-
Fungi represent the second largest group of eukaryotic organisms on earth, with estimates ranging from 1.5 to 5.1 million species[1]. Fungal fruiting bodies (often associated with basidiomycetes and called mushrooms) of macroscopic, filamentous, and epigeal fungi have structures called filamentous hyphae, which form interwoven web of tissue that is known as mycelium[2]. In the substrate where the fungus obtains its nutrition, most often, their mycelia are buried and usually found growing on fallen logs, decomposing piles of straw, lawns, beneath leaf litter, meadows, and gardens[3]. Accordingly, there are over 150,000 different types of mushrooms, but only 10% of which are known and named. The Philippines is rich with mycological resources that have promising pharmacological and nutraceutical potentials. These mycological resources are proliferating on different substrates at different geographical locations across the archipelago[3]. The growing interest of researchers in the field paved the way for the continued efforts and research works in the discovery of their potential use that would benefit society as a whole. InteresHtingly, as Filipinos are becoming acquainted with the good taste and health benefits of consuming mushrooms and other mushroom products, mushroom production in the Philippines is progressively improving[4].
Interestingly, aside from basidiomycetes, there are also several groups of fungi that produce considerable macroscopic fruiting bodies, as in the case of some members belonging to ascomycetes. One of which is the genus Xylaria. The genus Xylaria Hill ex Schrank (1786) is recognized as the largest and the first described genus under the family Xylariaceae Tul. & C. Tul. (1861), in which most members of this genus are found in the tropics and subtropics[5]. Xylaria species are saprophytic, may be weakly to strongly pathogenic to plants, and endophytic in nature which appears to exist in live plant tissues, decayed wood, dung, leaf litter, and soil[6,7].
In light of the discovery of various fungal species, it is important to determine their nutraceutical and pharmacological benefits. Hence, it is then necessary to optimize their growth conditions. Nowadays, there is a growing interest in mycelial production for nutraceutical or pharmacological purposes due to the promising potential various fungal species can offer. However, the production or cultivation of fungal fruiting bodies is not practical, for it is time-consuming to harvest fungal fruiting bodies for commercial purposes. The use of submerged culture conditions in mycelial biomass production has sparked attention for its number of advantages, including higher mycelial yield in a more compact space and shorter time, with fewer chances of contamination[8]. Aside from that, several researchers also stated that various compounds with significant biological activities could diffuse and can be derived from the culture medium itself[9,10].
Available research indicates that several optimization and functional studies have been carried out in various Xylaria species however, many species in this genus still remain unexplored, like Xylaria papulis Lloyd; hence, this study was conceptualized. This study evaluated the optimum cultural conditions of Xylaria papulis Lloyd in indigenous culture conditions making use of readily available raw materials such as rice bran, coconut water, potato, and corn grit in consideration of physical factors such as pH, agitation, temperature, and illumination conditions. Evaluation of which is important for the purpose of mass producing the mycelia of Xylaria papulis Lloyd locally in the attempts of producing available mycelia as a source of bioactive compounds with a wide range of benefits and potential.
-
Pure culture of X. papulis Lloyd (Sample_ID_4/GU300100.1) was obtained from the Tuklas Lunas Center and the Center for Tropical Mushroom Research and Development (CTMRD), Department of Biological Sciences, College of Arts and Sciences, Central Luzon State University (CLSU), Science City of Muñoz, Nueva Ecija, Philippines. The identity of X. papulis Lloyd used in this study was ascertained and confirmed molecularly using ITS nrDNA by Undan et al.[11] The said fungus was subcultured using potato dextrose agar.
Preparation of Potato Dextrose Agar (PDA) and mycelial inoculants
-
PDA was prepared by dissolving 39 g of commercially available PDA in 1 L of distilled water. Forty mL (1/3 of the volume capacity of the flat bottle used) of the prepared medium was dispensed in a clean flat bottle. It was covered with a cotton plug and wrapped with clean recycled paper secured with a rubber band. The prepared media were sterilized in an autoclave at 100 °C and 121 psi for 30 min. After sterilization, the bottled media were cooled in a slanted position to solidify.
Culture inoculants were prepared by aseptically inoculating agar blocks from the pure culture of X. papulis Lloyd into prepared PDA plates. The culture plates were incubated at room temperature to allow mycelial growth. After seven days of incubation, mycelial discs were taken using a sterilized 10-mm diameter cork borer which served as the source of inoculants for the evaluation of optimum submerged culture conditions.
Evaluation of culture media
-
The mycelial biomass production in different indigenous liquid media broths was evaluated. The treatments were designated as follows: T1 = coconut water from mature coconut fruit (Cocos nucifera); T2 = rice bran decoction broth (50 g of Oryza sativa bran in 1 L of H2O); T3 = local yellow corn grit broth (50 g of Zea mays cracklings in 1 L of H2O); and T4 = potato sucrose broth (200 g of Solanum tuberosum + 10 g of table sugar in 1 L of H2O) following the methods of Dulay et al.[12] with slight modifications.
The culture broths were adjusted to pH 6.5, and a volume of 100 mL was dispensed in each catsup bottle which served as the culture container, followed by sterilization at 100 °C, 121 psi for 30 min. After cooling, each medium was aseptically inoculated with a 10-mm mycelial disc and was incubated at room temperature to allow fungal growth. After 10 d of incubation, the mycelia were harvested, and the mycelial thickness, as well as the mycelial dry weight, were measured to determine the most favorable broth medium for mycelial biomass production. The experiment was carried out using five replicates. The liquid medium that yielded the highest mycelial biomass was evaluated for different pH levels, agitation conditions, temperatures, and illumination conditions.
Evaluation for pH of optimum liquid medium
-
The optimum liquid culture medium was used in the evaluation for optimum pH. The pH of the liquid medium was adjusted using 1 M NaOH and 1 M HCl. The varying pH levels evaluated were from pH 5.0 to pH 8.5 with an interval of 0.5 (T1 = 5.0, T2 = 5.5, T3 = 6.0, T4 = 6.5, T5 = 7.0, T6 = 7.5, T7 = 8.0, and T8 = 8.5). One hundred mL of the optimum broth medium with the specific pH was dispensed in a culture container and autoclaved at 100 °C, 121 psi for 30 min. After cooling, it was inoculated with a 10-mm mycelial disc and incubated for 10 d under the optimum liquid culture conditions. The experiment was carried out using five replicates. Mycelial biomasses were harvested, and the dry weights were recorded to determine the most favorable pH. Likewise, the mycelial thickness was also recorded.
Evaluation for agitation conditions
-
The optimum liquid culture medium and the best pH level were used in optimizing agitation conditions. One hundred mL of the favorable culture broth was dispensed in a culture container that was sterilized at 100 °C, 121 psi for 30 min, and cooled prior to the inoculation with mycelia. It was then incubated at room temperature while the shaking condition was incubated in a shaker at 70 rpm. The experiment was carried out using five replicates. After 10 d, the mycelial thickness was recorded. Likewise, the dry mycelial biomass was also weighed.
Evaluation for temperature
-
In the evaluation for the optimum temperature, the mycelial disc was inoculated in the best culture medium and pH level, incubated at optimum agitation and at different temperature conditions such as room temperature (28 °C), air-conditioned (23 °C), and refrigerated (10 °C). The experiment was carried out using five replicates, and after 10 d, the mycelial biomasses were harvested, and the dry weight was recorded. Likewise, the mycelial thickness was also measured using a vernier caliper.
Evaluation for illumination
-
In the evaluation for illumination, the mycelial disc was inoculated in the best culture medium and optimum pH level incubated in a static condition at optimum temperature and at different illumination conditions namely: i) continued lighting; ii) alternating 12-h dark and 12-h light; and iii) totally dark. Under continued lighting conditions, the inoculated culture containers were incubated in a chamber with artificial light (137 lux) at room temperature. For the dark condition, inoculated culture containers were covered with clean black paper and incubated at room temperature. In the alternating light and dark condition, cultures were covered with clean black paper for 12 h and then manually removed to facilitate lit conditions for 12 h. The experiment was carried out using five replicates, and after 10 d, the mycelial biomasses were harvested, and the dry weight was recorded. Likewise, the mycelial thickness was also measured. The sequential evaluation of optimum indigenous liquid culture conditions of Xylaria papulis Lloyd is shown in Fig. 1.
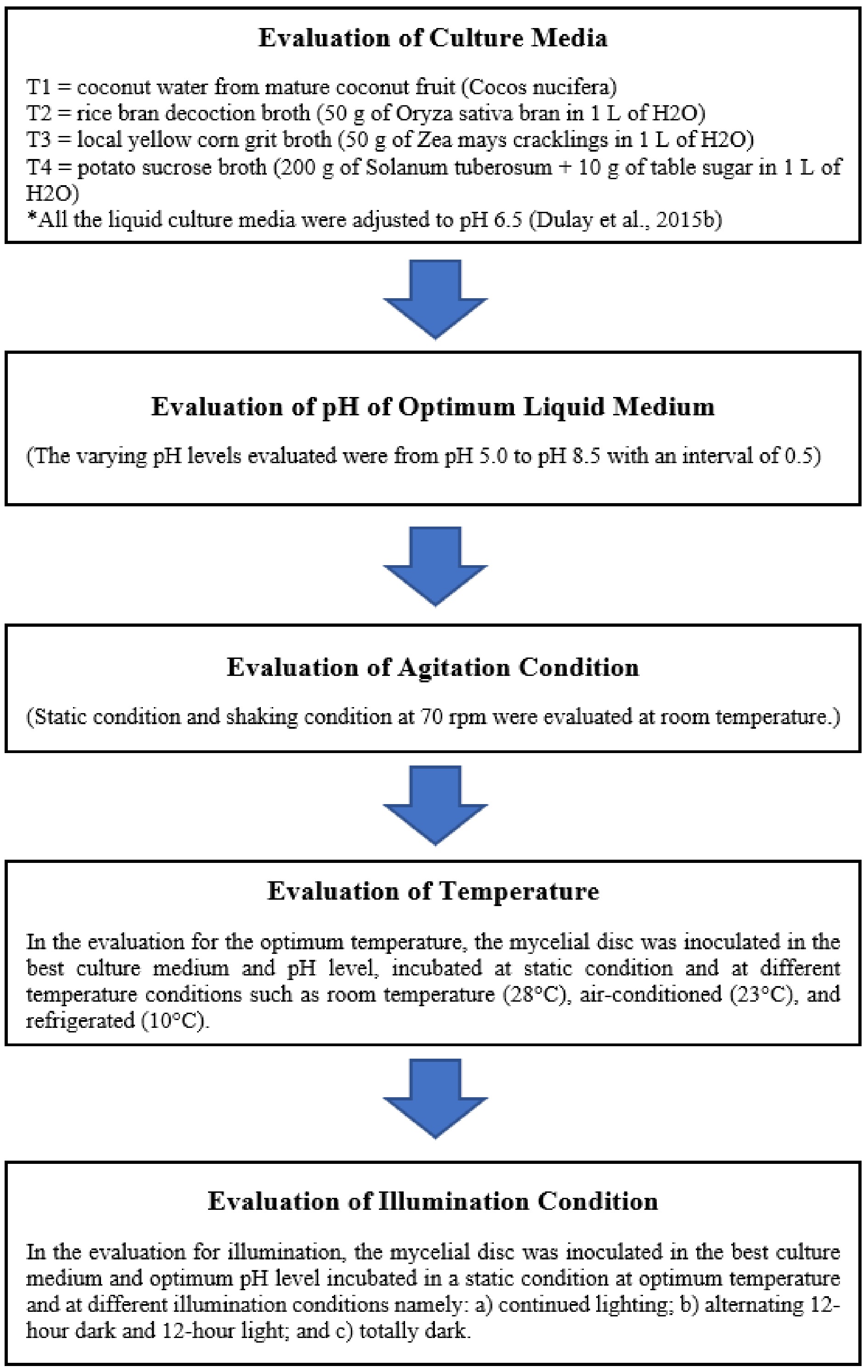
Figure 1.
Flow chart showing the step-by-step procedure done in the evaluation of optimum culture conditions for the mycelial growth of Xylaria papulis Lloyd.
Statistical analyses
-
The Minitab ver. 16 statistical software was used for statistical analyses of data. The experiments were laid out in a completely randomized design (CRD). Analysis of variance (ANOVA) was used, and treatments that were declared significant by ANOVA were further compared using Tukey’s comparison of means. Likewise, the T-test was used in the statistical analysis for the optimization of agitation conditions.
-
Four indigenous media, namely CW, CGB, RBB, and PSB, were used in the evaluation of culture media. After 10 d of incubation, the mycelial thickness was recorded. In this study, the dry mycelial weight was recorded after the mycelia were air-dried for three days through a desiccator. Table 1 presents the mean mycelial thickness and mean mycelial dry weight of the mycelial biomass of X. papulis Lloyd after 10 d of incubation in various indigenous liquid culture media. It can be inferred from Table 1 that CW exhibited the lowest mycelial thickness with a mean value of 1.02 mm, while PSB recorded the highest mean value of 2.31 mm. However, the latter is not significantly different from RBB, with a mean value of 1.70 mm. Interestingly, in the mean dry mycelial weight, PSB showed the highest mean value of 814.80 mg, while CGB had the lowest mean value of 88.00 mg. The result of mean dry mycelial weight is more reliable as an indication of growth than using fresh mycelial weight since water is eliminated from the sample. Based on the results of the present study, the PSB is the most suitable medium for the production of mycelial biomass since this liquid culture medium yielded the highest mycelial biomass. Moreover, fruiting bodies were observed growing in the PSB culture medium, as shown in Fig. 2.
Table 1. Mycelial growth performance of X. papulis Lloyd after 10 d of incubation in different indigenous liquid culture media.
Culture media Mycelial thickness*
(mm)Dry mycelial weight*
(mg)Coconut Water (CW) 1.02 ± 0.40c 315.00 ± 22.60b Corn Grit Broth (CGB) 1.42 ± 0.12bc 88.00 ± 55.40c Rice Bran Broth (RBB) 1.70 ± 0.51ab 345.10 ± 24.70b Potato Sucrose Broth (PSB) 2.31 ± 0.22a 814.80 ± 57.60a * Values are mean ± SD. Means in a column that do not share a letter are significantly different at a 5% level of significance using Tukey's Comparison of Means.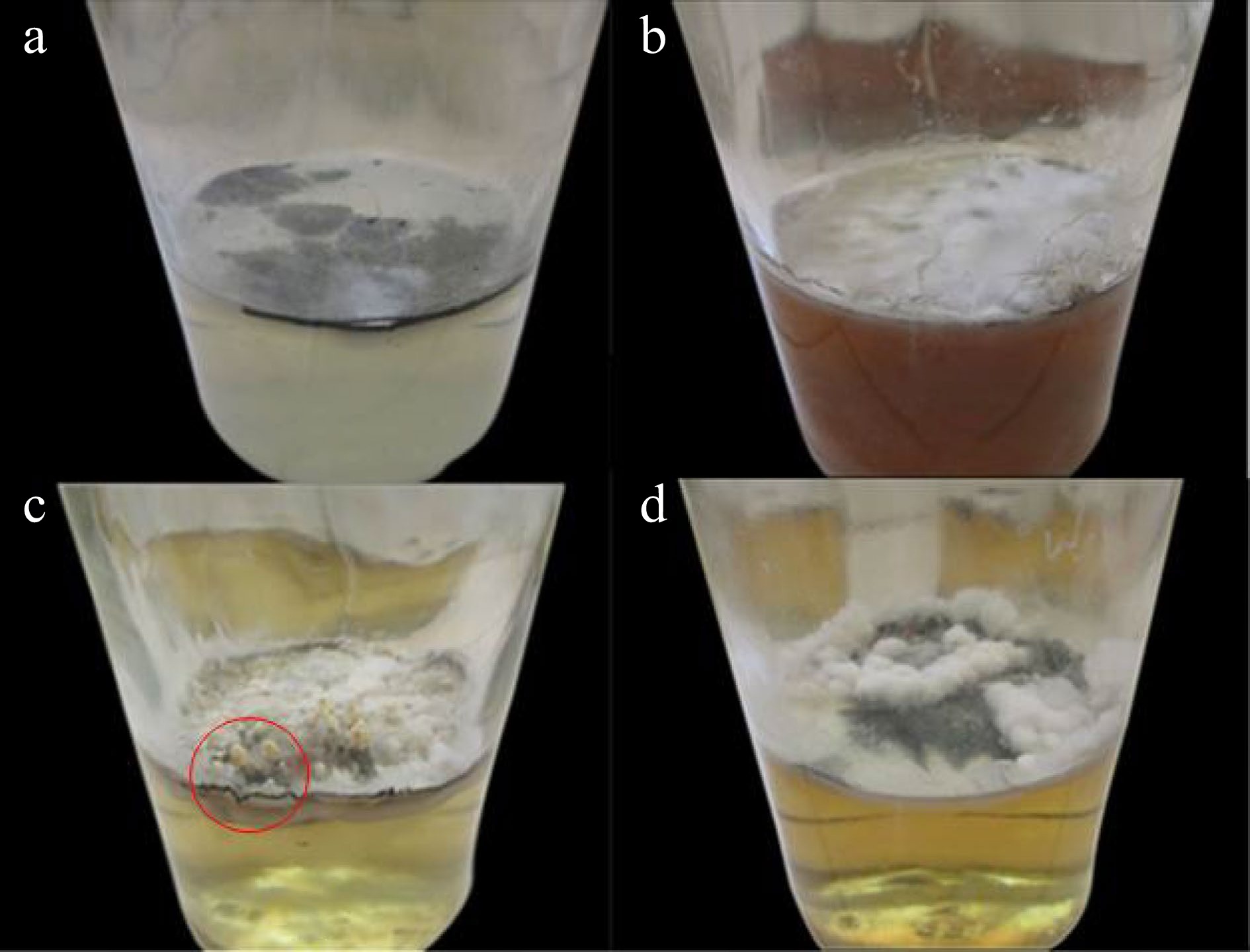 
Figure 2.
Mycelia of X. papulis Lloyd produced from (a) Corn Grit Broth, (b) Rice Bran Broth, (c) Potato Sucrose Broth (sprouting young fruiting bodies encircled), and (d) Coconut Water.
Evaluation for pH of optimum liquid medium
-
The optimum pH level is a critical factor associated with the growth of fungi because it will affect the cell membrane function, cell morphology and structure, the solubility of salts, the ionic state of substrates, the uptake of various nutrients and product biosynthesis[7,13]. In the evaluation for pH, PSB with pH levels ranging from 5.0 to 8.5 with an interval of 0.5 was investigated. Table 2 presents the mean mycelial thickness and mean mycelial dry weight of the mycelial biomass of X. papulis Lloyd after 10 d of incubation at different pH levels of potato sucrose broth. The mean mycelial thickness at different pH levels varies significantly from each other such that pH 6.0 recorded the highest mean value of 2.53 mm while pH 8.5 exhibited the lowest mean value of 1.07 mm. However, the former is comparable to pH 6.5, with a mean value of 1.96 mm. In terms of mean dry mycelial weight, pH 6.5 showed the highest mean value of 676.10 mg, while pH 8.5 had the lowest mean value of 242.52 mg. Comparison among means showed that pH 6.5 is not significantly different from pH 6.0 and pH 5.5; however, the latter is statistically comparable to pH 7.0. This result suggests that pH 5.5−6.5 is the optimum pH range for mycelial growth, although it can grow under a wide range of pH, as shown in Fig. 3.
Table 2. Mycelial growth performance of X. papulis Lloyd in optimum liquid medium (PSB) at different pH levels.
pH level (PSB) Mycelial thickness* (mm) Dry mycelial eeight* (mg) 5.0 1.51 ± 0.38bc 431.20 ± 26.50c 5.5 1.68 ± 0.43b 584.00 ± 57.30ab 6.0 2.53 ± 0.44a 651.10 ± 26.40a 6.5 1.96 ± 0.63ab 676.10 ± 39.10a 7.0 1.79 ± 0.67b 506.00 ± 76.00bc 7.5 1.78 ± 0.50b 458.00 ± 10.77c 8.0 1.57 ± 0.43bc 414.30 ± 21.30c 8.5 1.07 ± 0.52c 242.52 ± 62.40d * Values are mean ± SD. Means in a column that do not share a letter are significantly different at a 5% level of significance using Tukey's Comparison of Means.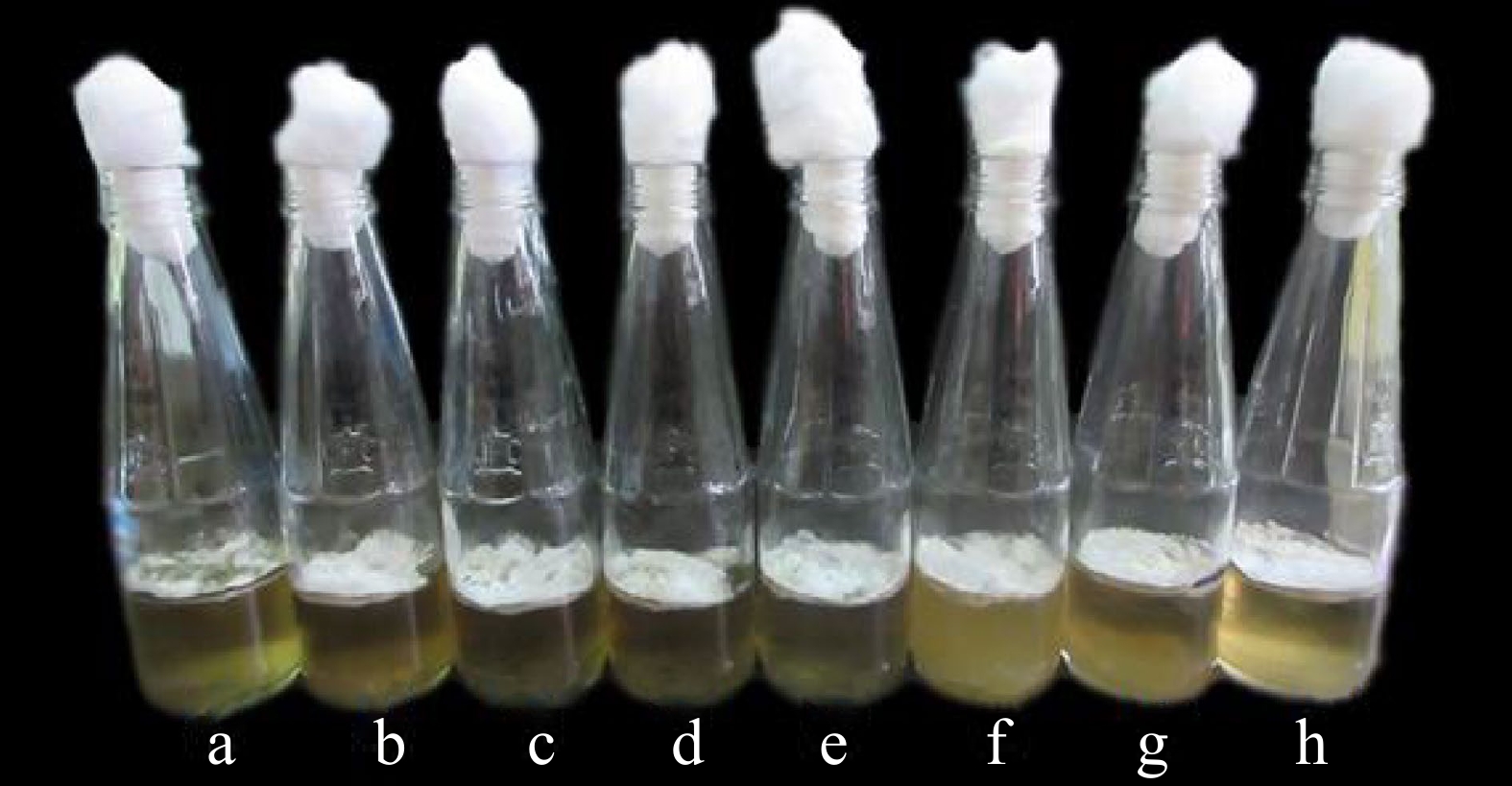 
Figure 3.
Mycelia of X. papulis Lloyd produced from different pH levels of PSB such as (a) pH 8.5, (b) pH 8.0, (c) pH 7.5, (d) pH 7.0, (e) pH 6.5, (f) pH 6.0, (g) pH 5.0, and (h) pH 5.5 after 10 d of incubation.
Evaluation of agitation condition
-
Agitation is an important factor making shear forces that affect fungal morphology, variation in mycelial growth, fragmentation of mycelia, and product formation or biosynthesis[14]. It also affects aeration, mass, and heat transfer, as well as the mixing of nutrients in the liquid medium[10]. In this study, the influence of agitation on mycelial growth was evaluated by inoculating a 10-mm mycelial disc in flasks containing PSB with a pH of 6.5 and incubated in static and agitated conditions. In agitated condition, the flasks were placed in a shaker set at 70 rpm. After 10 d of incubation, the mycelial thickness was recorded. Likewise, dry mycelial weight was recorded after the mycelia were air-dried for three days. Table 3 presents the mean mycelial thickness and mean dry mycelial weight after 10 d of incubation in static and agitated conditions using PSB and pH of 6.5. As seen in Table 3, the mean mycelial thickness is higher in static condition (1.94 mm) compared to shaking condition (0.93 mm). Moreover, mean dry mycelial weight in static condition (381.50 mg) is higher compared to shaking condition (233.90 mg). In this regard, static condition is better compared to shaking condition for mycelial growth since it yielded higher mycelial biomass as clearly shown in Fig. 4.
Table 3. Mycelial growth performance of X. papulis Lloyd on different agitation conditions after 10 d of incubation.
Agitation condition Mycelial thickness* (mm) Dry mycelial weight* (mg) Shaking (70 rpm) 0.93 ± 0.45b 233.90 ± 53.4b Static 1.94 ± 0.72a 381.50 ± 65.0a * Values are mean ± SD. Means in a column that do not share a letter are significantly different at 5% level of significance using t-test.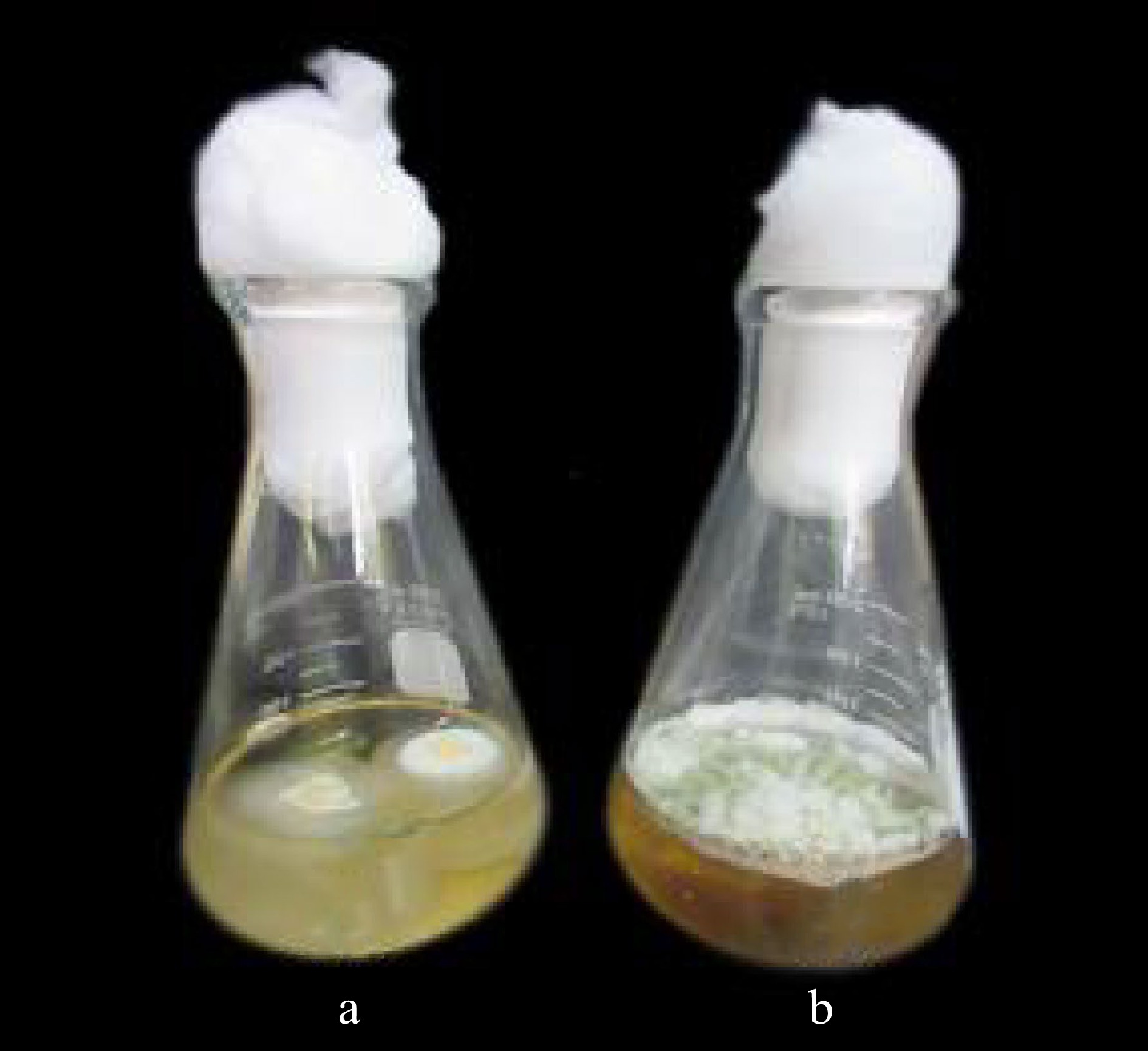 
Figure 4.
Mycelial biomass of X. papulis Lloyd produced after 10 d from shaking condition at (a) 70 rpm and (b) static condition.
Evaluation of temperature
-
The influence of temperature on mycelial growth was determined by inoculating 10-mm mycelial discs in PSB with pH of 6.5 incubated in static condition at different temperature conditions such as refrigerated (10 °C), airconditioned (23 °C) and room temperature (28 °C). After 10 d of incubation, the mycelial thickness was recorded. Likewise, dry mycelial weight was recorded after the mycelia were air-dried for three days. Table 4 presents the mean mycelial thickness and mean dry mycelial weight after 10 d of incubation in different temperature conditions using potato sucrose broth at pH of 6.5. The X. papulis Lloyd incubated at room temperature produced the highest mycelial thickness and mycelial dry weight with means of 1.99 mm and 320.00 mg, respectively. As expected, it is noted that no mycelial growth was observed in the refrigerated condition. Comparison among means revealed no significant difference between room temperature and air-conditioned temperature both in terms of mean mycelial thickness and mean dry mycelial weight. This result suggests the X. papulis Lloyd can be incubated in either condition to attain the optimum mycelial biomass production. Figure 5 shows the mycelial biomass produced from the different temperature conditions and static conditions using potato sucrose broth and pH of 6.5 after 10 d of incubation.
Table 4. Mycelial growth performance of X. papulis Lloyd in different temperature conditions after 10 d of incubation.
Temperature condition Mycelial thickness* (mm) Dry mycelial weight* (mg) Refrigerated (10 °C) 0.00 ± 0.00b 0.00 ± 0.00b Airconditioned (23 °C) 1.80 ± 0.54a 318.00 ± 47.60a Room temperature (28 °C) 1.99 ± 0.79a 320.00 ± 36.70a * Values are mean ± SD. Means in a column that do not share a letter are significantly different at 5% level of significance using Tukey's Comparison of Means.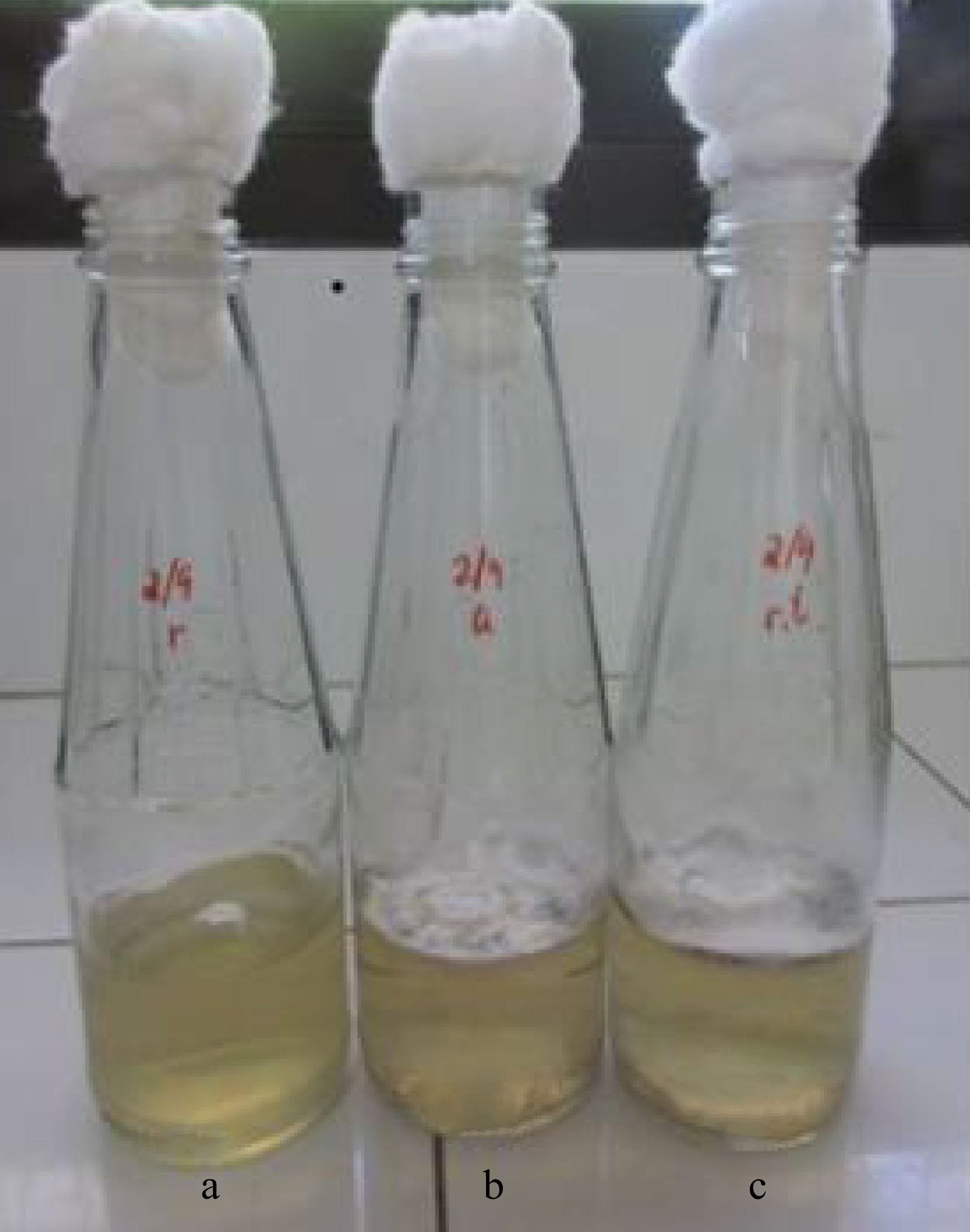 
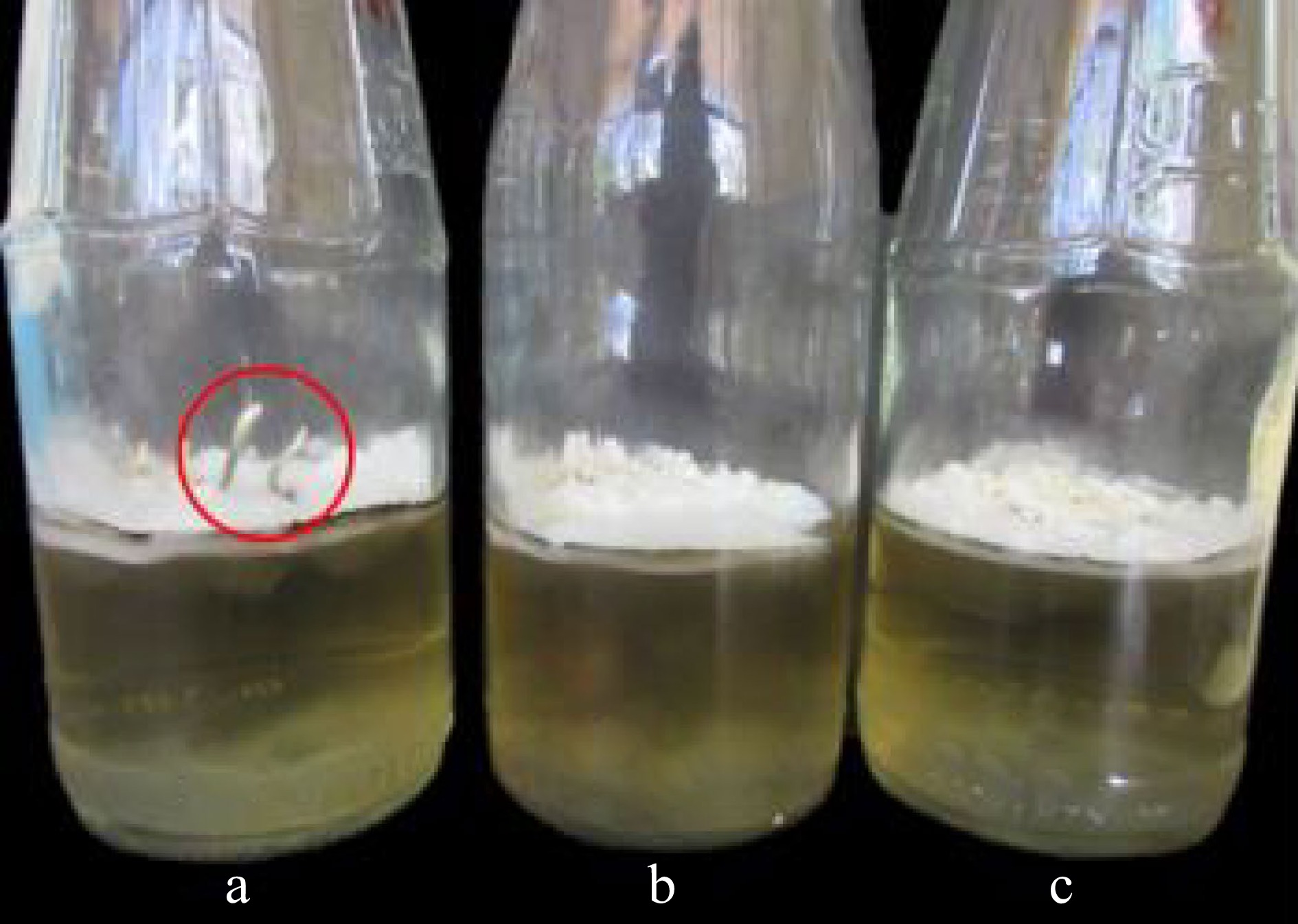
Figure 5.
Mycelia of X. papulis Lloyd produced after 10 d of incubation at different temperature conditions such as (a) refrigerated at 10 °C, (b) airconditioned at 23 °C, and (c) room temperature at 28 °C.
Evaluation of illumination
-
Statistical analysis revealed that there is no significant effect of various illumination conditions (lighted, alternating light and dark, and dark) on mycelial dry weights although in terms of mycelial thickness the lit condition (1.99 mm) is significantly higher compared with both light and dark as well as dark condition with means of 1.25 mm and 1.17 mm respectively (Table 5). The highest mycelial thickness and dry mycelial weight were observed in lit condition, on the other hand, those incubated in dark condition produced the lowest mean mycelial thickness and mean mycelial dry weight. Moreover, it is noteworthy to consider the fruiting bodies that were observed growing only in lit condition as shown in Fig. 6. Nevertheless, these results indicate that X. papulis Lloyd can be incubated in lit, alternating light and dark (every 12 h) and dark conditions. However, in the mass production of mycelia of X. papulis Lloyd the lit condition was considered. Figure 6 shows the mycelial biomass of X. papulis Lloyd produced after 10 d of incubation in different illumination conditions such as lit, alternating light and dark as well as totally dark.
Table 5. Mycelial growth performance of X. papulis Lloyd in different illumination conditions after 10 d of incubation.
Illumination condition Mycelial thickness* (mm) Dry mycelial weight* (mg) Lit 1.99 ± 0.41a 405.20 ± 57.30a Alternate light and dark (every 12 h) 1.25 ± 0.50b 380.80 ± 20.80a Dark 1.17 ± 0.48b 403.70 ± 13.70a *Values are mean ± SD. Means in a column that do not share a letter are significantly different at 5% level of significance using Tukey's Comparison of Means. -
The luxuriant growth of X. papulis on PSB could be attributed to the nutrient composition of the medium. Potato (Solanum tuberosum) tuber is rich in carbohydrates and fibers particularly the mixture of amylose and amylopectin that serve as a source of carbon for fungal growth[15]. They also mentioned that it contains various vitamins and minerals such as vitamin B6, potassium, phosphorus, magnesium, iron and low amount of sodium needed in many metabolic processes essential for growth. Potato tubers are often used to prepare various media for commercial use like potato dextrose agar (PDA) for culturing a wide range of fungi as they also provide significant amounts of protein with high biological values of 90 to 100 (with 100 being the biological value of eggs) as nitrogen sources[16].
The result obtained in the present study is in agreement with the findings of several studies on the optimization of various Xylaria species. Ramesh et al.[7] reported that among the culture media they have evaluated, Xylaria sp. R006 exhibited a maximum biomass production on potato dextrose broth. Rebbapragada & Kalyanaraman[13] also used potato dextrose broth with yeast extract in their optimization studies on Xylaria feejeensis HMJAU22039. Moreover, Chaichanan et al[17] reported that sucrose is one of the critical components of the culture medium in the optimization study of an endophytic fungus, Xylaria sp. Acra L38 for the production of an antifungal compound known as zofimarin. Sucrose is one of the components of the PSB culture medium used in this study.
Previous studies showed that various Xylaria species grew best from slightly acidic conditions to neutral pH which coincides with the results obtained in the present study, i.e., pH 5.0 to 6.5. In the findings of Ma et al.[18], the highest yield in exopolysaccharide (EPS) production from Xylaria nigripes was obtained in pH 7.0 along with other factors. Liu et al.[19] reported that the highest mycelial biomass production for Xylaria striata under liquid culture conditions was also noted at pH 7.0 incubated under shaking conditions for three days. However, Ramesh et al.[7] reported that maximum mycelial biomass production of Xylaria sp. Strain R006 was attained at pH 5.5 using potato dextrose broth. Moreover, Rebbapragada & Kalyanaraman[13] found that pH 6.0 was one of the optimum conditions for high antioxidant activity of Xylaria feejeensis HMJAU22039. Filamentous fungi are generally known to be tolerant to acidic pH and most of them have an optimum pH between 5.0 to 6.0 for cellular growth and several metabolic activities[20].
The results of this study do not conform with the findings of Liu et al.[19], who reported that the highest production of mycelial biomass of Xylaria striata was obtained at shaking condition for 13 d. Kim et al.[10] reported highest mycelial biomass production of Paecilomyces sinclairii in high agitation rate (250 rpm). Moreover, Wei et al.[21] found out that shaking conditions with agitation rate ranging from 80 rpm to 280 rpm, produced the highest biomass of Daldinia eschscholzii at the highest agitation rate. Subsequently, they explained that low biomass observed at the lowest speed could be due to insufficient oxygen concentration that may affect mycelial growth associated with shaking. In this present study, the low amount of mycelial biomass production using 70 rpm might suggest that higher agitation rates should also be studied.
Temperature is an important factor that affects mycelial growth, production of metabolic products and sporulation of fungi. Increasing temperatures can accelerate enzyme activity but high temperatures can inactivate them affecting metabolism and growth[22]. Temperature affects the growth of all organisms because it controls the rate of metabolic reactions[23]. The results of the present study conforms with the findings of Ramesh et al.[7] in which the optimum temperatures for maximal biomass production of Xylaria sp. Strain R006 was 25 to 30 °C. Moreover, Dulay et al.[24] also reported that optimum temperature conditions ranging from 28 to 30 °C for maximum mycelial biomass were achieved in various mushroom species (Ganoderma lucidum, Pleurotus cystidiosus, Volvariella volvacea, and Schizophyllum commune). Similarly, optimal temperature for macrofungal growth as reported by several studies was between 20 to 30 °C[7].
White light produces changes in the concentration of some metabolites and alters enzyme activities in several related metabolic pathways[25]. Wu et al.[26] reported that controlled illumination has positive effects in bioactive compound production in an ascomycete named Cordyceps militaris in its various cultivation stages. Moreover, Sanchez-Murillo et al.[25] found that continuous illumination resulted in the formation of fruiting structures while total darkness resulted only in continued vegetative growth of an ascomycete Paecilomyces fumosoroseus. The results of the present study are in agreement with the findings of the previous study such that presence of fruiting structures of X. papulis was observed only in continued lighting conditions.
Based on the findings of this study, the optimum indigenous liquid medium for mycelial growth of X. papulis Lloyd is potato sucrose broth (PSB) while optimum physical culture conditions are pH of 6.5 incubated at room temperature of 28 oC under continued lighting and static conditions. It is suggested, however, that future studies should elucidate the nutrient composition of potato sucrose broth for a more thorough evaluation of the nutrient composition of the optimum liquid media for the mycelial growth of X. papulis Lloyd. Also, future studies should consider the relationship between the combined factor of pH and temperature since this study only performed sequential evaluation, as shown in Fig. 1. Interestingly, the biological studies on the nutraceutical attributes of mycelial biomass of X. papulis Lloyd grown in indigenous liquid culture media is soon to be published.
This work was financed by the Commission on Higher Education (CHED) and special credits are accorded to the Central Luzon State University (CLSU), particularly to the Center for Tropical Mushroom Research and Development (CTMRD). Thanks also to the Don Mariano Marcos Memorial State University – North La Union Campus (DMMMSU-NLUC).
-
The authors declare that they have no conflict of interest.
- Copyright: © 2023 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Lopez MKS, Kalaw SP, Dulay RMR, De Leon AM, Reyes RG. 2022. Optimization of mycelial growth of Xylaria papulis Lloyd (Xylariaceae) in indigenous liquid culture conditions, science city of Muñoz, Nueva Ecija, Philippines. Studies in Fungi 7:21 doi: 10.48130/SIF-2022-0021
Optimization of mycelial growth of Xylaria papulis Lloyd (Xylariaceae) in indigenous liquid culture conditions, science city of Muñoz, Nueva Ecija, Philippines
- Received: 11 October 2022
- Accepted: 13 December 2022
- Published online: 28 December 2022
Abstract: The search for potential and inexhaustible sources of bioactive compounds with great functional activities is imperative for potential drug leads in treating various human diseases. In this regard, this study investigated the optimum liquid medium and physical culture conditions for the mycelial growth of X. papulis Lloyd for nutraceutical studies. The greatest mycelial biomass was achieved in PSB liquid medium among the indigenous media evaluated, such as potato sucrose broth (PSB), corn meal broth (CMB), rice bran broth (RBB), and coconut water (CW). In terms of pH, X. papulis Lloyd was observed to grow in a wide range of pH (5.0−8.5), but the best mycelial growth was observed at pH 6.5. Room temperature of 28 °C, lighted (137 lux), and static conditions were the other optimum physical culture conditions for mycelial growth of X. papulis Lloyd.
-
Key words:
- Liquid culture /
- Optimization /
- Xylaria papulis Lloyd













