HTML
-
The moist chamber culture technique (Gilbert & Martin 1933) enables researchers to document myxomycete species at a faster rate than collecting sporocarps that have developed under natural conditions in the field (Alexopoulos 1953). As such, the technique is strongly recommended and widely employed in conjunction with field-based methods to record sporocarps for diversity surveys (Wrigley de Basanta & Estrada-Torres 2017). Several researchers have noted that data generated using the moist chamber technique are taxonomically biased and exclude species with longer developmental times and larger fruiting structures (i.e., Alexopoulos 1964, Stephenson et al. 2004). The assumption has been that this bias exerts minimal impact on diversity studies if the associated field component is comprehensive with multiple sampling periods strategically planned to span the environmental and seasonal variation across the study areas. To date, very little effort has been made to quantify the extent of the bias imposed by using the moist chamber technique.
Most biological work with myxomycetes has centered on documenting the diversity and distribution of species (Novozhilov et al. 2017). The resultant data has often been used to infer ecological attributes even though the studies were not explicitly designed to quantify such characteristics. Interest in designing observational and experimental studies to explicitly test ecological hypotheses and produce reliable and robust information that can be used to understand general ecological principles and processes as well as inform the management of natural resources has increased. Toward these means, projects aimed at elucidating ecological attributes cannot continue to rely on traditional diversity approaches. Thorough evaluations of existing diversity-based approaches to quantify the sources and extent of bias, error, and variability are required to inform study design, statistical approaches, and interpretation for ecological studies. The use of the moist chamber culture technique, based on Gilbert and Martin's approach, does not provide a true replicable standard appropriate for ecological studies. There are several factors, not considered in biodiversity assessments, that can influence the results such as substrate mass, moisture levels, laboratory climate, and temporal extent. Rojas et al. (2021) provided quantified data validating that some of these factors can explain differences among datasets despite the large influence of a system's natural variability.
The sampling schemes used to collect substrate materials (e.g., plot, plotless, random, systematic, etc.), the precise parameters used for moist chamber preparation (e.g., mass, heterogeneity, quality of substrate material(s) etc.), the laboratory incubation conditions (temperature, humidity, quantity of electromagnetic radiation, air flow, etc), and the data collection parameters (e.g., number of sporocarps, number of spores produced, days until fully developed, etc.) have been highly variable among modern investigations and a standardized methodological approach has not been proposed. An opportunistic approach as described by Cannon & Sutton (2004) has been highly recommended for diversity studies as it maximizes the probability of surveying the widest range of potential productive niches; however, biodiversity assessments regularly utilize other sampling schemes (e.g., Campbell & Dooley 2008). Plot-based sampling approaches, which are designed to document the density and frequency of organisms, have been used as a framework to collect substrates for myxomycete studies even though the studies being conducted were not intended to study such parameters (i.e., Dagamac et al. 2014). These discrepancies occur due to the absence of a widely accepted standardized methodological approach developed from the results of quantitative studies of the moist chamber technique and associated sampling schemes. Despite the issues mentioned above, the technique is remarkably useful for macroecological purposes (Wellman 2020) or ecological studies that require researchers to execute a higher level of control on ecological forces (i.e., Walker et al. 2019).
The study presented herein discusses the results of three distinct investigations conducted in a Costa Rican tropical forest where a wider, long-term, effort to understand the ecology of myxomycetes is ongoing (Rojas & Stephenson 2021). The main objective of the present discussion is to elucidate relevant parameters that can be used to assess the utility and limitations of the moist chamber culture technique for ecological and applied studies using myxomycetes as focus organisms. The three experiments explained herein are integrated within the idea that the moist chamber technique is a relevant method to address the effect of global changes on myxomycetes. By using the sporocarp phase as a proxy to quantify their ecological associations with natural and non-natural environments, myxomycete data is remarkably relevant for easy-to-implement monitoring of microbiota.
-
The study presented herein was carried out in the Finca Experimental Interdisciplinaria de Modelos Agroecológicos (FEIMA) in Turrialba, Costa Rica during 2018 and 2019. This research station, administrated by the University of Costa Rica, encompasses a 28-ha successional forest patch in the premontane tropical moist forest life zone. In this area, a thorough 30-month assessment of myxomycete occurrence, based on field collections, was carried out to evaluate phenological patterns in tropical species (Rojas & Stephenson 2021). Based on the general recommendations for myxomycete surveys (Wrigley de Basanta & Estrada-Torres 2017), the moist chamber technique was utilized to increase the number of species recorded from the study area.
With the latter technique, three discrete experiments were carried out with the goal of evaluating the effect of methodological modifications on the generation of data (Fig. 1). As such, the first experiment tested the idea that different results can be obtained when the isolation process takes place in different laboratories in a similar manner to Rojas et al. (2021). For this experiment a total of 432 substrate samples were collected in the form of 12 samples of both litter and bark at ground level, at 1 m and at 2 m of distance from the ground in six independent locations. Once collected, the material was divided in two sets of 216 samples corresponding to exactly half of the collecting effort and taken to different laboratories (named laboratory one and two in this study) for isolation with the moist chamber technique. In this case, litter samples collected at the ground were considered ground litter, whereas litter samples collected above the ground were considered aerial litter.
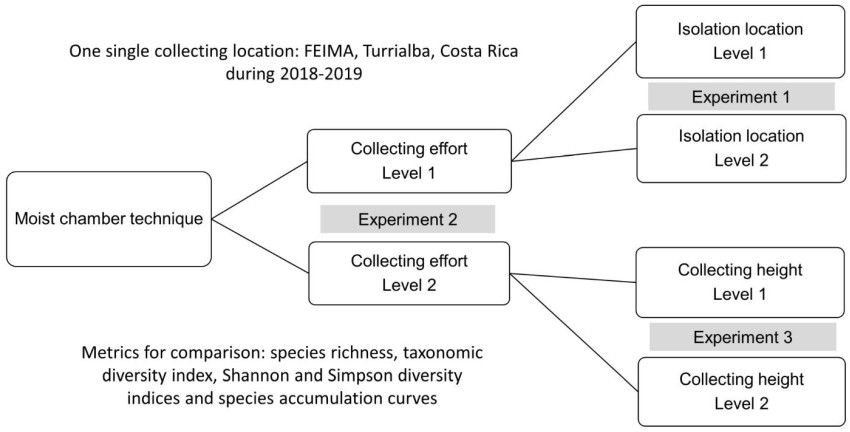
Figure 1. Organization of the present study in three discrete experiments based on comparisons of moist chamber data obtained at two levels for each variable under evaluation.
The second experiment was designed with the goal of testing the effect of sampling effort. For this, a similar design to the first experiment was used but with an 67% reduction in collection effort. As such, only 72 total samples were collected in the form of two samples (instead of 12) of both litter and bark at ground level, at 1m and at 2m above the ground, in the same six independent locations. The material was taken to one of the laboratories used in the previous experiment for the isolation process with the moist chamber technique. In this case, the resulting dataset was only compared with the extended sub dataset of the first experiment isolated in the same laboratory.
The third experiment was implemented with the goal of evaluating whether the original height at which the material was collected in the field could influence the resulting data. As such, this experiment utilized the reduced effort strategy consisting of 72 samples, but with this approach, six samples of twigs and six of litter were collected only at ground level in each of the six independent locations. For analysis, this dataset was compared with the dataset of 72 samples used in the second experiment that reflected the reduced effort.
In all cases, substrate samples were used to set up moist chamber cultures in the manner described by Stephenson & Stempen (1994). The results from all experiments were compared in terms of species richness, the Shannon and Simpson (1-D) diversity indices, and the taxonomic diversity index as response variables. Compositional similarity was analyzed using the Bray Curtis distance ranging from 0 for two datasets that are completely different to 1 for two datasets that were exactly the same. A cluster analysis was constructed in the second experiment using these data. With the first and second experiments, the analysis of the same parameters by height was also performed. Analyses of Variance was used to evaluate differences in the number of records and species by height categories in the first experiment. In all cases, the software PAST (Hammer et al. 2001), v 4.06b was used.
-
Thirty-two species of myxomycetes were recorded across all three experiments. From these, 25 were recorded in the first experiment (432 moist chambers), 22 in the second experiment (288 moist chambers) and 25 in the third experiment (144 moist chambers). Similarly, 224 records of myxomycetes were made in the first experiment, 175 records in the second one and 136 records in the third one. As such, the observed productivity was 0.51, 0.60 and 0.94 records/moist chamber for the first, second and third experiments, respectively. In both terms of species and records, the results of the three experiments did not show a particular pattern based on the number of studied moist chambers. The relationship between effort and productivity was strong and negative, but not significant (r = -0.94, p = 0.2).
The first experiment produced 108 records representing 18 species at laboratory one and 116 records representing 20 species at laboratory two (Table 1). The Shannon index of diversity was 2.63, the Simpson index of diversity was 0.91, and the taxonomic diversity index was 1.8 in both cases. As such, no differences in diversity were observed. The Bray Curtis distance between datasets was 0.47 indicating that the species assemblages recovered between the labs were compositionally different.
Table 1. Records of myxomycetes by species observed in the first experiment of the present study arranged by height above the ground
Species Laboratory one Laboratory two 0 m 1 m 2 m Subtotal 0 m 1 m 2 m Subtotal Arcyria afroalpina 2 1 1 4 Arcyria cinerea 2 5 4 11 7 6 5 18 Arcyria insignis 1 1 Comatricha elegans 1 1 Comatricha nigra 2 2 1 1 Comatricha tenerrima 1 1 2 Cribraria violacea 1 1 2 3 5 Didymium anellus 1 1 2 Didymium bahiense 5 7 7 19 1 1 Didymium difforme 1 2 3 2 2 4 Didymium minus 2 7 7 16 Didymium squamulosum 3 2 5 3 1 1 5 Diderma hemisphaericum 1 2 4 7 4 4 2 10 Hemitrichia minor 3 1 4 Hemitrichia pardina 1 1 Lamproderma scintillans 1 1 3 5 2 2 4 Perichaena chrysosperma 4 3 6 13 3 3 4 10 Perichaena pedata 2 4 4 10 1 1 Physarum compressum 3 6 5 14 5 5 6 16 Physarum didermoides 2 1 3 6 3 6 4 13 Physarum pusillum 1 1 2 2 2 Physarum superbum 1 1 Stemonitis axifera 1 1 Stemonitis fusca 1 1 2 Stemonitopsis aequalis 1 1 With respect to the three heights from which the samples were collected, 56 records representing 17 species were recorded at ground level, 81 records representing 19 species were collected at 1 m above the ground and 87 records representing 21 species were collected at 2 m above the ground. Both the number of records and the number of species were significantly higher at 2 m when compared to the other two heights (F(2, 33) = 2.6, p = 0.03, F(2, 33) = 3.9, p = 0.03, Tukey p = 0.03). The Shannon index of diversity was 2.7 for the ground level and 2.8 for the other two heights, and the Simpson index of diversity was 0.92 for the ground level and 0.93 for the other two heights. The taxonomic diversity index had a value of 2.1 in all cases. The Bray Curtis distance calculated for the comparison between datasets collected at the ground level and 1 m above the ground was 0.80, whereas the same value for the comparison between datasets at 1 and 2 m from the ground was 0.89. The same value for the comparison between datasets from ground level and 2 m above the ground was 0.77.
In the second experiment, 108 records representing 18 species were observed with the full effort of 216 samples and 67 records representing 19 species were observed with the reduced effort of 72 samples (Table 2). The productivity of the full dataset was 0.5 records and 0.08 species/moist chamber and for the reduced effort were 0.93 and 0.26, respectively. The Shannon diversity index was 2.7 in both cases, whereas the Simpson diversity index was 0.91 for the full dataset and 0.90 for the reduced dataset. The taxonomic diversity index was 1.8 for the full dataset and 2.1 for the reduced effort. The Bray Curtis distance between datasets was 0.64.
Table 2. Records of myxomycetes by species observed in the second experiment of the present study arranged by height above the ground
Species Full effort Reduced effort 0 m 1 m 2 m Subtotal 0 m 1 m 2 m Subtotal Arcyria afroalpina 1 2 3 Arcyria cinerea 2 5 4 11 4 2 1 7 Arcyria insignis 1 1 2 2 Comatricha nigra 2 2 Comatricha tenerrima 1 1 2 1 1 2 Cribraria violacea 1 1 2 2 Didymium bahiense 5 7 7 19 1 2 2 5 Didymium clavus 1 1 2 Didymium difforme 1 2 3 1 2 3 Didymium iridis 1 1 Didymium squamulosum 3 2 5 1 4 3 8 Diderma hemisphaericum 1 2 4 7 2 2 Hemitrichia minor 3 1 4 1 1 2 Lamproderma scintillans 1 1 3 5 1 2 3 Perichaena chrysosperma 4 3 6 13 1 1 1 3 Perichaena depressa 1 1 Perichaena pedata 2 4 4 10 1 1 Physarum compressum 3 6 5 14 5 7 5 17 Physarum didermoides 2 1 3 6 1 1 Physarum pusillum 1 1 2 1 1 2 Physarum superbum 1 1 Stemonitis fusca 1 1 2 In terms of height, at ground level, 11 species were observed in both datasets, but the full and reduced dataset were associated with 21 and 31 records, respectively. At the height of 1 m above ground, the full dataset documented 40 records representing 14 species, whereas the reduced dataset was associated with 41 records representing 13 species. At the height of 2 m above ground, 45 records representing 15 species were observed in the full dataset, whereas 42 records representing 16 species were associated with the reduced dataset. Both the Shannon and Simpson indices of diversity were similar between datasets for all heights, with values of 2.4 and 0.91 for ground level, 2.5 and 0.91 for 1 m and 2.7 and 0.93 for 2 m above the ground. The Bray Curtis distances were used to create a cluster dendrogram illustrating that the datasets were compositionally distinct, with the assemblages associated with the ground layer being the most distinct in both cases (Fig. 2).
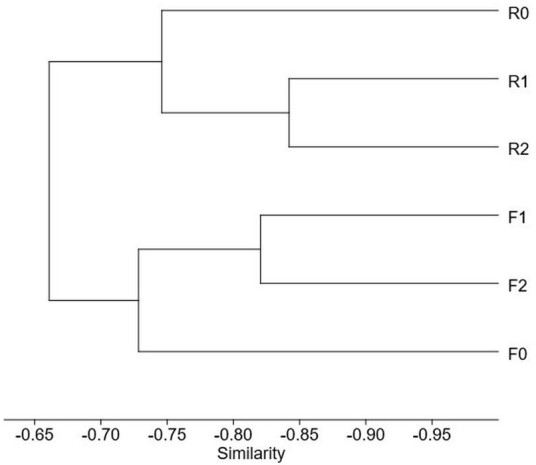
Figure 2. Cluster analysis carried out with the data from the second experiment showing the Bray Curtis similarities among sub datasets collected at different heights for both studied datasets. F = full dataset, R = reduced dataset. 0, 1 and 2 refer to ground level, 1 m and 2 m above the ground.
In the third experiment, both datasets showed similar results. A total of 19 species representing 67 records were observed in the dataset with the full height sampling scheme, whereas 19 species representing 69 records were observed in the dataset that only evaluated ground-level substrates. With these results, an average of 0.94 records/moist chamber and 0.26 species/moist chamber were observed in both cases. The Shannon index was 2.7 and the Simpson index was 0.90 in both cases. The taxonomic diversity index was 1.9 for the dataset with full height and 2.1 for the ground-level dataset. The Bray Curtis distance between datasets was 0.52 suggesting differences in the structural composition that are visible on Table 3.
Table 3. Records of myxomycetes by species observed in the third experiment of the present study arranged by sampling type
Species Full height sampling Ground level sampling Arcyria afroalpina 3 11 Arcyria cinerea 7 16 Arcyria insignis 2 6 Comatricha nigra 3 Comatricha tenerrima 2 8 Cribraria violacea 2 2 Didymium anellus 1 Didymium bahiense 5 1 Didymium clavus 2 Didymium difforme 3 1 Didymium iridis 1 Didymium minus 2 Didymium squamulosum 8 Diderma hemisphaericum 2 3 Hemitricha minor 2 1 Lamproderma scintillans 3 2 Perichaena chrysosperma 3 2 Perichaena corticalis 3 Perichaena depressa 1 Perichaena pedata 1 2 Perichaena vermicularis 1 Physarum compressum 17 3 Physarum didermoides 1 Physarum oblatum 1 Physarum pusillum 2
-
The experiments discussed above illustrated clear differences between using the moist chamber technique for diversity or ecological studies using myxomycetes. The main goal of the first approach is to record as many species as possible with a given effort, whereas the goal of the latter was to elucidate patterns and discover the underlying mechanisms producing them through species-environment interactions (Tucker et al. 2005). The moist chamber culture technique was not developed to generate diversity or ecological data, but its original intent was more aligned with the former. The original objective of the experiment explained by Gilbert & Martin (1933) was to promote algae grow on pieces of bark, and the observation of myxomycetes was a surprising secondary result. These researchers eventually experimented with different bark samples and started recording myxomycetes that were rarely collected at the time (see comments from Alexopoulos 1964), which ultimately led them to recommend the technique for diversity surveys. In this manner, the moist chamber technique was conceptually conceived as a successful strategy to record myxomycetes, but from a methodological point of view, the technique only generates presence-based data on the group of myxomycetes that can form sporocarps in the microcosm of the culture.
The limitations associated with the use of the technique, however, work as a stabilizing force, that albeit artificial, facilitates the formation of myxomycete sporocarps by increasing the homogeneity of the microcosms. As such, the results shown herein were not particularly unexpected. The data from all three experiments carried out in the present study were consistent in showing the lack of differences in response metrics between the two levels associated with the studied factors in each case. As such, diversity-based variables such as species richness, species diversity (calculated with the two indices) and the taxonomic diversity index were equivalent in all cases. This result has two implications since it demonstrates the consistency of the moist chamber technique to generate data, but it also shows, as observed in the experiment two, that sampling effort is a factor of consideration in the design of biodiversity surveys (see the cost-benefit balance of Rollins & Stephenson 2012). The only meaningful differences for data interpretation observed herein were based on compositional aspects of the myxomycete assemblages associated with the different evaluated levels (see Tables 1, 2 and 3 and low values of the Bray Curtis distances).
Interestingly, differences in the taxonomic structure of datasets, both in terms of species richness and number of observations per species, provide ecological meaning to results. However, as discussed earlier, if the moist chamber technique is implemented using the original simplistic protocol, empirically based ecological conclusions cannot be obtained and substantiated. For instance, Cavalcanti et al. (2015) recorded 10 species of floricolous myxomycetes from 368 moist chambers, but Rojas et al. (2016) recorded an average of 12.8 species in six groups of 90 moist chambers each. This means that with an effort of 24%, the second authors recorded as many species growing on tropical inflorescences as the first ones, demonstrating again that smaller efforts can yield greater productivity in a given situation. In fact, Schnittler & Stephenson (2002) carried out a study with a similar effort to Cavalcanti et al. (2015), but recorded three times as many species. In all these studies, however, it seems likely that experimental considerations such as the time of the year when substrates were collected, the chemical quality of the substrates (see Stephenson et al. 2020), or even the distance from the ground at which samples were collected (observed herein), could be associated with the differences in results. Moreover, other methodological aspects of the laboratory procedures may have affected results as well. In this manner, it is impossible to make a meaningful ecological comparison of results among these three studies since the only methodological elements in common among all of them were the use of Petri dishes, filter paper, distilled water, and floral parts of Heliconia plants. A large number of other "comparable" studies would be subject to the same limitations with respect to drawing valid comparisons.
Significant compositional evaluations of myxomycete assemblages associated with specific compartments of ecosystems require a refined protocol of the moist chamber technique. In recent years, some applications where myxomycete data is useful have been evaluated, including urban assessments (Hosokawa et al. 2019), bioremediation (Kryvomaz & Maximenko 2016), and climate interactions (see Li et al. 2021). For these ecological studies, and many others, a standardized technique would have generated data with increased broader impacts as they could truly be compared among studies. In this manner results from each experiment in the present study are comparable at the different set levels because researchers used standardized conditions throughout the entire process, but they are not truly comparable with similar assessments made by other researchers in other parts of the world. In other words, the lack of a standardized moist chamber technique imposes an inherent constraint on the use of ecological data generated with the technique.
In general, the other three lessons that can be drawn with data from the present study are that moist chamber productivity decreases at higher levels of sample coverage, that aerial litter is a highly productive substrate in tropical forests, and that different substrates generate data with different compositional structure. None of these observations are new and they all support the results from previous assessments using moist chambers. For instance, lower effort levels generate data at a faster pace than higher ones, simply because accumulated data tends to stabilize after a particular effort threshold (see results from Rojas & Stephenson 2020). Also, aerial litter has been recognized as a very good substrate for myxomycetes (Novozhilov et al. 2017) and on some occasions, such as this study, response variable values can be greater on it. However, ground litter is often associated with a high number of records and species within the genus Didymium (also observed herein) and studies targeting such genus could use this information for design purposes.
Results from the present study suggest that a collecting effort designed to record system variability (i.e., spatial and temporal heterogeneity) represents a superior cost-benefit situation for synecological studies than a more intense effort designed only for a limited spatial or temporal purpose. However, common practices such as substrate differentiation and the establishment of different sampling areas can have varied implications for the results obtained from diversity or ecological surveys. In the former, they simply increase the probability of recording more myxomycetes, but for the latter they increase the probability of recording more variability and thus, generate data more appropriate for pattern detection. As such, substrate collection for moist chamber evaluations should be opportunistic (not based on plots or transects) when a floristic or diversity study is intended. In those cases, less samples per collecting site, but more (and different) collecting sites would likely generate a better picture of the myxobiota. Density or frequency evaluations that require plot sampling, should be restricted for ecological studies. However, for these, more analyses on the use of the moist chamber technique are needed, particularly at the level of laboratory procedure, in order to generate a more complete picture of the utility and limitations associated with the use of this technique. Only then will meaningful and truly comparable data be generated for ecological studies of myxomycetes.
- This work was financed by the University of Costa Rica (Vicerrectoría de Investigación 570-B8-006). Appreciation is extended to Shiori Nakajima and Werner Rodríguez at FEIMA. This study was carried out as part of a research agreement between the University of Costa Rica and Lincon Memorial University.
- Copyright: © 2021 by the author(s). This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
| C Rojas, AW Rollins, R Valverde. 2021. Generation of myxomycete data from three discrete experiments using moist chamber cultures in a Neotropical forest. Studies in Fungi 6(1):450−459 doi: 10.5943/sif/6/1/34 |












