafen reductive-oxidative steps. More than building, this approach is about thinking behind construction.
Machine learning and AI-based SynCom optimization
-
Artificial intelligence tools are emerging as powerful predictors of SynCom behavior. Machine learning models trained on large-scale omics and environmental datasets can predict key aspects of SynCom performance. They also account for real-world challenges such as the accumulation of toxic intermediates, and the need for distributed catabolic pathways across community members[40,41].
The EWSS and SRL
-
The EWSS is a comprehensive framework designed to evaluate candidate microbial consortia based on seven key environmental characteristics. These characteristics include herbicide class and chemical structure, distribution of catabolic functions, intermediate toxicity and degradation pathways, cross-feeding capacity, contributions from helper organisms (e.g., biofilms, extracellular polymeric substances [EPS] and stress buffering), compatibility with the available ecological niches and resources and resilience to environmental stressors. The framework systematically maps these traits to specific SynCom architectures, moving away from trial-and-error approaches towards evidence-based design that maximizes field performance.
We propose SRL as a scale (1–9) adapted from NASA's Technology Readiness Level (TRL) and Bioindustrial Manufacturing Readiness Levels (BioMRL)[40,41] to track progress for SynCom from lab concept to field deployment[40]. SRL1–3: mechanistic discovery identification of key catabolic genes, pathways, and strains using omics technologies; SRL4–5: in-vitro assembly, build and test the SynCom in controlled lab conditions, assessing flux, stability and degradation efficiency; SRL6–7: microcosm and mesocosm testing validation in sterile and non-sterile soil systems for colonization, robustness, and minimal off-target impact; SRL8–9: pre-deployment and field trials conducted biosafety assessments, regulatory checks and monitored filed releases with quantitative efficacy tracking.
The SRL scale provides a structured framework to guide the DBTD pipeline by setting clear milestones. SynComs reaching SRL ≥ 6 are typically able to establish themselves in soil environments and deliver rapid herbicide degradation without harming native microbial communities[48,49].
It is worth mentioning that spatial and temporal heterogeneity in soil herbicide concentrations plays a critical role in influencing SynCom performance. Herbicide distribution within soil is rarely uniform and concentration gradients, especially between surface and deeper soil layers, creates distinct microenvironments that affect microbial activity. For instance, vertical gradients in oxygen availability, moisture and organic matter content shape degradation kinetics by favoring different microbial populations at various soil depths. Furthermore, soil aggregation and diffusion limitations generate localized concentration hotspots and cold spots, leading to uneven substrate availability for SynCom members. Such spatial heterogeneity can significantly disrupt metabolic interactions within SynComs. In high-concentration zones, primary degraders may dominate due to sufficient substrate availability, while downstream metabolizers that rely on intermediates for degradation may become spatially separated or limited by substrate scarcity in low-concentration areas[3]. This spatial decoupling of metabolic functions can reduce cross-feeding efficiency, promote intermediate accumulation and hinder overall mineralization rates. Temporal fluctuations in herbicide concentration, driven by rainfall events, adsorption–desorption processes and plant uptake further contribute to shifts in herbicide bioavailability, causing SynCom members to experience substrate-limited or inhibitory conditions at different times[34].
Mechanistically, these dynamics underscore the importance of spatial niche partitioning, bioavailability constraints and microbial dispersal traits (e.g., motility, biofilm formation, and attachment to soil particles) in SynCom design. These factors are critical for maintaining metabolic fluxes across diverse soil microhabitats and ensuring continuous herbicide degradation. From an EWSS perspective, integrating traits related to stress tolerance, metabolite diffusion compatibility and ecological resilience is essential under heterogeneous soil conditions. By explicitly accounting for spatiotemporal concentration gradients, SynCom design can more accurately predict field performance and bridge the gap between controlled laboratory systems (SRL4–5) and real-world applications (SRL6–7)[4,24].
EWSS-guided SynCom design across representative herbicides (atrazine, metolachlor, and fomesafen)
-
To demonstrate the general applicability of the EWSS framework across structurally distinct herbicides, a unified design logic can be applied to atrazine, metolachlor and fomesafen by systematically linking molecular structure to required catabolic and supportive traits. Atrazine, a triazine herbicide, undergoes stepwise biodegradation through hydrolytic dechlorination, N-dealkylation and subsequent cyanuric acid ring cleavage, requiring coordinated activity of upper- and lower-pathway degraders[9]. In contrast, metolachlor, a chloroacetanilide herbicide, is initially detoxified via glutathione S-transferase (GST)-mediated conjugation, followed by downstream oxidation, dechlorination, and transformation of key metabolites such as MESA and MOXA[34]. Fomesafen, a diphenyl ether herbicide with nitroaromatic functional groups requires more complex transformation involving nitro-reduction and oxidative cleavage, often necessitating interkingdom cooperation between bacterial and fungal partners[29].
Despite these structural differences, a consistent EWSS-guided workflow emerges: (1) identification of key transformation pathways based on chemical structure; (2) prioritization of functional traits such as specific degradative enzymes, intermediate detoxification capacity, and stress tolerance[34]; (3) selection of complementary microbial taxa enabling division of labor and metabolic handoffs; (4) assembly of SynComs that balance upstream and downstream metabolic fluxes to prevent intermediate accumulation; and (5) validation under soil-relevant conditions to ensure stability, functional persistence, and complete mineralization[29]. Collectively, these examples demonstrate that EWSS provides a generalizable and mechanistically grounded framework for designing robust SynComs across herbicides with diverse chemical structures.
Integration within the DBTD workflow
-
These predictive tools and frameworks are inductively integrated into a workflow that combines classical microbiology with systems biology approaches. This enables iterative refinement of SynCom composition guided by soil-relevant constraints, ultimately converging on robust designs.
Future opportunities
-
Opportunities to exploit these include the use of archaea, fungi, and protists for diversity; delivery systems for persistence; synthetic signaling for responses; isotopic tracing for fate; and microbiome vaults for preservation and activation. The outcome enables SynComs that are resilient in the face of the persistent herbicides used, and simultaneously scale bioremediation, enhancing performance predictability in complex agroecosystems.
-
Typical herbicide residues, including atrazine, metolachlor and fomesafen pose significantly challenges for soil health and agroecosystem functioning. This review highlighits that rationally designed SynComs can overcome key limitations of single-strain bioremediation through metabolic division of labor, cross-feeding of intermediates and enhanced community stability. Evidence from studies published between 2019 and 2025 indicate that SynComs generally achieve more efficient and complete herbicide degradation (approximately 1.5–3-flod) than single strains or natural attenuation, particularly in soil microcosms system. The proposed EWSS and SRL framework provide structured tools to guide predictive design and maturation of consortia from laboratory proof of concept, to field deployment.
Although promising, the majority of current evidence derives from controlled laboratory studies and microcosms. Field-scale validation, including long-term stability, interactions with native microbiomes, and impacts on soil multifunctionality remains a critical priority. Future research should focus on: (1) complete mineralization pathways, especially for metolachlor and fomesafen; (2) models of how SynComs interacts with the environment; (3) high-throughput formulation strategies for field delivery; and (4) regulatory frameworks that facilitate safe deployment. With continued integration of synthetic biology, multi-omics, and ecological principles, SynCom-based bioremediation holds great potential for restoring contaminated agricultural soils and supporting sustainable agriculture.
Challenges, failures, and risks in SynCom deployment
-
Key limitations:
(1) Poor long-term persistence in non-sterile soils due to competition, predation, phages, and abiotic variability.
(2) Community instability (loss of keystone degraders, composition drift, reduced function).
(3) Potential unintended effects (transient nutrient cycling disruption, enrichment of opportunists, network shifts).
(4) Scarcity of published negative results due to bias.
(5) The need for transparent failure reporting and long-term ecological monitoring.
-
The authors confirm their contributions to the paper as follows: Hiba Adil Mahjoob: investigation, data curation, visualization, writing – original draft; Xueling Yang: supervision, investigation, writing – review & editing; Yan He: conceptualization, supervision, funding acquisition, writing – review & editing. All authors reviewed the results and approved the final version of the manuscript.
-
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
-
This research was financially supported by the Zhejiang Provincial Key Research and Development Program of China (Grant No. 2023C02004), the Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (Grant No. JYB2025XDXM703), the National Natural Science Foundation of China (Grant Nos 42177006, 42307018), the Fundamental Research Funds for the Central Universities (Grant No. 226-2025-00064), and the China Agriculture Research System of MOF and MARA (Grant No. CARS-04).
-
The authors declare that they have no conflict of interest.
-
Full list of author information is available at the end of the article.
- Copyright: © 2026 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Mahjoob HA, Yang X, He Y. 2026. Mechanistic insights into designing synthetic microbial communities to accelerate herbicide biodegradation in soils. Agricultural Ecology and Environment 2: e014 doi: 10.48130/aee-0026-0010
Mechanistic insights into designing synthetic microbial communities to accelerate herbicide biodegradation in soils
- Received: 14 January 2026
- Revised: 25 March 2026
- Accepted: 11 April 2026
- Published online: 06 May 2026
Abstract: Persistent herbicide residues, including atrazine, metolachlor, fomesafen, etc., pose significant threats to soil microbial diversity, nutrient cycling and long-term agroecosystem sustainability. Conventional single-strain bioremediation frequently results in incomplete degradation, accumulation of toxic intermediates and limited ecological fitness under field conditions. Synthetic microbial communities (SynComs) provide a promising alternative by harnessing metabolic division of labor, cross-feeding and niche complementarity. This review synthesizes advances from 2019 to 2025, demonstrating that rationally designed SynComs substantially outperform single strains or natural attenuation, with most evidence derived from laboratory and soil microcosm studies conducted in non-sterile agricultural soils (e.g., loamy soils with moderate organic matter content and neutral pH). SynComs generally exhibit enhanced degradation performance compared to single strains, owing to metabolic complementarity and cooperative interactions. Reported improvements vary widely across studies, ranging from 1.5 to 3-fold, in specific optimized cases, depending on the pollutants, community design and experimental conditions. Enhanced performance is attributed to: (1) distribution of catabolic steps across community members, (2) cooperative detoxification of intermediates, and (3) improved resilience via mutualistic interactions. Emerging tools, including high-throughput isolation, multi-omics profiling, genome-scale metabolic modelling and machine-learning guided assembly, enable increasingly predictive SynCom design. To support translation from laboratory optimization to field application, we propose two complementary frameworks: (1) the Evidence-Weighted SynCom Strategy (EWSS), which links herbicide structure to essential catabolic and supportive traits; and (2) the SynCom Readiness Level (SRL1–9), which tracks consortium maturity from mechanistic discovery to field-scale validation. While field-scale validation remains limited, existing mesocosm and field studies consistently demonstrate improved in-situ dissipation relative to natural attenuation.