-
A sustainable increase in agricultural production is needed due to a predicted increase of 100%−110% in demand for crops globally from the year 2005 to 2050[1]. Abiotic stresses can physically damage the plant or weaken the plant's defense and health, thus assisting the entry or attack of pathogens on the plant[2]. Biotic stresses mainly involve infections caused by pathogens, such as viruses, fungi, nematodes, bacteria, and protozoa[3]. Disease in plants is defined as the response of plant cells and tissues to any environmental stress or a pathogen, resulting in abnormality in plant health[4]. Disease endemics in plants occur yearly on various crops in different parts of the world[5]. Phytopathogens have a severe negative impact on the quality and quantity of agricultural products, parallelly a bad effect on the economy of a country. Thus, a sustainable food supply is at risk, due to the rapid propagation of pathogens with an increase in the incidences and severity of infectious diseases[6].
At present, the increasing demand for crop production depends on synthetic agrochemicals to reduce the risks posed by plant diseases and ensure crop yield[7]. The Food and Agriculture Organization of the United Nations (FAO) Statistics database shows that the use of fungicides, bactericides, and pesticides worldwide is increasing rapidly[8]. However, excessive use of agrochemicals without discrimination contributes to global warming and has several adverse effects—such as making pathogens more resistant, causing toxicity to non-target organisms, and posing severe risks to human and environmental health[9,10]. Consequently, innovative plant disease control approaches and advancements in technology need to be established and employed to enhance the effectiveness of plant disease control measurements and minimize environmental damage. Nowadays, antimicrobial nanomaterials are attracting more attention in the scientific community as they can unlock the limitations experienced by other antimicrobial agents and traditional pesticides. Nanomedicine is a new approach to overcoming the challenges of conventional treatments, based on the manufacturing and fabrication of nanoparticulate[11,12]. Numerous types of nanostructures such as metal nanoparticles, nanogels, biodegradable polymeric nanoparticles, nanoliposomes, and solid lipid nanoparticles, have been endeavored as probable drug delivery systems[13,14]. Nanoparticles are generally no larger than 100 nm in size and their smaller size along with their higher surface-to-volume ratio, govern them, as effective biocidal agents, as both these combined effects facilitate the intimate interaction on the microbial membrane[13,15]. The fungicidal ability of biosynthesized metallic nanoparticles was found to be more aggressive than commercially applied antibiotics such as amphotericin and fluconazole. Obvious membrane damage in Candida sp. has been observed after the application of plant-mediated silver nanoparticles, which not only damaged the intercellular components of the fungus but also destroyed the cell functions[16,17]. In antibacterial activity, the negative charge on the bacterial cell wall interacts with the positive charge nanoparticles due to their electrostatic forces resulting in the disruption of the bacterial cell walls. Furthermore, nanoparticles also release metal ions from their extracellular place, which then enter the cell wall and disrupt the normal biological process of bacteria. Inside the bacterial cell, either nanoparticles or metal ions, persuade the ROS level and damage the protoplasm. The generation of oxidative stress leads to the oxidation of glutathione, which results in the destruction of the antioxidative defense mechanism of organisms against ROS. The metal ions are therefore free to interact with cellular systems (e.g., membranes DNA, proteins, etc.), disrupting cellular functions. Metal ions can develop a solid coordination bond with O, S, or N atoms which are present in abundance in biomolecules and organic systems. Meanwhile, the bond between biomolecules and metal ions is generally non-specific, ultimately, metallic nanoparticles exhibit a broad spectrum of potential[18,19].
To synthesize eco-friendly and low-cost metallic nanoparticles, researchers are utilizing the capabilities of biological materials to manufacture metallic nanoparticles[20]. In the green-fabrication of metallic nanoparticles, the reduction of metals involves the biological mass as reducers—either intra-cellularly or extra-cellularly[21]. Apart from eco-friendliness and cost-effectiveness, the advantage of the biological approaches over classical approaches (physical and chemical methods), comprise the efficiency of the technique in catalyzing the reactions in the aqueous environment at standard pressure and temperature conditions, as well as the flexibility of the process by implemented in almost at any scale and any settings[22]. The elements of biological sources are responsible for the reduction and the process may often be triggered by various compounds and constituents such as proteins, terpenoids, flavonoids, alkaloids, terpenoids, phenols, amines, carbonyl, amide groups, different pigments, and other reducing agents. One or more of these reducing agents may be responsible for the synthesis of metallic nanoparticles[23]. Applications of nanomaterials are extended to the fields of both human and plant pathology and biotechnology[24,25]. Based on their unique characteristics, various biogenically synthesized nanoparticles serve as anticancer and antimicrobial agents in various fields[26,27]. CuO nanoparticles have vast unique characteristics from pharmacology to pest ecology for overcoming various pathogenic diseases[28].
Mangifera indica is a famous tree that belongs to the family Anacardiaceae used in traditional medicines, especially for skin problems, cough, diarrhea, malaria, dysentery jaundice, and their anti-microbial properties[29,30]. Thus, due to the antioxidant, antimicrobial, and pharmacological importance of M. indica, it was assumed that employing plant leaves as a reducing and capping agent would enable the formation of CuO.NPs with vital phytochemicals. The main objective of the current study is to synthesize eco-friendly and economically viable M. indica-mediated CuO.NPs to explore their antifungal and antibacterial potential. This current work shows that CuO.NPs synthesized by M. indica are biocompatible, eco-friendly, and environmentally sustainable Nano fungicides against phytopathogens. Therefore, improving the yield of nanoscale metal particles, using low-cost raw materials, and greensource, and employing simple energy-saving technology are the research directions needed in the future. At present, there have been successful cases of using grass to synthesize CuO.NPs. Therefore, green synthesis of nanoscale metals may have a broad prospect and a great potential for development.
-
The chemicals used in this work were Copper (II) sulphate pentahydrate (CuSo4·5H2O), deionized H2O. Escherichia coli, Staphylococcus aureus and Rhizopus oryzae.
Plant material processing
-
The healthy leaves of Mangifera indica were collected from Punjab province, Pakistan, and identified from the well-reputed plant herbarium center of the taxonomy laboratory department at Quaid-i-Azam University (QAU) Islamabad Pakistan. The fresh plant leaves were thoroughly washed with de-ionized water and dried under ambient temperature (1−2 weeks). After plant materials had completely air dried, extracts were pulverized with a blender to a fine powder. Afterward, 30 g of plant powder was added to 300 mL of distilled water and boiled for half an hour at 74 °C. The material was then shifted in an incubator shaker for 2−3 h at 50 rpm at 40 °C. The resultant mixture was then filtered with Whattman No. 1 filter paper. The extract filtered was kept at 4 °C for further synthesis process of CuO.NPs[31].
Preparation of CuO.NPs
-
The formulation of green CuO.NPs were mediated by the co-precipitate method by applying Mangifera indica leaf extract. For the green synthesis of CuO.NPs, 40 mL copper (II) sulphate pentahydrate (CuSo4·5H2O) solution (1 M) was prepared by adding in 10 mL aqueous plant extract and stirring on a hot plate at 80 °C for 4 h at 200 rpm. The brownish color CuO.NPs in precipitate form were collected via 12,000 rpm of centrifugation (Velocity 14R, 220VAC 50/60HZ, 10A China), for 10 min. The resulting pellet was washed thrice with deionized water and dried in a drying oven for 4 h at 60 °C. Finally, the radish-brown NPs were collected, and stored at room temperature for further use[32].
Phytochemicals screening
-
The antioxidant, antifungal, and antibacterial compounds in plant extracts were carried out by manual procedure in a lab using chemicals. The phytochemical screening of phenols and saponins, alkaloids triterpenes, flavonoids, and tannins according to the already well-established protocol[33].
Phenols and tannins test verification
-
To confirm the presence of phenols and tannins in M. indica plant extract solution. A few drops of FeCl3 3% were added drop by drop to a 1 mL extract solution. After some time, deep blue coloration formed which was the first identification of phenolic and tannins.
Test for triterpenes
-
For triterpene compound analysis, 1 mL of chloroform was added into the solutions, and then 1 mL of concentrated H2SO4 was carefully mixed into the test tubes by sliding it down the walls. The confirmation of triterpenes' presence was indicated by the development of red coloration in the solution.
Test for flavonoids
-
Lead acetate solution was added to 1 mL of the extract solutions and the presence of flavonoids was confirmed by the formation of a yellow precipitate in the solution.
Test for alkaloids
-
The alkaloid test was carried out by adding 5 mL of HCl to the extract solution. The solution was agitated, filtered, and kept for further analysis. Meyer's test was then carried out by adding 2 mL of the filtrate to 5 mL of Meyer's reagent. The formation of a yellow precipitate indicates the presence of alkaloids.
Characterizations of CuO.NPs
-
The crystallographic nature and intrinsic characteristics of green synthesized M. indica-mediated NPs were confirmed by XRD analysis (Schimadzu-Model Kyoto, Japan) ranging 2θ from 10° to 90° via Cu/Kɑ radiations with wavelength 1.5406 Å ran at 40 kV with 30 mA at room temperature. The size of synthesized particles was determined using the Debye-Scherer equation (D = K/cos), where D is the crystal size and is vertical to the reflecting planes. K is constant in this equation (0.9), and the variables are the X-ray wavelength (1.5406), the angular full-width at half-maximum in radians, the Bragg's angle, and the angular full-width at half-maximum in radians.
FT-IR analysis for the green synthesized M. indica-mediated CuO.NPs were analyzed to confirm the functional groups of biomolecules in the prepared NPs. The wavelength was set in the range of 400−4,000 cm−1 using an FT-IR spectroscope (Schimadzu-Model FT-IR, Kyoto, Japan) having a resolution of 4 cm−1.
The surface morphology of green synthesized CuO.NPs were assessed by scanning electron microscopy. The drop of metal NP solution sample was loaded on copper-coated stubs and then evaporated at light for full drying. Finally sample was loaded in the transmission electron microscope (JEOL JSM-5910), with 10 KV of accelerating voltage for 4 min[34]. The morphology of synthesized samples was further proven with a high-resolution application of the protocol. The size of the synthesized particles was measured by determining the diameter of the nanoparticles[35,36].
Antibacterial assay
-
The antibacterial potential of synthesized green CuO.NPs were evaluated against E. coli and S. aureus bacteria via the disc diffusion method[37]. The bacterial inoculum was prepared by culturing a single colony of E. coli in nutrient broth at 37 °C for 24 h. Then the prepared fresh inoculum was swabbed evenly on media plates (nutrient broth agar), and 5 mm of wells were created in the plates by using an autoclaved cork borer. A total volume of 20 µL of the sample having various concentrations (100, 50, and 30 µg·mL−1) was added according to the wells. The cefixime was applied as a positive control and DMSO as a negative control. The culture plates were incubated at 37 °C for 24 h and their zones of inhibitions were measured. The experiment was performed three times.
Antifungal assay
-
The antifungal potential of synthesized green CuO.NPs were explored in vivo against R. oryzae. In vivo was accomplished by adding R. oryzae mycelia in Czapek Dox Broth medium. The components of Czapek Dox Broth media were sucrose (7.5 g), potassium chloride (0.25 g), sodium nitrate (0.5 g), magnesium sulphate (0.25 g), dipotassium phosphate (0.25 g), ferrous sulphate (0.005 g), and 250 mL distilled H2O. The media was incubated at 32 ± 2 °C for 24 h at 50 rpm with continuous shaking. The D. kaki was surface sterilized via 50 % sodium hypochlorite suspension before treatment with synthesized CuO.NPs. Then the artificial wounds were created in the center of the fruits by a 2 mm sterile cork borer. In the wound of fruits, 5 µL spore suspension was inoculated into the wounds of each fruit, further treated with 30 µL volume of both concentrations (100 and 60 µg·mL−1) of CuO.NPs. To provide moisture and avoid contamination, the treated fruits were covered with a wet muslin cloth. After a week of post-inoculation, the diameter of the lesion was measured[38,39].
Antioxidant activity
-
The antioxidant activity was assessed using the 2,2-Diphenyl-1-picrylhydrazyl (DPPH) radical scavenging experiment, 100 μL of green synthesized CuO.NPs at a concentration of 1 mg·mL−1 were serially diluted with 100 μL of methanol. The well was filled with 100 μL of 0.1 mM methanolic DPPH, and the plates were then let to sit at room temperature in the dark for 30 min. M. indica extract, CuO.NPs, and ascorbic acid concentrations were evaluated at an ELISA plate reader were used to read the plates at 517 nm. Where A0 and A1 are the absorbances of the control and treatment, respectively, were used to calculate the DPPH inhibitory percentage[34].
${DPPH\; inhibitory}\; {\text{%}} = \dfrac{{\rm A}_0 - {\rm A}_1 }{{\rm A}_0}\times 100 $ Where A0 and A1 are absorbances of control and treated, respectively.
-
In the last few years, nanotechnology has attained great impetus in applied material due to its unique physiochemical properties. Numerous methods for synthesizing metal nanoparticles are currently documented in the literature and employed by researchers in their quest to discover novel and emerging uses for these nanoparticles. Nanoparticles are the upcoming plant pathogen weapons in modern agriculture, and green-produced nanoparticles have shown significant potential for disease prevention[40]. The biological approach encompasses the creation of nanoparticles through the utilization of living organisms, including plants, bacteria, fungi, and algae. When plants are involved in the synthesis of nanoparticles, it is referred to as green nanotechnology[41]. The utilization of plant biomaterials for the synthesis of metallic nanoparticles is now recognized as a highly emerging field. Plants possess inherent organic reducing agents, which render them well-suited and adaptable for nanoparticle synthesis[42]. Numerous plant extracts have been effectively tested for the production of metallic nanoparticles. Extracts from various plants such as Camellia sinensis, Azadirachta indica, Halymenia dilatata, Stachys lavandilfolia, Eucalyptus, and Mentha have been documented as capable of synthesizing diverse types of CuO.NPs, each with distinct and specific applications[38,43,44].
Mangifera indica, a significant plant with numerous medicinal uses, has gained recognition. Phoenix dactylifera has previously been employed for the production of various metallic nanoparticles. To investigate the antimicrobial and antioxidant activity, researchers have utilized leaf extract from M. indica to synthesize silver oxide nanoparticles (AgNPs)[45]. In the present study, M. indica was utilized as a source to generate copper oxide nanoparticles (CuO.NPs). The synthesis of green M. indica-mediated CuO.NPs were confirmed via a color change from yellow to copper-reddish indicating CuO.NPs synthesis. The main objective of green synthesis was the fabrication of cost-effective, environmentally friendly, and biocompatible nanoparticles[46]. The production of green CuO.NPs are a complicated process that depends on various variables due to their colloidal properties. The main objective is to develop a phyto-fabricated approach that can be commercialized without using complex processes such as magnetic filtration, ultracentrifugation, co-precipitation, and flow field gradient. Another objective is to specify the reaction conditions that aid in controlling the size of the particle. Already reported studies confirmed the suitability of the co-precipitation method to synthesize reduced size and homogenous texture. The results of this study align with previous research, demonstrating that this method can meet the increasing need for well-dispersed green CuO.NPs in biomedicine and engineering[47]. In this present work, green synthesized CuO.NPs were characterized through different techniques. After visual confirmation of biomolecules from M. indica plant extract. To confirm the antifungal and antibacterial biological molecules in green source plant extract visual confirmatory tests were performed in the laboratory. The Mangifera indica filtrate solution contained all antioxidant flavonoids necessary for the reduction of green synthesized CuO.NPs (Fig.1).
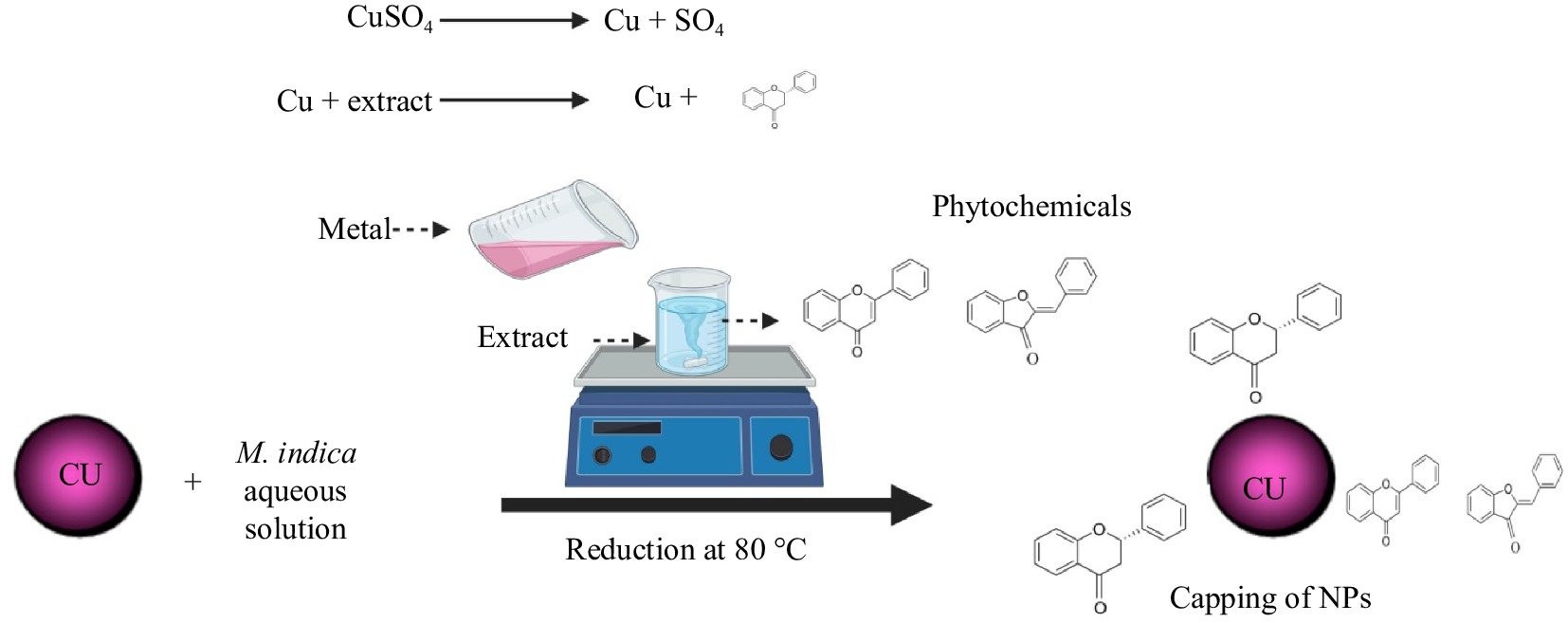
Figure 1.
Proposed hypothetical picture of the reduction mechanism of copper sulphate by the aqueous leaf extract solution of M. indica on a hot plate at 80 °C for 4 h at 200 rpm.
FTIR spectroscopy was carried out to investigate biomolecules responsible for the reduction of Cu nanoparticles. The functional groups act as capping and reducing agents in synthesizing nanoparticles and were analyzed by FT-IR (Fig. 2). FT-IR exhibited absorption peaks 3,404.13, 300.37, 2,382.35, 1,641.84, 1,476.90, 1,072.42, 872.66, and 651.51 cm−1 matching to several functional groups that are C-N stretching aromatic amino groups, C-O carboxylic anions, alcohol O-H stretching, and amine N-H stretching groups respectively, while peaks for CuO.NPs were instituted nearby 550–600 cm−1. The fabricated CuO.NPs were covered by proteins and various metabolites occupying functional groups. The FT-IR result confirmed that the proteins and amino acids residues built a strong connection to attract the metals and prevent clusters due to the capping of CuO.NPs and stabilizing the respective medium (Table 1).
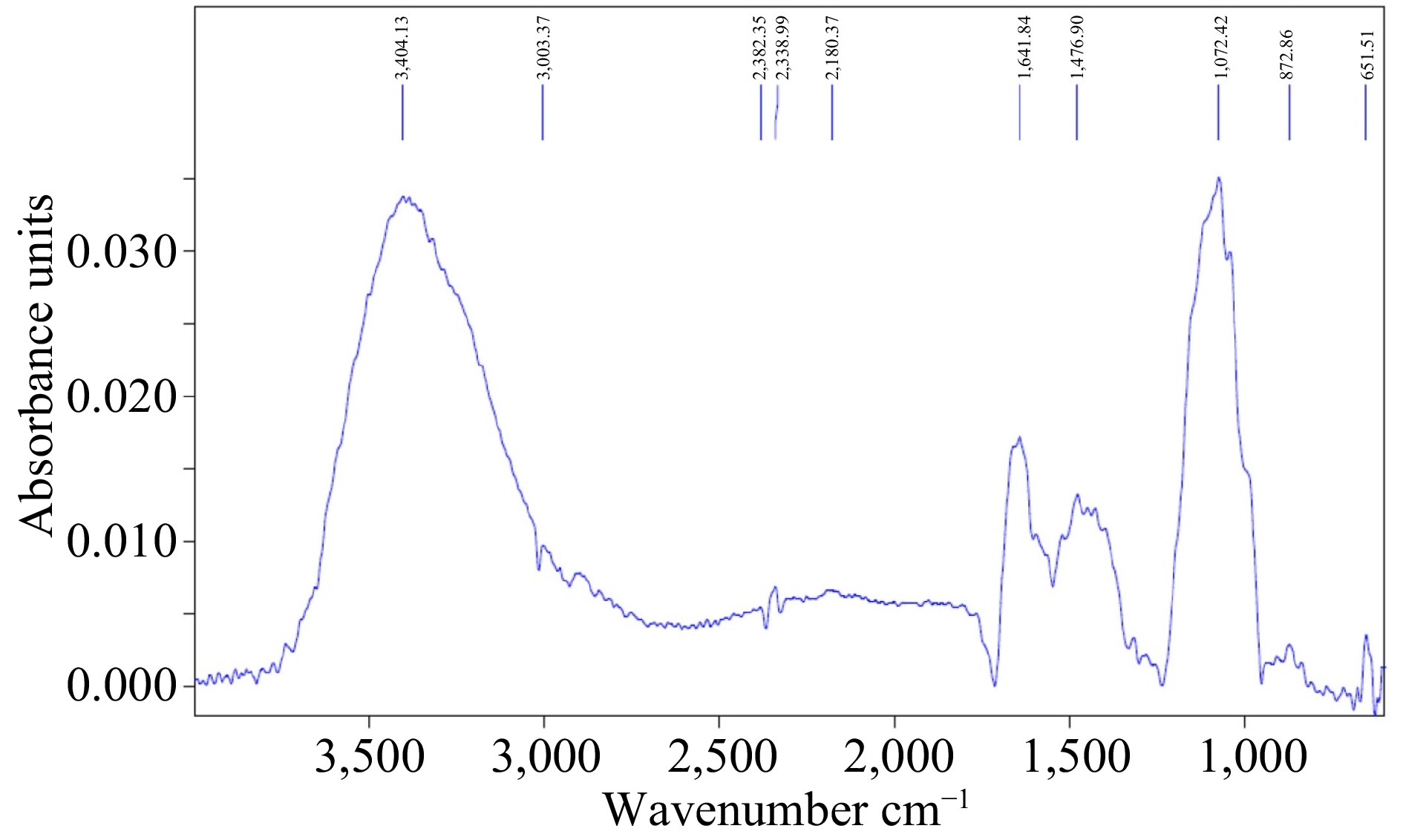
Figure 2.
FTIR spectra of M. indica mediated CuO.NPs. FTIR showed corresponding peaks 3,404.13, 300.37, 2,382.35, 1,641.84, 1,476.90, 1,072.42, 872.66, and 651.51cm−1 corresponding to various functional groups.
Table 1. Possible functional groups in M. indica plant extract.
S.
No.Functional groups Compounds Wave.
no.Vibration Bonding Peaks 1 OH Alcohol 3,404.13 Stretching Strong Broadband 2 O-H Carboxylic 300.37 Stretching Strong Broadband 3 O=C=O Carbon oxide 2,382.35 Stretching Medium Broadband 4 −C=C Alkanes 1,641.84 Stretching Medium Broadband 5 N-H Amine 1,476.90 Bending Medium Broadband 6 N-H Anhydride 1,072.42 Stretching Strong Broadband 7 C-Cl Aldehyde 872.66 Bending Medium Broadband 9 C-Br Alkyl Halides 651.51 Stretching Strong Broadband The current FT-IR analysis demonstrates that green synthesized CuO.NPs have potential as an organic core through the presence of various functional groups, as well as an inorganic support to penetrate the cell wall of microorganisms. Previous studies have indicated that green synthesized CuO.NPs have a variety of reactive functional groups, providing enough potential to be utilized as an effective means of combating pathogens in the agricultural sector[48]. The FT-IR analysis verified the existence of functional groups in the green synthesized CuO.NPs, which possess the potential to effectively inhibit pathogenic growth.
The XRD spectra of synthesized green CuO.NPs demonstrated six major peaks at 2θ angles of 49.67, 39.36, 36.75, 29.56, and 25.34 corresponding to 113, 202, 111, 002, −111, and 110 plans respectively depicting the crystalline nature of nanoparticles (Fig. 3). However, a few unknown peaks were also noticed denoted by *. The average size of the particles was evaluated by Scherrer's equation (D = kλ/β Cosθ) and was retrieved from XRD findings to be 55.29 nm. The XRD results are compatible with the previously studied XRD patterns[49]. The XRD patterns portray the crystalline texture of synthesized particles which play an important role in the interaction of nanoparticles with microbial cell walls[50]. Moreover, the crystalline nature of small-size particles helps to control the formation of biofilm by producing oxidative stress[38].
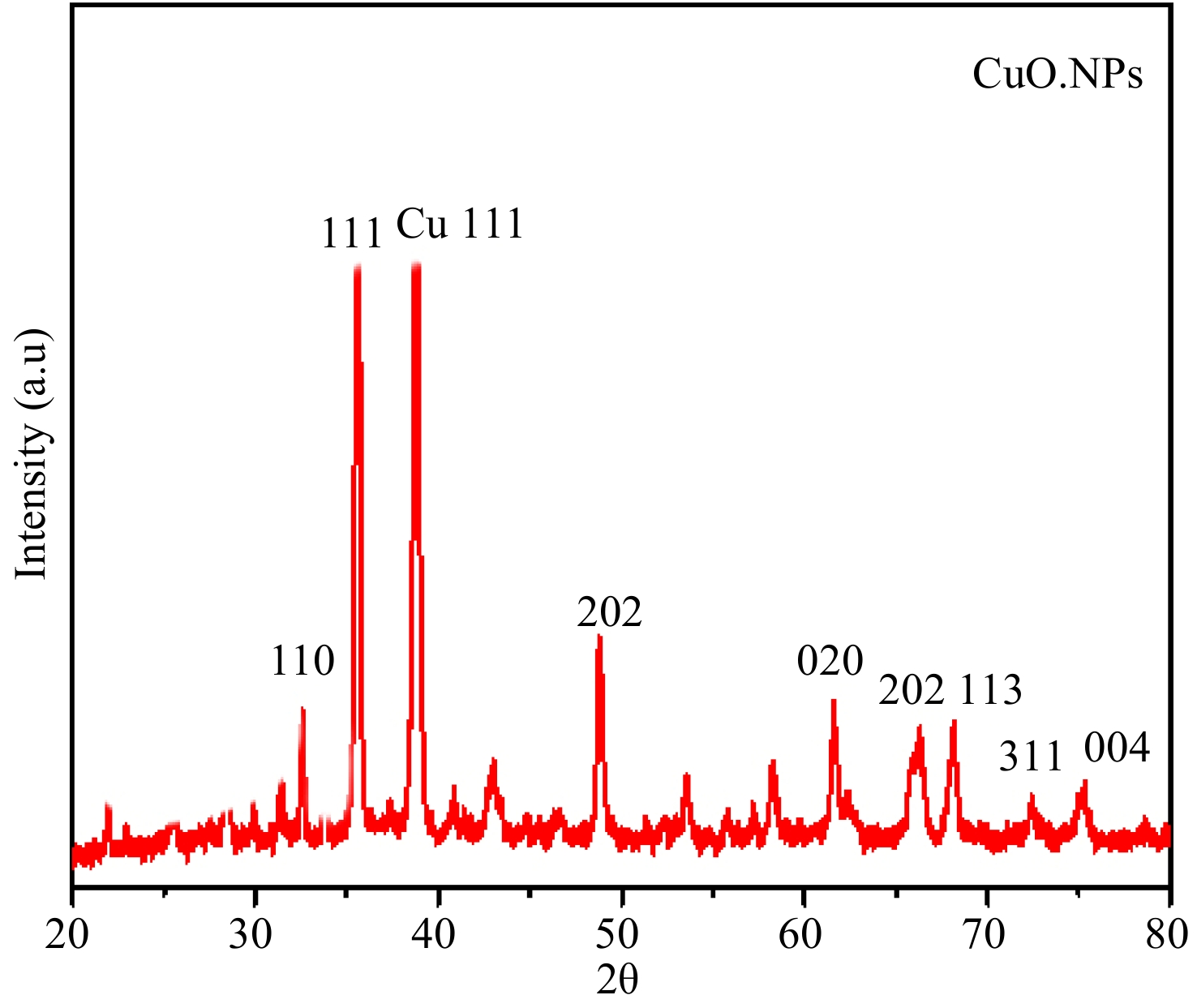
Figure 3.
XRD spectra of M. indica mediated CuO.NPs. The XRD spectra of the green synthesized M. indica mediated CuO.NPs exhibit seven main peaks at 2θ angles of 49.67, 39.36, 36.75, 29.56, and 25.34 plains respectively.
The TEM analysis indicated that the CuO.NPs display a spherical morphology and are effectively dispersed without aggregating, with sizes ranging from 40−80 nm, corresponding closely to the sizes determined from the XRD analysis (Fig. 4a & b). The current analysis is in correspondence with the already reported analysis of green fabricated CuO.NPs. The particle aggregation can be attributed to the electrostatic attraction forces between them. Previous studies suggest that a spherical object can easily enter the microbial cell wall which is mainly responsible for sustaining the integrity of microbes and disturbances in cell walls eventually leading to microbial death[51,52].
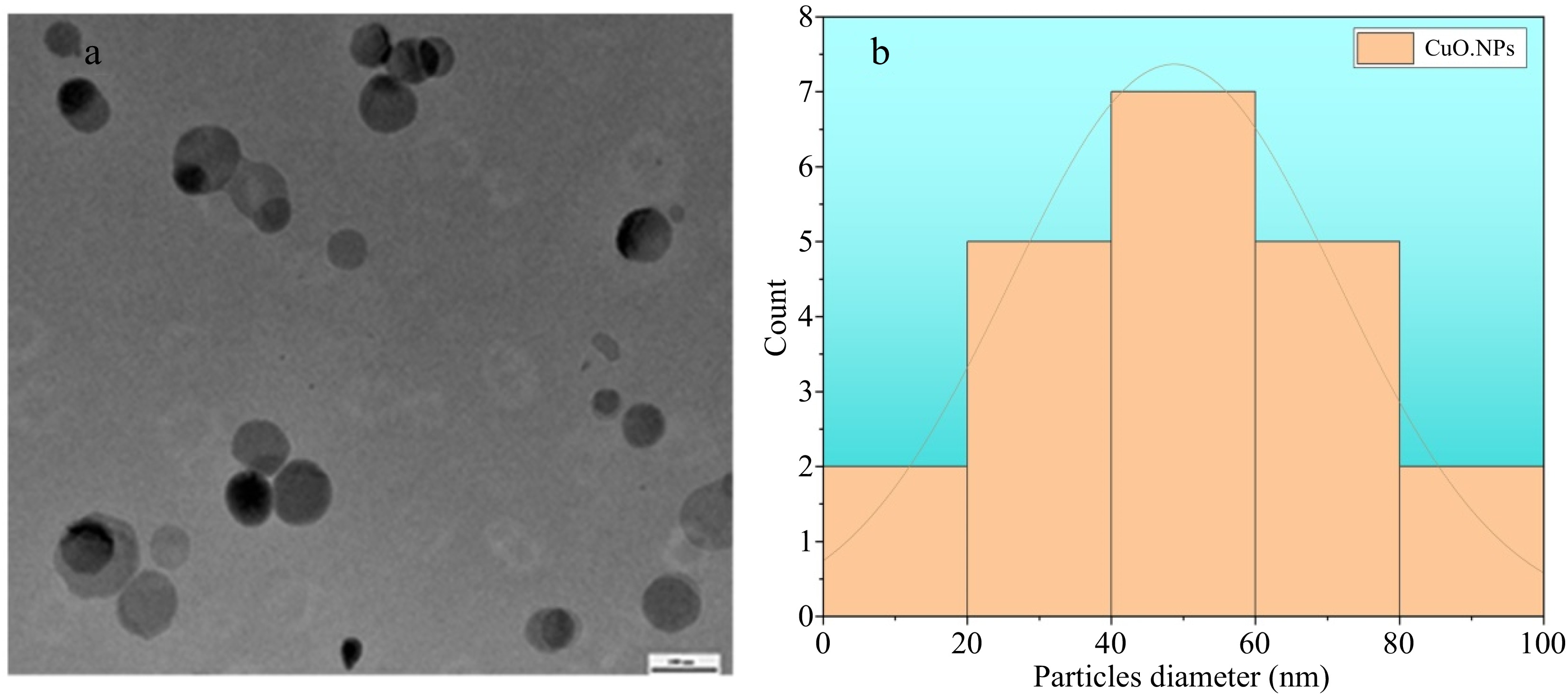
Figure 4.
TEM analysis of green synthesized M. indica CuO.NPs. Spherical shape with a diameter ranging between 30−90 nm.
The TEM results of CuO.NPs shown in Fig. 4 demonstrated a spherical shape having a diameter in the range of 30−90 nm. The electrostatic forces among the particles result in slight agglomeration, but the presence of organic core and plant extract material provides a clear demarcation for each particle (Fig. 4). The shape is likely to increase the surface area of both the particles and the substrate, thereby enhancing their ability to attach and ultimately leading to the antimicrobial effects of CuO.NPs[53].
Antimicrobial activities of CuO.NPS
-
The antibacterial activity of CuO.NPs were evaluated at different doses (30, 60, and 100 µg·mL−1) against E. coli and S. aureus (Fig. 5). In E. coli, the results showed the highest inhibition antibacterial zone at 100 µg·mL−1 of CuO.NPs at 10.8 ± 0.41 (mean ± SD) while 60 and 30 µg·mL−1 from three repeated trials showed inhibitions respectively relative to control 6.4 ± 0.20 and 3.6 ± 0.20 (mean ± SD). Similarly, it was found that green CuO.NPs inhibited the growth of gram-positive bacterial stain efficiently 8.89 ± 0.69 at 100 µg·mL−1 compared with other treatments. While 60 µg·mL−1 of NPs showed a 6.0 ± 0.2 mm zone of inhibition. In the current findings, 3.8 ± 0.2 inhibition zone was observed at the lowest concentration of 30 µg·mL−1 of CuO.NPs. Previously reported studies showed that the bactericidal effects of green fabricated NPs are due to the small size of particles and the availability of various organic functional groups on the nanoparticle's surface[54]. It was reported in a previous study, CuO.NPs found to have highly bactericidal potential to stop their growth. The organic base of nanoparticles attracts the bacterial cells due to strong electrostatic forces of attraction, resulting in the deactivation of the cellular enzymes and causing damage to plasma membrane permeability ultimately causing cell death[55,56]. Moreover, the accumulation of different phytocompounds on the surface of CuO nanoparticles causes damage to the microbial DNA and their proteins formulating enzymes and inhibiting microbial growth. Additionally, the particles' small sizes and crystalline texture were verified by XRD, SEM, and TEM causing the inhibitory effects of the CuO.NPs[57].
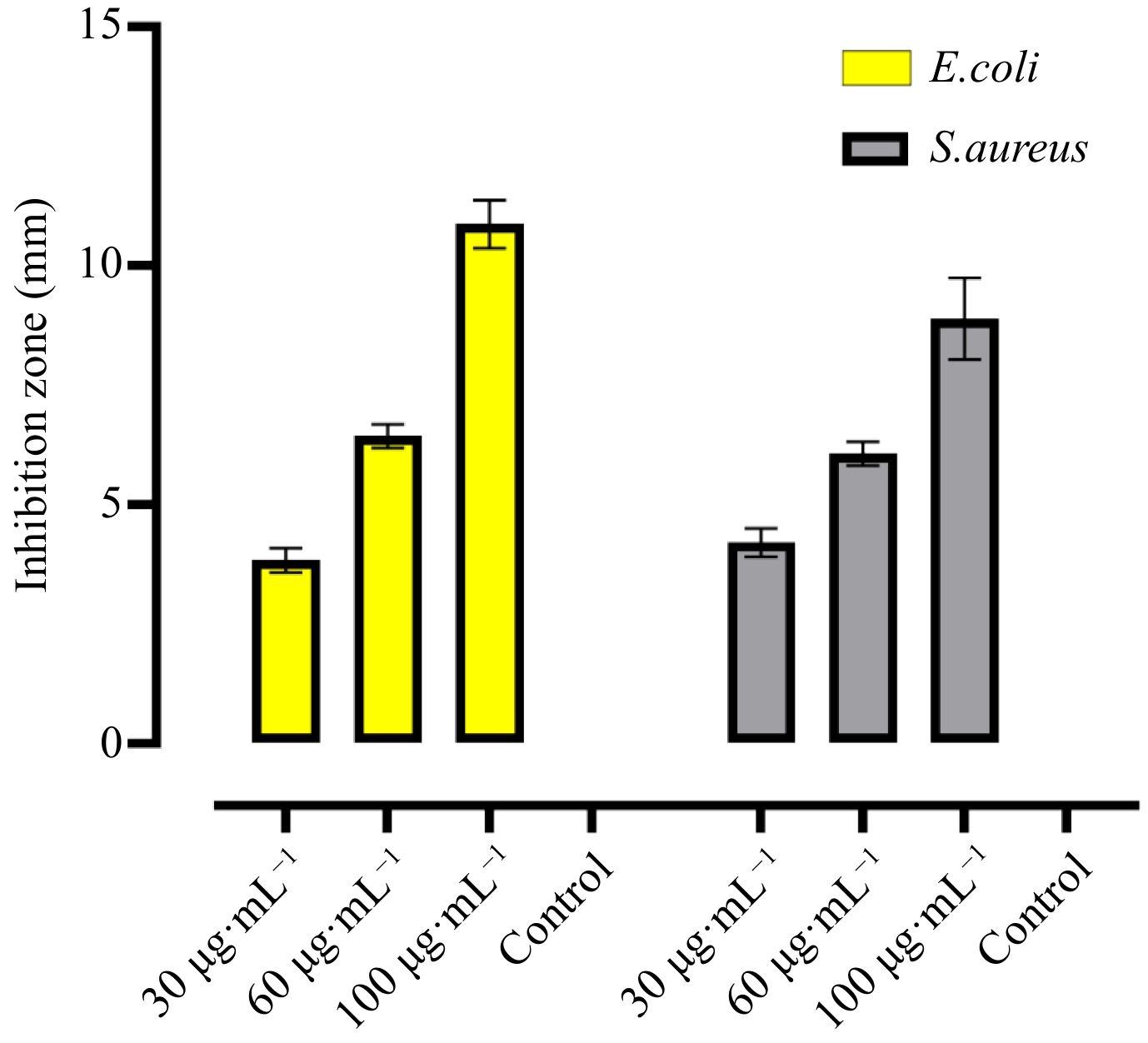
Figure 5.
In-vitro antibacterial activity of CuO.NPs were evaluated at different doses (30, 60, and 100 µg·mL−1) against E. coli and S. aureus.
The antifungal activity in vivo of green synthesized CuO.NPs were evaluated on detached fruit assay. The in vivo study on persimmon fruit resulted in an obvious reduction of disease with an increase in the doses of synthesized CuO.NPs (100, 60, and 30 µg·mL−1) (Fig. 6). In vitro, the highest concentration of 100 µg·mL−1 of particles showed maximum inhibition of disease incidence 23.4 ± 1.81 (mean ± SD) against R. oryzae as compared to control 97.0 ± 0.81 (mean ± SD) (Table 2).
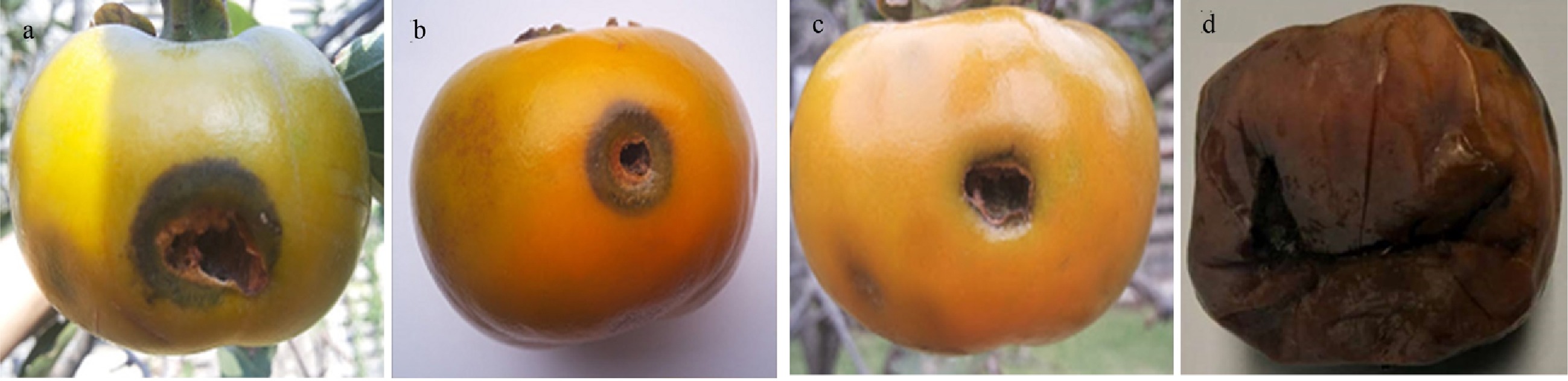

Figure 6.
Fruit detached antifungal in vivo assay. Persimmon fruits were infested and treated with different concentrations of CuO.NPs including: (a) 30 mg·L−1, (b) 60 mg·L−1, (c) and (d) 100 mg·L−1. Control fruit were without any NP.
Table 2. Effectiveness of green CuO.NPs on fruit against disease.
Treatment Diseased area (mm) 30 mg·L−1 64.6 ± 1.6 60 mg·L−1 44.2 ± 0.6 100 mg·L−1 23.4 ± 1.8 Control 97.0 ± 0.81 In this current work, antifungal potential has been shown by green fabricated CuO.NPs both in vivo and in vitro. The presence of plant phytocompounds on the surface of nanoparticles is the main reason for their potential and the ability of CuO.NPs to penetrate fungal cell walls are also dependent on their small size and crystalline texture. The small size of the particles provides a large surface area to adsorb the biomolecules to the cell wall and results in disruption of the cell wall and cellular components, leading to the death of microorganisms. The green fabricated CuO.NPs exhibit potential microbicidal effects, cost-effectiveness, and reproducibility over chemically generated nanoparticles[58]. To identify synergistic effects, copper nanocomposites were tested against the phytopathogenic fungus Alternaria alternata, Rhizoctonia solani, and Botrytis cinerea. According to the findings, nanocomposite showed greater activity at a concentration of 90 g·mL−1. Bimetallic blends antifungal properties were similarly successful in controlling Rhizoctonia solani growth at doses of 30, 60, and 90 g·mL−1. In a greenhouse setting, they also demonstrated successful management of cotton seedling damping-off[59,60].
Antioxidant activity
-
In humans, free radicals are created by several metabolic pathways and cause degenerative diseases and lowered immune function (Liu et al.)[61]. To combat free radicals, medicinal plants with phenolic components are employed as antioxidants. The proportion of scavenging action was substantially higher in the standard and plant extracts, Das et al.[62] reported comparable findings. While Mangifera indica contain polyphenols and other phytochemicals with significant antioxidant activity and are employed as natural antioxidants to control degenerative diseases (Ssekatawa et al.)[63], ascorbic acid is a recognized antioxidant. The mechanism behind the antioxidant activity of inorganic nanoparticles involves the binding of transition metal ion catalysts by free radicals, which results in the enhancement of radical scavenging activity. Antioxidant activity was noticeably elevated in the green synthesized NPs and ascorbic acid. The capping of the CuO.NPs with the phytochemicals utilized in their production were confirmed by FTIR analysis. The biosynthesized CuO.NPs antioxidant activity against the DPPH radical was assessed, and its efficacy was plant extract and conventional ascorbic acid. CuO.NPs had a potent inhibitory effect on DPPH, with an IC50 value of 10.68 0.03 g·mL−1. A dose-dependent reaction of CuO.NPs and ascorbic acid to the DPPH radical are shown in Fig. 7.
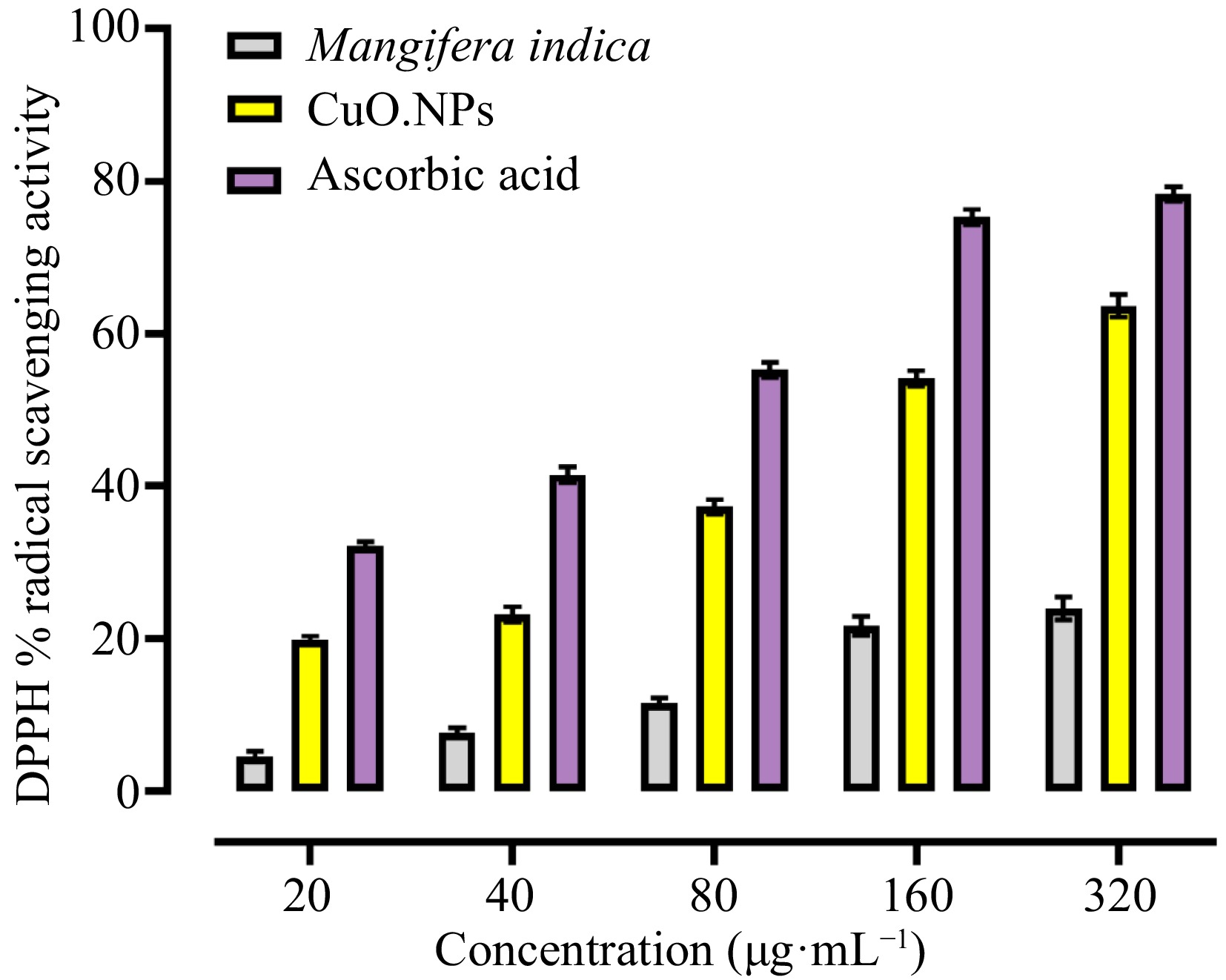
Figure 7.
Antioxidant activity of M. indica CuO.NPs. The DPPH assay was carried out with 20, 40, 80, 160, and 320 µg·mL−1 concentrations of M.indica. CuO.NPs and Ascorbic acid were used as a standard.
CuO.NPs biosynthesized with Sargassum longifolium demonstrated an inhibitory percentage of 20% when tested at 5 g·mL−1, which is consistent with our previous findings[64,65]. On the other hand, copper mixed oxide (CuO/Cu2O) nanoparticles made from Phoenix dactylifera leaves' antioxidant properties showed strong DPPH inhibition at 4 mM[66]. Green chemistry of inorganic metal and metal oxide nanoparticles (NPs) have a wide range of applications in agriculture, particularly in the management and treatment of plant diseases due to fungi. For the correct uses, the biocompatibility and toxicity of the nanomaterials should be investigated[67].
Nanoparticles exhibit antimicrobial and antioxidant potential through multifaceted penetration mechanisms. The nanoparticles bind to microbial membrane and their penetration inside the protoplasm of pathogenic cells has been documented as the most prominent mode of antimicrobial action. NPS released into the microbial cells react with thiol groups causing a denaturation of the proteins and enzymes Fig. 8. The results demonstrated that green modified CuO.NPs can represent a successful alternative treatment for fungi and bacterial infections.
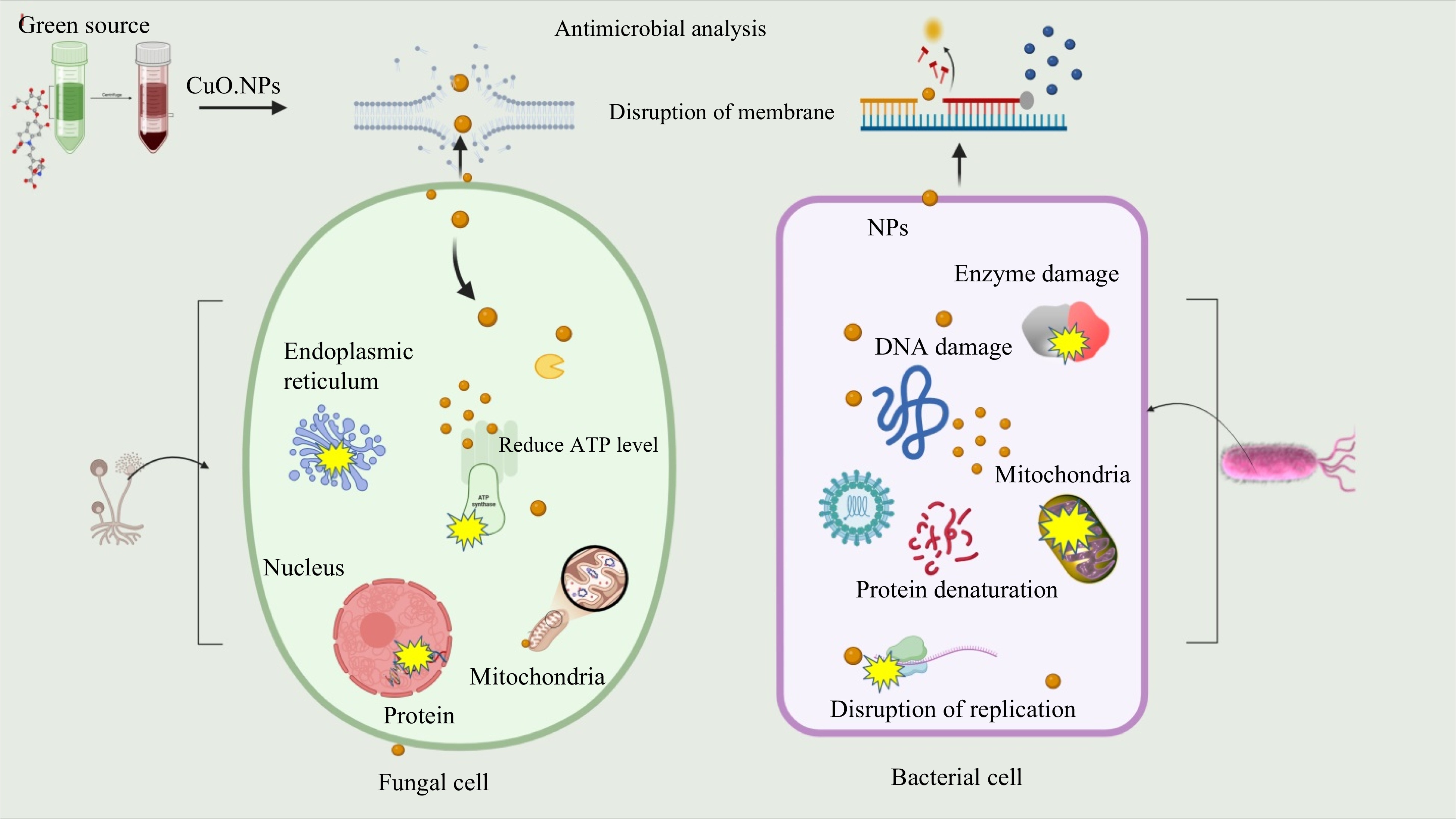
Figure 8.
Expected antimicrobial mechanism of CuO.NPs. Disruption of cell wall and cytoplasmic membrane nanoparticles adhere to or pass through cell wall and cytoplasmic membrane. Cu ions denature ribosomes and inhibit protein synthesis. Interruption of adenosine triphosphate (ATP) production: ATP production is terminated because Cu ions deactivate respiratory enzymes on the cytoplasmic membrane. Reactive oxygen species (ROS) produced by the broken electron transport chain can cause membrane disruption. Cu and reactive oxygen species bind to deoxyribonucleic acid and prevent its replication and cell multiplication. CuO.NPs directly move across the cytoplasmic membrane, which can release organelles from the cell.
-
The current work provides an efficient and cost-effective protocol for the synthesis of CuO nanoparticles by the green approach and its safe use in phytopathology. Using plant materials for particle synthesis allows for the safe and effective use of these particles as antimicrobial agents to combat diseases. This approach is essential in meeting the growing demand for biocompatible solutions in food and agriculture. In this current work, the nanoparticles used against pathogens suppressed their growth by damaging their cell walls and delaying their spore germination. Based on our findings, it is highly recommended to apply green synthesized CuO.NPs mitigate devastating and lethal plant diseases due to their significant antimicrobial effects. This work offers a new avenue for additional investigations of metallic nanoparticles in vital domains like phytopathology and agriculture. The use of this environmentally friendly method of biogenic nanoparticles may help to reduce the fungal infection in sweet fruit and improve the socio-economic standing of farmers.
-
The authors confirm contribution to the paper as follows: study conception, literature analysis data and figures desigen: Shah IH, Ashraf M, Ashraf GA, Rasheed HU, Li G, Mouna J, Faizan M, Altaf MA, Shakoor A; experiments, writing, and prepared the original draft: Shah IH; reviewing data and table creation: Sabir IA, Azam M, Rehman A, Ahmad Z; providing guidance on the whole manuscript: Song C, Manzoor MA. All authors reviewed the results and approved the final version of the manuscript.
-
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
This work was supported by the Key R&D Project of the 14th Five Year Plan of China (2023YFC3503804), Startup fund for high-level talents of West Anhui University (WGKQ2022025), the Open Fund of Anhui Engineering Research Center for Eco-agriculture of Traditional Chinese Medicine (WXZR202318), and Demonstration Experiment Training Center of Anhui Provincial Department of Education (2022sysx033). The authors extend their appreciation to the Deanship of Scientific Research at King Khalid University, Saudi Arabia for funding this work through Small Groups Project under Grant Number (RGP.2/176/44). We thank Dr. Aamir Hasan Shah from the University of California Loss Angles USA for CuO.NPs characterization analysis of XRD and FTIR.
-
The authors declare that they have no conflict of interest.
-
# Authors contributed equally: Iftikhar Hussain Shah, Irfan Ali Sabir
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Shah IH, Sabir IA, Ashraf M, Rehman A, Ahmad Z, et al. 2024. Phyto-fabrication of copper oxide nanoparticles (NPs) utilizing the green approach exhibits antioxidant, antimicrobial, and antifungal activity in Diospyros kaki fruit. Fruit Research 4: e022 doi: 10.48130/frures-0024-0015
Phyto-fabrication of copper oxide nanoparticles (NPs) utilizing the green approach exhibits antioxidant, antimicrobial, and antifungal activity in Diospyros kaki fruit
- Received: 08 November 2023
- Revised: 01 March 2024
- Accepted: 12 March 2024
- Published online: 03 June 2024
Abstract: Nanotechnology has emerged as a prominent field in recent times. The fabrication of biocompatible materials has taken on highlighted significance owing to their requisite application in diverse sectors including medicine, water treatment and purification, health, and other related fields. There has been a lot of research done recently on the green synthesis of various nanoparticles (NPs). Copper a high-performance metal used in agriculture to combat pathogenic attacks, has received less attention. The current work demonstrates the successful preparation of green synthesized copper oxide nanoparticles (CuO.NPs) from Mangifera indica (M. indica) leaf extract. The spectral and morphological characterization biosynthesized were observed using, FTIR, XRD, and TEM analysis. The FTIR analysis revealed the functional groups present in plant extracts. XRD was carried out to demonstrate the crystalline nature and size of nanoparticles using the Scherrer formula. UV was performed to observe the optical properties of NPs. Further, Transmission electron spectroscopy (TEM) was carried out to confirm the physical shape of CuO.NPs with 50 nm. The M. indica mediated NPs were evaluated against gram-negative and positive bacteria Escherichia coli (E. coli) and Staphylococcus aureus (S. aureus) at different concentrations. The in vivo fungicidal activity was performed against Rhizophus oryzae (R. oryzae) on Diospyros kaki (persimmon fruit). The detach fruit method was applied to evaluate the potential of NPs. Higher exposure of 100 µg·mL−1 CuO.NPs showed mycelia inhibition followed by 30, 60, and control treatments. Furthermore, Green CuO.NPs showed prominent antioxidant activities as compared to plant sources. The findings obtained suggest that the green-formulated CuO.NPs could be further investigated for the treatment of many phytopathogenic diseases.
-
Key words:
- M. indica /
- Green synthesis /
- CuO.NPs /
- Antibacterial /
- Antifungal /
- Antioxidant /
- Diospyros kaki












