-
Freezing is one of the oldest and most widely used food preservation methods, which enables better retention of taste and nutritional value of most foods compared to other methods such as thermal processing[1]. This low temperature inhibits the growth of microorganisms and reduces the rate of chemical reactions, thus maintaining the quality of foods. With the advances in freezing operations, there are still several drawbacks of freezing preservations which deteriorate the quality of food products. Ice recrystallization is one of the major causes of quality deterioration of frozen products, since the size and quantity of ice crystals closely relate to the quality of frozen foods[2,3].
To minimize the formation of large ice crystals, it is critical for frozen food manufacturers to select appropriate processing parameters (e.g. quick freezing) and utilize ice crystal growth inhibiting ingredients[3]. For instance, polysaccharide gums are used in ice cream production to prevent the coarse and icy texture after heat shock during storage and transportation by enhancing viscosity. However, this could lead to undesirable melting characteristics such as higher mix viscosity and a chewy body. Hence, there is still a need to identify new ingredients which can inhibit ice recrystallization in frozen foods with or without minimal quality changes.
With an increased consumer demand for healthy and natural foods, food industries are striving to meet these trends by offering 'clean-label' products which consist of ingredients perceived as natural and known by consumers. This brings the interest in finding natural anti-freezing alternatives such as anti-freezing proteins (AFPs). AFPs are a group of proteins that protect organisms from deep freezing temperatures and are naturally present in many foods, such as fish, peach, winter wheat, and kale[4,5]. The presence of AFPs enable the survival of many organisms under extreme cold conditions[6,7]. APFs bind to the surface of nascent ice crystals thus depressing the freezing point of water in a non-colligative manner[8]. They are believed to inhibit the growth of ice crystals by an adsorption-inhibition mechanism[5]. The adsorption of AFPs at the ice-solution interface results in local surface curvature effects, leading to the inhibition of ice crystal growth[9,10]. This ability of AFPs to retard ice recrystallization enables their potential use as a natural ice growth inhibitor in frozen products such as ice cream[11] and sucrose solutions[12]. However, the high cost and low yield of natural AFPs limit their applications in food manufacturing[5].
Hence, an economical and practical source with ice recrystallization inhibition (IRI) activity needs to be explored for application in food processing. Some studies have demonstrated that protein hydrolysates could be potentially utilized to inhibit ice recrystallization in food products[2,13−15]. Mueller & Liceaga[13] have reported the cryoprotective activity of silver carp protein hydrolysates. Hydrolyzed fish gelatin and bovine collagen, both rich in proline, have demonstrated ice crystallization inhibition activity in ice creams[2,14,15]. However, besides the studies on gelatin, collagen, and fish hydrolysates, there have not been many other studies on the IRI activity of protein hydrolysates. Hence, this study focused on investigating the IRI activity of protein hydrolysates obtained from different types of animal- and plant-based proteins via enzymatic hydrolysis.
The objective of this study is to evaluate the IRI capabilities of different animal- or plant-based protein hydrolysates for their applications in food processing. Studies have demonstrated the importance of amino acid composition on the IRI activity of AFPs[2,14,16,17]. Specifically, threonine[16,17], asparagine[18,19], alanine[18,20], and proline[2,14] are found to play an important role in ice binding. In addition, the amphiphilic structure of AFPs is considered to also contribute greatly to their IRI activity[21,22]. Hence, six proteins of different amino acid compositions were examined in this study, including sodium caseinate (SC), whey protein isolate (WPI), egg white protein (EWP), pea protein isolate (PPI), soy protein isolate (SPI), and commercial gelatin (GE). These proteins are common food proteins with adequate amounts of the four IRI contributing amino acids[23−26], thus making them economical and practical sources to produce IRI protein hydrolysates which can be utilized to ensure the quality and extend the shelf-life of frozen food products. Moreover, succinylation reaction was performed on selected peptides to examine the effect of chemical modification and amphiphilicity change on the IRI activity of hydrolysates.
-
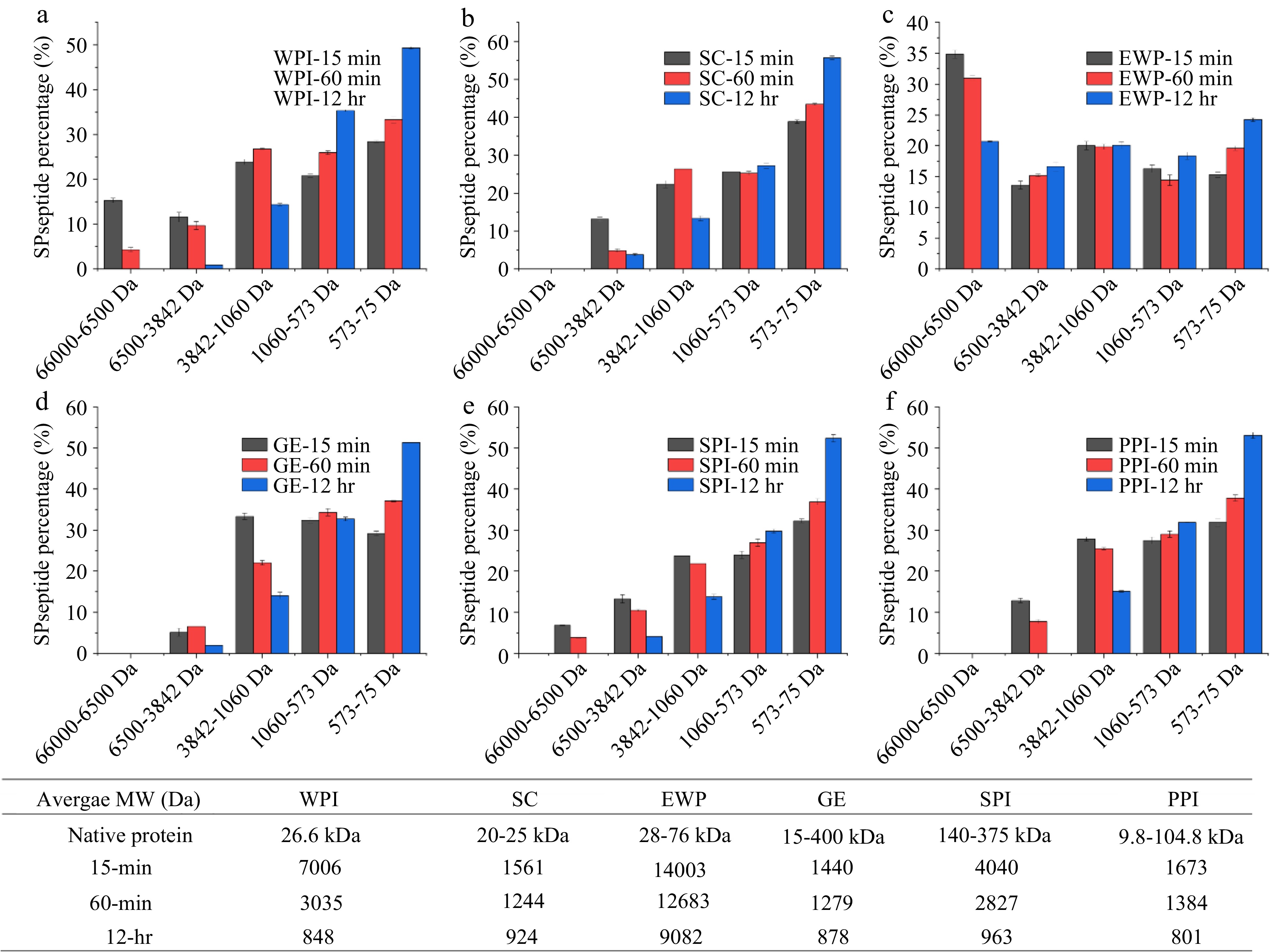
To evaluate the potential IRI activity of protein hydrolysates, alcalase, a non-specific protease, was used to hydrolyze six different types of protein including SC, WPI, EWP, GE, PPI and SPI. The hydrolyzed peptide underwent molecular size distribution measurement using HPLC and IRI activity assessment by splat assay. Figure 1 demonstrates the peptide molecular weight distribution of the hydrolyzed protein samples and their weighed average molecular weight (Da) at different hydrolyzing times. Overall, increased hydrolysis time led to a decrease in the average molecular weight of sample as shown in Fig. 1. Except for EWP, the extensive 12-hr hydrolysis has resulted in an average molecular weight of the hydrolysates less than 1,000 Da. This indicates the lower efficiency of alcalase in hydrolyzing EWP compared to the hydrolysis of the other five proteins examined in this study. EWP has been reported to contain proteinase inhibitors which could lead to the reduced hydrolysis activity of alcalase[27,28].
Figures 2 & 3 demonstrate the ice crystal microscopic images and mean ice crystal sizes of various hydrolysate samples in PBS buffer at 4% (w/w) after 30 min annealing at −8 °C. Polyethylene glycol (PEG) in PBS buffer at 4% (w/w) was the negative control[22]. Without enzymatic hydrolysis, except WPI, the five types of native proteins were not soluble in PBS buffer at 4% (w/w) concentration, and thus could not be tested for IRI activity. WPI was soluble but did not show IRI activity at 4% (w/w) concentration. As shown in Fig. 2, compared to the 4% PEG, much smaller ice crystals were observed for the treatment of 4% SPI-15min hydrolysate, which indicates this sample is IRI active, whereas similar or larger ice crystals were observed in SC, WPI, EWP, GE and PPI after 15 min enzymatic hydrolysis. Quantitative ice crystal results are presented in Fig. 3 which indicates that after 15 min hydrolysis, except SPI hydrolysates, the other five samples tested did not have an IRI activity compared to PEG.
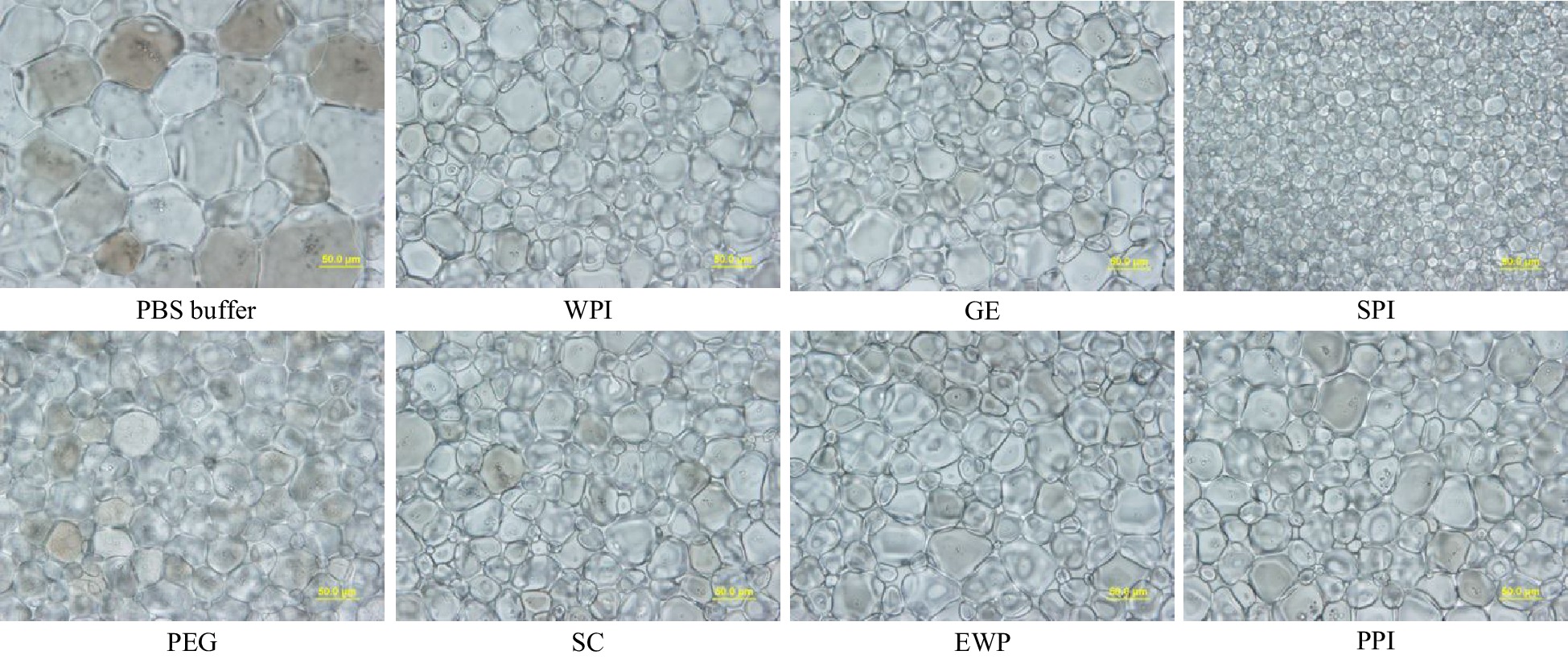
Figure 2.
Standard splat assay microscopic image at 40× for PEG (negative control), and GE, SPI, SC, EWP and PPI hydrolysates after 15-min alcalase hydrolysis at 4% (w/w) concentration, and phosphate buffer saline (PBS – the buffer solvent) annealed at −8 °C for 30 min (scale bar = 50 μm).
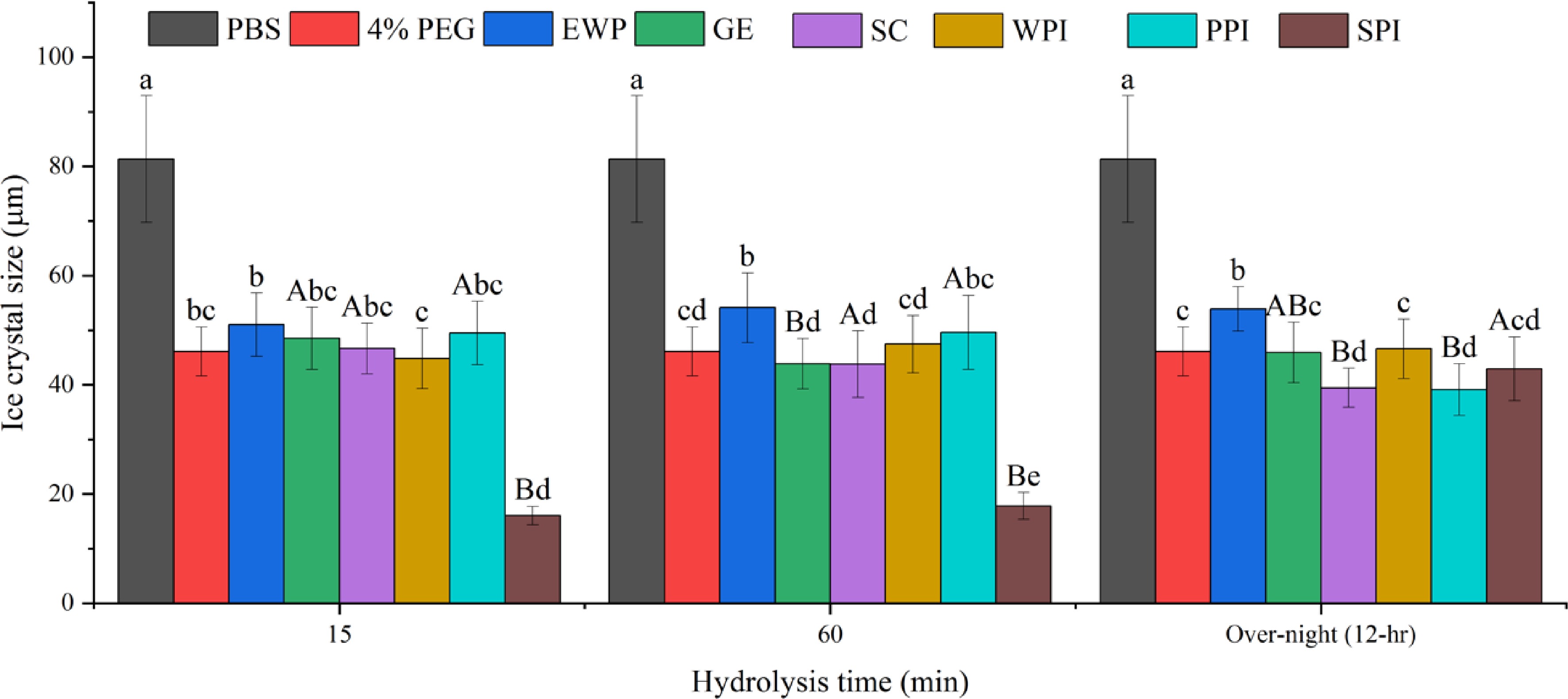
Figure 3.
Mean average ice crystal size (μm) from different protein hydrolysates (4% w/w) solutions under −8 °C. Same lower-case letter indicate no significant difference (p > 0.05) within the same hydrolysis time, while same upper-case letters indicate no significant difference within the same protein (p > 0.05).
As shown in Fig. 3, after 15- and 60-min hydrolysis, only SPI hydrolysates resulted in the formation of significantly (p < 0.05) smaller ice crystals compared to the negative control PEG and the solvent, PBS. However, after 12-hr hydrolysis, SPI-15 and SPI-60 hydrolysates at 4% (w/w) concentration lost its IRI activity. This is likely due to extensive hydrolysis of SPI resulting in a higher amount of small peptides and single amino acids which are not IRI active. As shown in Fig. 1e, the IRI active SPI hydrolysates (15- and 60-min hydrolysis), contained a large molecular fraction (66,000 – 6,500 Da), whereas the 12-hr hydrolyzed sample did not, which indicates that this large molecular fraction might have contributed to the IRI activity of the SPI-15 and SPI-60. In order to have an IRI activity, a molecule needs to bind to the ice surface through electrostatic interaction followed by structural realignment to optimally hydrogen bond with the oxygen-oxygen lattice on the ice surface, and then a partial nonpolar environment is ideally created by neighboring hydrophobic residues of the peptide[5]. Thus, it is critical for the hydrolyzed peptide to dynamically adapt its conformation when it approaches an ice surface and align its oxygen-containing groups with that of the ice surface to bind and inhibit the ice crystal growth[14]. Based on the current literature, peptides with IRI activity should have at least a molecular size ranging from 2 to 3.5 kDa[15,29]. The average molecular weights of SPI-15 and SPI-60 are within and close to this range as shown in Fig. 1. In this case, a single amino acid or small peptide (di- or tri-peptide) might not be efficient enough in inhibiting ice crystal growth.
After extensive hydrolysis (12-hr), SC and PPI hydrolysates resulted in statistically significant (p < 0.05) smaller ice crystal sizes compared to PEG (Fig. 3), although not as effective as SPI-15 and SPI-60. As shown in Fig. 1b & f, after 12-hr hydrolysis, large and medium sized peptide fractions were broken down to produce smaller peptide fractions with a significant increase in the small peptide and amino acids fraction (573–75 Da) indicating the IRI active peptide/amino acid, such as proline, might present in this fraction for SC and PPI. Primacella et al.[30] observed the gelation inhibition activity of proline in egg yolk upon freezing storage due to the formation of proline-water hydrogen bonding and shielding of the hydrophobic sites of proteins. Proline is also known as a natural cryoprotectant which is found in plants and marine invertebrates[31], and can be used in cryopreservation of red blood cells[32].
In addition, WPI-60 and SPI-60 shared very similar peptide molecular weight distribution with a comparable average molecular weight as shown in Fig. 1a & e. However, WPI-60 was found insignificant (p > 0.05) in ice crystal inhibition compared to 4% PEG (the negative control). This might be due to the differences in amino acid composition and peptide sequence of the hydrolyzed products. Studies have shown that threonine, asparagine, alanine, and proline were critical in ice binding for anti-freezing proteins (AFPs) and IRI active hydrolysates[2,14,16−20]. Table 1 indicates the four IRI contributing amino acid compositions of the six examined proteins. Overall, all six types of protein contain an adequate amount of these four IRI contributing amino acids with a total content of about 25% (w/w). Compared to WPI, SPI only contains a higher amount of asparagine but has a lower content of the other three amino acids. SPI and PPI have a similar content of these four amino acids, but PPI did not produce IRI active hydrolysates after 15- and 60-min enzymatic hydrolysis. Bovine bone gelatin is the most proline abundant among these six proteins, but did not exhibit an IRI activity in this study. This indicates the importance of the peptide sequence on its IRI activity. As discussed earlier, it is important for IRI active proteins/peptides to bind the ice surface via electrostatic interaction and structural realignments. For example, studies have found that the repeated tripeptide sequence of (Ala-Ala-Thr)n with a disaccharide (3-O-(β-D-galactosyl)-D-N-acetylgalactosamine) bonded to the β-oxygen of the threonine residue is essential for the IRI activity of AFPs[33,34]. Hence, an IRI active hydrolysate needs to not only contain the IRI active amino acids but also have them in the correct sequence in order to efficiently bind to the ice surface thus inhibiting ice crystal growth.
Amino acids Egg white protein Bovine bones gelatin Whey protein Casein protein Soy protein Pea protein Threonine 4.75 1.77 6.87 4.05 3.60 3.70 Alanine 8.93 8.69 5.55 2.76 4.10 4.40 Asparagine 10.08 4.20 9.18 7.57 11.60 11.50 Proline 4.25 12.54 6.66 9.33 5.60 5.00 Total 28.01 27.2 28.26 23.71 24.9 24.6 Overall, among the six types of protein, SPI hydrolysates, after 15- and 60-min enzymatic hydrolysis, demonstrate a great potential for further anti-freezing applications in food processing. While no significant difference (p > 0.05) in IRI activity was observed between SPI-15 and SPI-60 (Fig. 3), both samples were further evaluated under different annealing temperatures, time, and concentration for their IRI activity evaluation.
IRI activity of SPI-15 and SPI-60 under various annealing temperatures, times, and peptide concentration
-
To investigate the effect of temperature on ice crystal growth, the SPI-15 and SPI-60 samples (at 4% w/w) were examined for their IRI activity under different annealing temperatures ranging from −10 to −4 °C. Overall, SPI-60 was found to be less temperature resistant compared to SPI-15 as shown in Fig. 4a & b. SPI-15, at 4% (w/w) concentration, was able to inhibit the growth of ice crystals, with sizes smaller than 23 μm (vs. 37 to 48 μm for PEG), between −10 to −6 °C, whereas SPI-60 was only able to retard ice crystal growth at −8 °C or lower. At −6 ºC, SPI-15 was able to maintain the average ice crystal size around 22 µm at and beyond 20 min annealing, whereas ice crystal in SPI-60 system continued to grow until the end of the 30 min annealing, resulting in an average ice crystal size of 35.9 µm. When the temperature continued to increase to −4 °C, both SPI-15 and SPI-60 lost their IRI activity, thus demonstrating no significant difference (p > 0.05) in ice crystal size compared to PEG under the same annealing condition. At −4 °C, ice crystals in SPI-15 and SPI-60 (4% w/w) continued growing within 30 min annealing duration. The amount of IRI active peptides applied (4% w/w) was probably not enough to ensure the IRI activity of SPI-15 and SPI-60 at increased unfrozen water fractions and high annealing temperature. As discussed in the previous section, the increased hydrolysis time resulted in a reduction of high molecular fraction (65000-6600 Da) which could be responsible for the IRI activity of SPI hydrolysates (SPI-15 and SPI-60). The reduced amount of the IRI active fraction in SPI-60 led to its lower temperature-resistant capability compared to SPI-15.
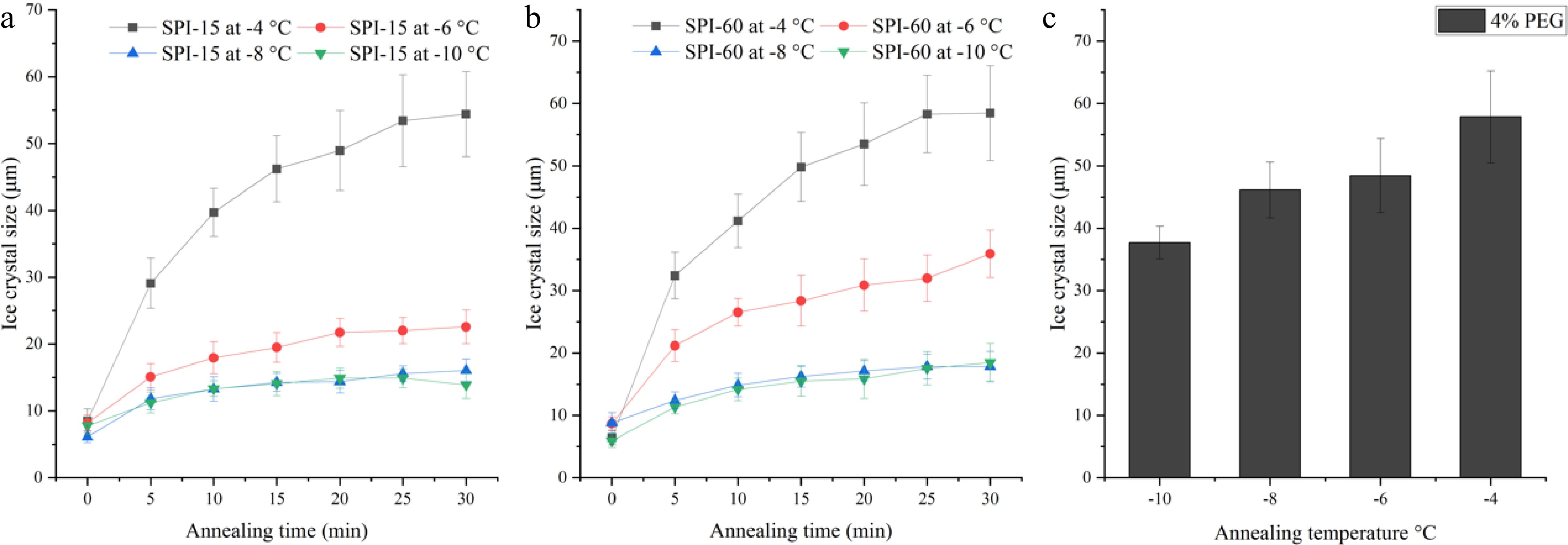
Figure 4.
Ice crystal size of 4% (a) SPI-15 and (b) SPI-60 hydrolysates in PBS buffer system annealed under −10 to −4 °C for various time. (c) Ice crystal size of 4% PEG (negative control) after 30 min annealing under various temperatures.
To evaluate the effect of peptide concentration on the IRI activity of SPI hydrolysates, different concentrations of SPI-15 (2%−4% w/w) were applied. As shown in Fig. 5, decreased concentration of SPI-15 resulted in a decreased temperature resistance, where SPI-15 at 2% (w/w) was not able to retard ice crystal growth even at −10 °C. In addition, SPI-15 at 3% (w/w) could not inhibit ice crystal growth at and beyond −6 °C compared to PEG and demonstrated lower effectiveness compared to 4% (w/w) SPI-15 under −10 and −8 °C, indicating the importance of the IRI active peptide content. Generally, in a commercial household freezer (−18 °C), the temperature is unlikely to fluctuate above −10 °C[35]. Thus, SPI-15, used at 3% or 4% (w/w), could be effective in withstanding temperature fluctuation and maintaining a small ice crystal for frozen products during storage and transportation.
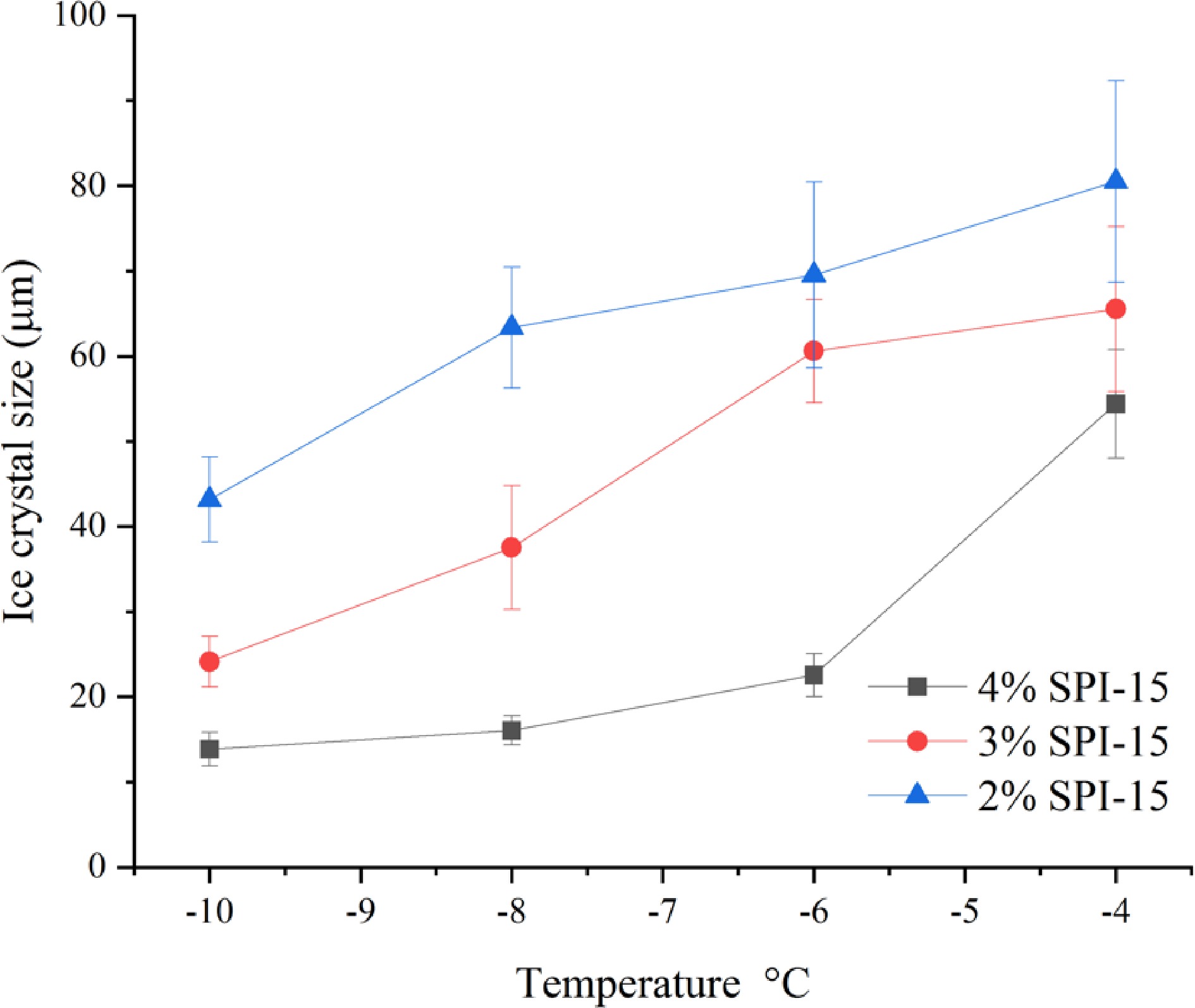
Figure 5.
Ice crystal size of SPI-15 hydrolysates in PBS buffer at varied concentrations (2%−4% w/w) annealed under different temperatures (−10 to −4 °C) for 30 min.
IRI activity of OSA-modified PPI and SPI hydrolysate after 12-hr hydrolysis
-
As discussed earlier, an enhanced IRI activity was observed in PPI hydrolysates after 12-hr enzymatic hydrolysis resulting in significantly smaller (p < 0.05) ice crystal size compared to PEG, whereas SPI hydrolysates lost their IRI activity after 12-hr hydrolysis. Hence, succinylation reaction was performed using OSA to modify SPI-12hr and PPI-12hr hydrolysates to evaluate the change in structure and amphiphilicity on the IRI activity of the modified samples. As shown in Table 2 and Fig. 6, both OSA-modified SPI-12hr and PPI-12hr, at 4% (w/w) concentration, demonstrated a significant increased IRI activity (p < 0.05) compared to their non-modified counterparts and the negative control (PEG) annealed at −8 °C. At −8 °C, the ice crystal size in OSA-modified hydrolysates was almost half of the size compared to the non-modified treatments. As the temperature increased to −6 °C, only OSA-SPI demonstrated a significant IRI activity compared to SPI-12hr and PEG (p < 0.05). This result supports the hypothesis on the significance of structural amphiphilicity in the IRI activity of AFPs which has one side hydrophilic for ice binding and on the other side hydrophobic feature preventing ice crystal growth[21,22]. Thus, the hydrophilic SPI and PPI hydrolysates modified with the hydrophobic octenyl succinic anhydride (OSA) through succinylation reaction had an enhanced amphiphilicity. Besides AFPs, synthetic polymers have been examined for their IRI activities with amphiphilicity[36]. Li et al.[37] have also observed the amphiphilicity of nanocelluloses with IRI activity.
Table 2. Ice crystal size of 4% legume protein (SPI and PPI) hydrolysates and their derivatives (OSA-modified) under annealing temperatures of −8 and −6 °C with 4% PEG as negative control.
Sample Ice crystal diameter
(μm) −8 °CIce crystal diameter
(μm) −6 °CPEG 46.13a 48.43a SPI-12hr 42.95b 48.45a OSA-SPI-12hr 16.93d 26.29b PPI-12hr 39.12c 49.37a OSA-PPI-12hr 17.47d 49.37a Same superscript indicates no significant difference (p > 0.05) within the same column. 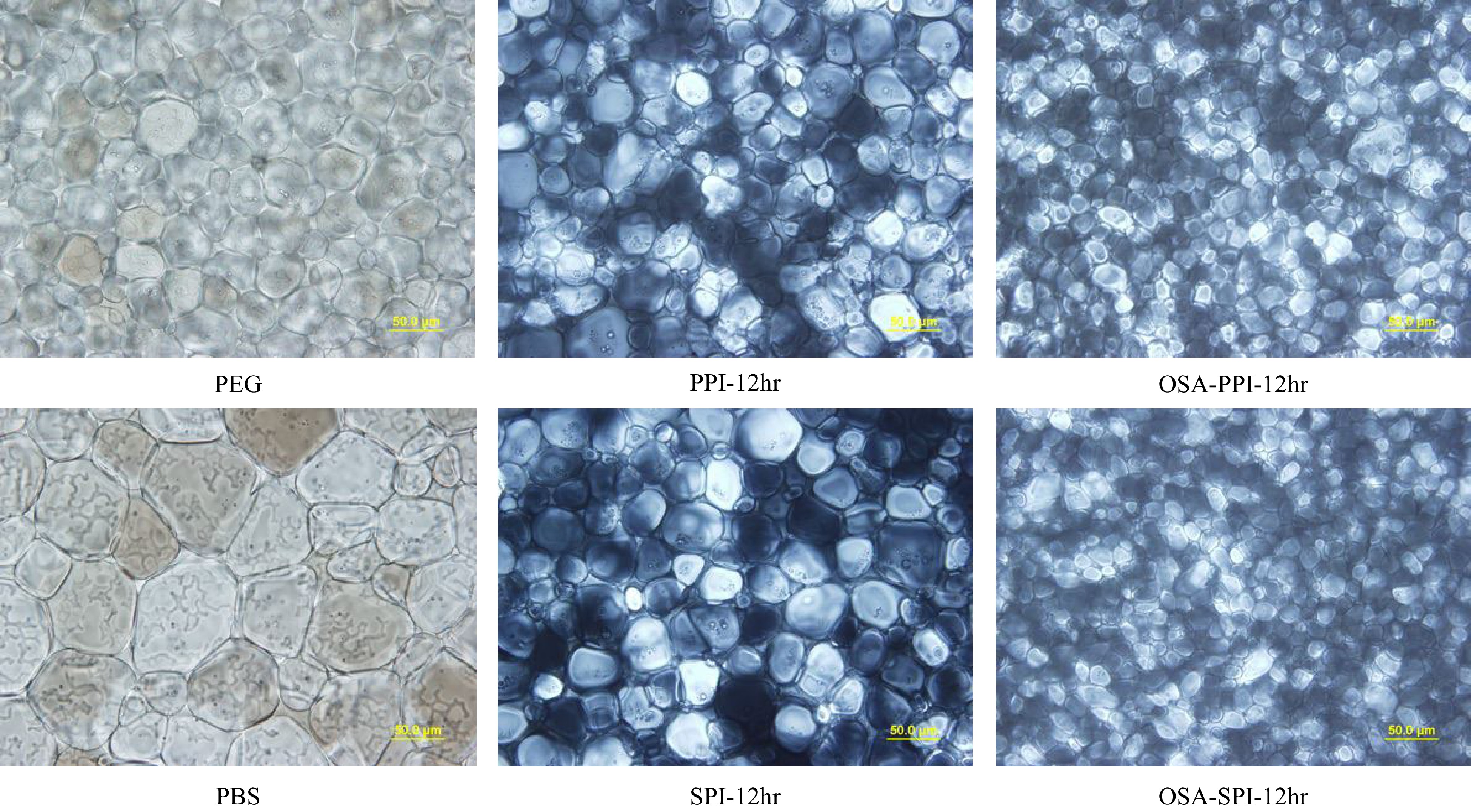
Figure 6.
Ice crystal image under polarized microscope at 40x for 4% SPI hydrolysates, PPI hydrolysates and their OSA-modified derivatives annealed at −8 °C compared to the negative control 4% PEG and PBS (the buffer solvent). (scale bar = 50 μm).
Figure 7 demonstrates the ice crystal growth of SPI hydrolysates and PPI hydrolysate with their OSA-modified derivatives suspended in PBS (4% w/w) within 30 min annealed at temperatures of −8 and −6 °C. For OSA-SPI-12hr, ice crystal size was stabilized after 15 min under both −8 and −6 °C, while ice crystal size in the non-modified SPI-12hr continued to grow beyond 15 min. While, OSA-PPI-12hr was only able to stabilize ice crystal growth at −8 °C at and beyond 15 min, it could not inhibit ice crystal growth when the temperature increased to −6 °C, indicating that the enhancement by the addition of OSA might not be enough to ensure the activity of OSA-PPI-12hr at increased fractions of unfrozen water at high annealing temperature. The ability of OSA-SPI-12hr in inhibiting ice crystal growth under increased temperature makes it a better agent in preventing ice recrystallization in frozen foods compared to OSA-PPI-12hr.
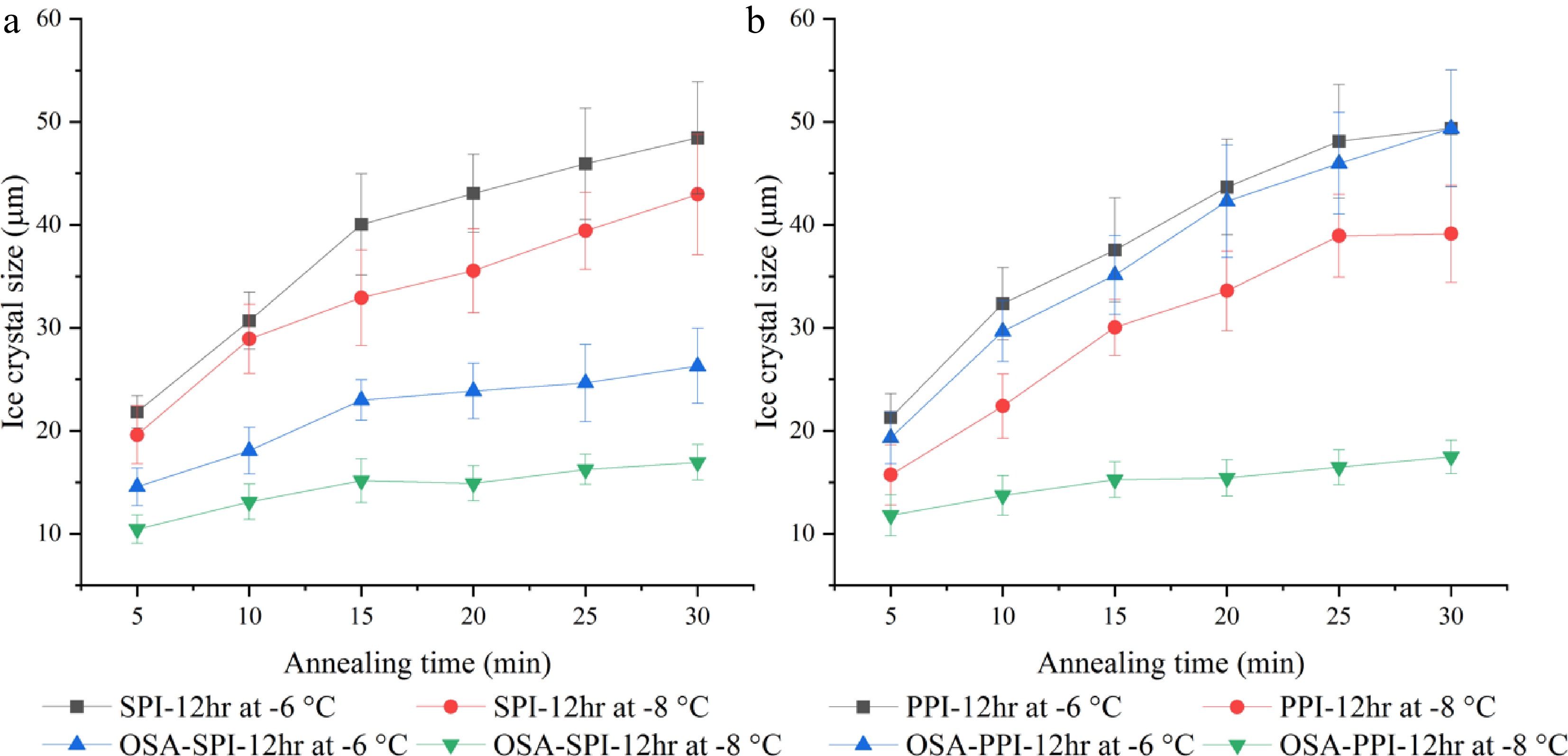
Figure 7.
Ice crystal growth within 30 min annealed at −8 °C and −6 °C in 4% (w/w) (a) SPI-12hr hydrolysate and OSA modified SPI-12hr, and (b) PPI-12hr hydrolysate and OSA modified PPI-12hr.
Surface hydrophobicity was also measured to assess the effect of OSA modification on the hydrophobicity of the SPI and PPI hydrolysates using the ANS fluorescent method. Table 3 shows the surface hydrophobicity of the modified and non-modified PPI and SPI hydrolysates. OSA-PPI-12hr was found to be significantly more hydrophobic (p < 0.05) compared to the other three samples in which no significant difference was observed (p > 0.05). As a fluorescent probe, ANS binds to the hydrophobic regions of proteins, thus leads to an increase in fluorescence, which is used as a measure of protein surface hydrophobicity[38]. In this case, the ANS method might not be sensitive enough to detect the changes by the modification, since protein hydrolysates were highly soluble in water and more hydrophilic than their native form, leading to the low surface hydrophobicity as measured by the ANS method even after the addition of OSA. In addition, the presence of carboxyl and carbonyl groups in OSA-modified samples might interfere with the ANS binding resulting in a low hydrophobicity[39]. Hence, in future studies, other methods should be identified to characterize the structural changes of the modified hydrolysates in order to understand the effect of OSA modifications on its IRI activity. Different degrees of modification will also be examined to optimize the modification process.
Table 3. Surface hydrophobicity of SPI and PPI hydrolysates and their OSA modified products.
Sample Surface hydrophobicity SPI-12hr 6685b OSA-SPI-12hr 6196b PPI-12hr 7123b OSA-PPI-12hr 9134a Same superscript indicates no significant difference (p > 0.05) among samples. -
Overall, this study has demonstrated the high potential of utilizing enzymatic hydrolysis to produce an IRI active hydrolysate from common food proteins. This will enable the utilization of cost-effective proteins for the production of IRI agents in a sustainable manner rather than using the high-cost and low-yield natural AFPs. Among the six proteins examined, SPI is found to be an effective protein for the production of IRI active hydrolysates. Its IRI activity is influenced by the annealing temperature and peptide concentration, where lower temperature and increased concentration result in enhanced IRI activity. The effectiveness of hydrolysates' IRI activity could be attributed to their amino acid composition and sequences. Moreover, chemical modification of SPI and PPI hydrolysates by the addition of OSA was found to enhance its IRI activity. OSA-SPI-12hr demonstrated better temperature resistance compared to OSA-PPI-12hr. Future studies will further evaluate the effect of degree of OSA modification on the IRI activity and characterize the structural changes of the modified hydrolysates.
-
All protein samples, sodium caseinate (SC; Sigma-Aldrich, MO, USA), whey protein isolate (WPI), egg white protein (EWP; prepared from freeze-dried egg white), pea protein isolate (PPI; BulkSuppliments.com, NV, USA), soy protein isolate (SPI; BulkSuppliments.com, NV), and commercial gelatin (GE; Kraft Foods, IL, USA), were adjusted to a standard moisture content before the hydrolysis treatment. Enzymatic hydrolysis of protein samples by alcalase was performed based on methods established in previous studies with some modifications[2,14]. Protein dispersion of 10% (w/w) was hydrolyzed at 45 °C for up to 60 min at pH 9.0 with 0.176 Anson unit enzyme/g protein added. Extensive, 12-hr, hydrolysis was also performed to evaluate the effect of enzymatic hydrolysis on the anti-freezing properties of the hydrolysates. After hydrolysis, enzyme was inactivated by incubating the solution for 10 min in boiling water followed by centrifugation at 10,000 x g for 10 min to obtain the supernatant of peptide products. The supernatants were collected and freeze-dried for ice recrystallization inhibition (IRI) activity measurement.
Ice recrystallization inhibition (IRI) assay
-
A standard splat assay was used to assess the IRI activity of the hydrolyzed peptides. For the assay[40], a 10 µl droplet of sample solution, with various concentrations (2%– 4% w/w), was dropped from a height of 1.5 m onto a glass slide surface that is precooled to −80 °C. The sample was then annealed under different temperatures between −10 °C and −4 °C for up to 30 min using a cryo-stage HCS 302 (Instec Instruments, Boulder, CO, USA) before collecting images using polarized light microscopy (Olympus BX51, Tokyo, Japan) with a built-in digital camera (DP 70, Olympus, Tokyo, Japan) to determine the size of the ice crystals and examine the effect of annealing temperature on IRI activity of each peptide. The size of the 10 largest ice crystals in the image was measured using Image J (National Institutes of Health, Bethesda, MD, USA). The average mean size of the ice crystals of each sample was calculated from three independent splat assay measurements. Polyethylene glycol (PEG) was used as a negative control[37,41].
Peptide molecular weight distribution
-
The molecular weight profiles of the hydrolyzed samples were analyzed by size exclusion chromatography on an Agilent HPLC system (Agilent Technologies, Santa Clara, CA, USA). The separation was performed on a BioSep-SEC-S2000 column (300 × 7.80 mm, Torrance, CA, USA) as described by Price et al.[42]. In brief, the mobile phase was 45% acetonitrile in water with 0.1% trifluoroacetic acid at a flow rate of 1.0 mL/min. The sample was injected at 20 µL with 20 min run time detected at 214 nm at ambient temperature. The elution profile was compared to the standards with known molecular weights (albumin, glycine, and a mixed peptide standard) to determine molecular size distribution of the hydrolysates. The average peptide size of the hydrolyzed sample was calculated as the sum of the multiplication of the mean molecular size and percentage of each molecular range obtained via HPLC analysis.
Modification of PPI and SPI hydrolysates through succinylation reaction
-
Modification of hydrolysates legume proteins was performed based on methods established in the previous studies with minor modifications[43,44]. Octenyl succinic anhydride (OSA) has been used to react with all nucleophilic groups, including amine groups and aliphatic hydroxyl groups[45,46]. Hydrolysate samples, 2 g, were dispersed in 250 mL 0.075 M phosphate buffer (pH 8). OSA was added to hydrolysates, with constant stirring over a 3-hr period at levels of 40% of the weight of the hydrolysate sample. During the reaction, pH was maintained at 8–8.1 with 2 N NaOH. After 3-hr reaction, pH of the reaction solution was adjusted to 6.5 followed by centrifugation to obtain derived protein and remove any insoluble compounds prior to dialysis. Modified hydrolysates then underwent dialysis with 100-500 MWCO membrane in deionized water for 3 d under refrigerated conditions (4 °C) to remove any unreacted OSA, the deionized water was changed daily. After dialysis, the modified hydrolysates were freeze-dried for IRI activity and surface hydrophobicity analysis.
Surface hydrophobicity
-
Surface hydrophobicity was determined using 1-phenylamino-2-napthalenesulfonic acid (ANS) as fluorescent probe as previously reported[47]. Briefly, 200 µL of different concentrations of legume proteins and derivatives (0.005 to 0.5 mg protein/mL in phosphate buffer, pH 7.4) was plated in 96-well black-walled microplate and 10 µL ANS probe (0.8 mM in phosphate buffer, pH 7.4) was added. The mixture was incubated in the dark for 10 min and fluorescence intensity was measured at excitation and emission wavelengths of 390 and 480 nm, respectively. The fluorescence intensity was plotted against protein concentration, and the slope of the linear regression curve was reported as surface hydrophobicity.
Statistical analysis
-
All experiments and measurements were performed in duplicates. Statistical analysis was carried out using the JMP statistical package. Analysis of variance was used, and the data were represented as mean value ± standard deviation. Tukey's multiple sample comparison tests were carried out to assess the significance of the difference between treatments. Statistical significance was indicated at a confidence level of 95% (p ≤ 0.05).
This study was supported by the Hatch Multi-State Project (Accession # 1023982).
-
The authors declare that they have no conflict of interest.
- Copyright: © 2022 by the author(s). Published by Maximum Academic Press on behalf of Nanjing Agricultural University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Wan Z, Fei T, Wang T. 2022. Inhibition of ice crystal growth by protein hydrolysates from different plant- and animal-based proteins. Food Materials Research 2:17 doi: 10.48130/FMR-2022-0017
Inhibition of ice crystal growth by protein hydrolysates from different plant- and animal-based proteins
- Received: 05 September 2022
- Accepted: 01 November 2022
- Published online: 24 November 2022
Abstract: Due to the quality deterioration and the corresponding waste of frozen foods, there is a constant need for an effective, consumer and environmentally friendly, and sustainable anti-freezing ingredient. In this study, enzymatic hydrolysis and food grade chemical modification (succinylation) were evaluated to produce ice re-crystallization inhibition (IRI) active peptide-based ingredients. Six different types of plant- and animal-based proteins were evaluated. Soy protein isolate (SPI), after 15- and 60-min alcalase hydrolysis, demonstrated a significant IRI activity compared to polyethylene glycol (PEG; as negative control) resulting in a two-third reduction of ice crystal size annealed at −8 °C. Annealing temperature and peptide concentration also affected the IRI activity of SPI hydrolysate where decreased temperature and increased concentration resulted in a higher IRI activity. Increased hydrolysis time decreased the IRI activity of SPI hydrolysates but increased the IRI activity of sodium caseinate (SC) and pea protein isolate (PPI) hydrolysates. SPI-60 was less temperature resistant compared to SPI-15 which inhibited the growth of ice crystals (smaller than 23 µm). Moreover, chemical modification via succinylation reaction with octenyl succinic anhydride (OSA) enhanced the IRI activity of the PPI and SPI hydrolysates leading to a 55.3% and 60.6% reduction in the size of ice crystals, respectively. Overall, this study demonstrates a great potential in the utilization of common food proteins for the production of value-added natural anti-freezing ingredients which could benefit the food industry by enhancing frozen foods' shelf-life and reducing waste.
-
Key words:
- protein hydrolysates /
- ice recrystallization inhibition /
- succinylation.