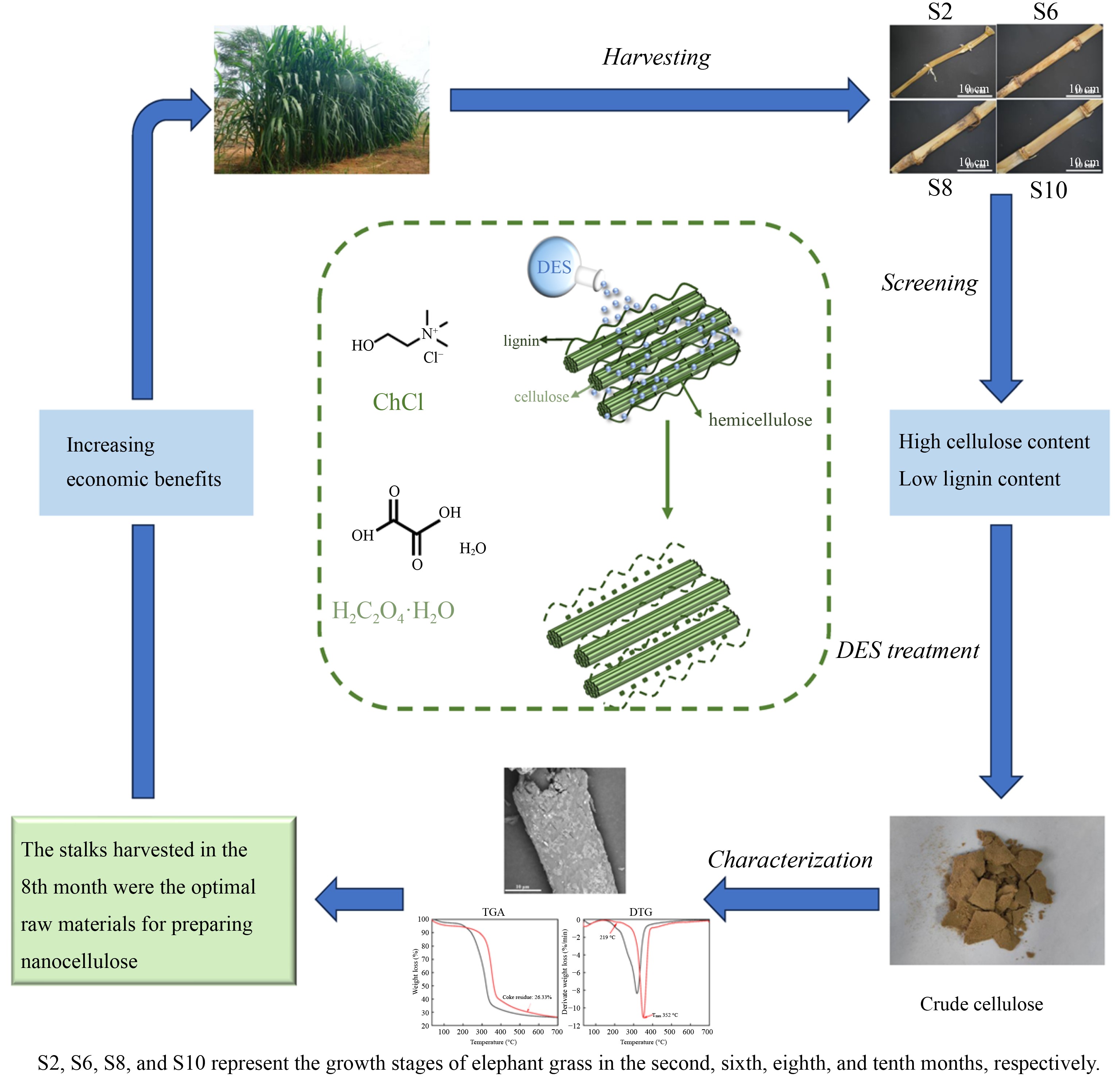
-
Elephant grass (Pennisetum purpureum), also known as Napier grass, belongs to the subfamily Panicoideae of the family Poaceae[1]. It is a large perennial C4 herb native to Africa and is considered the most important forage and energy grass in the tropics and subtropics[2]. It has high resistance and adaptability to various environmental stresses, such as high temperature and drought[1]. Additionally, it demonstrates high photosynthetic efficiency, strong dry matter accumulation capacity, and rich nutritional content[3]. Elephant grass is a lignocellulose-rich plant with a high cellulose and hemicellulose content, surpassing that of most straws, such as corn stover, wheat straw, and reed straw. Therefore, it holds great potential for applications in bioenergy, papermaking, and the preparation of cellulose nanofibers[4−6].
Cellulose nanofibrils are materials derived from lignocellulosic biomass by reducing cellulose to nanoscale dimensions using a variety of physical, chemical, and biological techniques[7]. Due to the abundance of functional groups in cellulose, it can be chemically modified to produce functionalized cellulose nanofibrils. This material finds applications in polymer enhancement, food, medicine, and various other fields[8]. Therefore, it is reasonable to infer that isolating cellulose from elephant grass stalks and preparing cellulose nanofibrils could significantly enhance the economic benefits of elephant grass. Researchers have successfully isolated cellulose from elephant grass stalks and prepared nanocellulose with excellent properties[6, 9]. However, no study has yet reported the effect of different growth stages of elephant grass on the performance of isolated cellulose.
When utilizing elephant grass as a bioenergy source, the objective is to achieve optimal biomass yield and maximum biomass conversion efficiency[2, 10]. For animal forage, a higher leaf rate, higher nitrogen concentration, and lower fiber content are required. However, these factors significantly impact biomass conversion[11, 12]. Plants exhibit different contents of chemical components at various growth stages. Previous studies have shown that harvest management is a crucial factor in determining the chemical composition of perennial grasses[13]. Lignocellulose primarily exists in the plant cell wall, where cellulose serves as the main component, hemicellulose plays a connecting role, and lignin acts as a binder. These components are interconnected by both covalent and non-covalent bonds. Among these, lignin is an aromatic compound that is hydrophobic and difficult to dissolve. Additionally, lignin contains a large number of chromophore groups. Therefore, lignin limits the efficient utilization of biomass to some extent[14, 15]. In conclusion, understanding the changes in the chemical composition of elephant grass at different growth stages can optimize the benefits of biomass conversion and maximize the use of elephant grass resources. This is essential for its sustainable and high-value-added utilization.
Deep eutectic solvent (DES) is a green chemical solvent consisting of hydrogen bond acceptors and hydrogen bond donors[16]. In comparison to traditional organic solvents, deep eutectic solvents offer numerous advantages, including low toxicity, low melting point, and biodegradability. Currently, they are widely utilized in the pretreatment of lignocellulosic biomass to remove a significant amount of hemicellulose and lignin while preventing excessive cellulose degradation[17].
In this study, elephant grass stalks were harvested at different growth stages. Subsequently, crude cellulose was isolated from these stalks using a deep eutectic solvent. Through comparative analysis of the chemical composition differences in elephant grass stalks at various growth stages and the performance of the isolated crude cellulose, the optimal elephant grass feedstock suitable for preparing cellulose nanofibrils were identified.
-
Elephant grass (Germplasm number: CF052696) was provided by the Tropical Crops Genetic Resources Institute, Chinese Academy of Tropical Agricultural Sciences (TCGRI, CATAS), located in Haikou, China. Stalks of elephant grass were collected during the 2nd (S2), 6th (S6), 8th (S8), and 10th (S10) months of growth. Stalk samples were collected from the same part of elephant grass stalks at different growth stages, with three biological replicates collected at each growth stage.
Choline chloride and oxalic acid dihydrate were procured from Shanghai Macklin Biochemical Co., Ltd., (Shanghai, China) and acetone was acquired from Xilong Chemical Co., Ltd. (China).
Methods
Pretreatment of elephant grass stalks
-
The stalks were pulverized using a crusher and screened through a 100-mesh screen to obtain the powder samples. These samples were hermetically sealed and stored in a dry, room-temperature environment.
Isolated of crude cellulose
-
Crude cellulose was isolated from elephant grass stalks using a deep eutectic solvent. The mixture of choline chloride and oxalic acid dihydrate (molar ratio 1:1) was heated and stirred at 60 °C for 1 h until a homogeneous solution was formed to obtain deep eutectic solvent. The elephant grass stalks powder (1.0 g) was added to a deep eutectic solvent (20 g) and heated at 110 °C for 3 h. When the reaction was complete, the flask was placed on ice to cool. The reaction mixture was washed with acetone solution (1:1, v/v), filtered and the solid residue was collected. It was then dried at 60 °C to obtain crude cellulose. The solid recovery of crude cellulose was calculated by the Eqn (1):
$ {\text{Solid recovery}} ({\text{%}})=\left(\dfrac{{Weight \;of\; crude \; cellulose}}{{Weight\; of\; raw\; material}}\right)\times \text{100} $ (1) Analyses and characterizations of elephant grass stalks and crude cellulose
Morphology characterization
-
After gold spraying, the morphology of elephant grass stalks and crude cellulose were characterized using a scanning electron microscope (SEM, Phenom ProX, The Netherlands) at 10 KV.
Determination of water content
-
After weighing the freshly harvested stalks, the samples were dried at 105 °C until reaching a constant weight, and then reweighed to determine their dry weight. The water content of stalks is calculated by the Eqn (2):
$ {\text{Water content}} ({\text{%}})=\left(\dfrac{{Fresh}\;{weight}-{Dry}\;{weight}}{{Fresh}\;{weight}}\right)\times{100} $ (2) Composition analysis of elephant grass stalks
-
The component test methods have been slightly modified according to the references: Cellulose and hemicellulose contents were determined by the phenol-sulfuric acid colorimetric method[18]. Lignin content was determined by the redox method[19]. The crude fiber content was determined by the residual weight method[20]. Acid detergent fiber and neutral detergent fiber were determined using the paradigm washing method[21]. Crude fat extraction was carried out using the soxhlet extraction method[20]. Determination of pectin content was by the carbazole colorimetric method[22]. Nitrogen content was determined by the Kjeldahl method, and crude protein content was calculated using Eqn (3)[20]:
$ {\text{Crude protein}} ({\text{%}})={Nitrogen\; content\; ({\text{%}})}\times {6.25} $ (3) Chemical structure analysis of isolated crude cellulose
-
The fourier transform infrared spectroscopy (FTIR) of crude cellulose was analyzed using a Tensor 27 (Bruker, Germany) spectrometer. The spectra were collected in the range of 4,000–500 cm–1. The spectra were recorded cumulatively in 32 scans with a resolution of 4 cm–1.
The X-ray diffraction (XRD) spectra of crude cellulose were scanned on a D8 Advance XRD diffractometer (40 kV CuKa) with scanning diffraction angles (2θ) from 10° to 45°. The crystallinity index (CrI) of crude cellulose was calculated using the Segal method[23] and Eqn (4):
$ {\text{CrI}} ({\text{%}})=\left(\dfrac{{{I}}_{{002}}-{{I}}_{{am}}}{{{I}}_{{002}}}\right)\times {100} $ (4) where I002 is the maximum diffraction intensity of the 002 crystalline peak at 22°, and Iam is the minimum intensity of the amorphous fraction around 18°.
Thermal gravimetric analysis (TGA)
-
TGA analyses of elephant grass stalks powder and crude cellulose were determined using a TG 209 F3 thermogravimetric analyzer (NETZSCH-Gerätebau GmbH). The temperature was increased from 30 to 700 °C with a heating rate of 10 °C min–1 under a nitrogen atmosphere.
Statistical analysis
-
Analysis of variance (ANOVA) was performed using SPSS 26.0 by Duncan multiple range test (p < 0.05).
-
As shown in Fig. 1a, the color of the leaves changes from light green to dark green as the elephant grass grows, and the lower leaves of the stalks began to dry out by S8. Notably, the stalks of S2 significantly shrink after drying, whereas S6, S8, and S10 hardly show any shrinkage. As elephant grass grows, the aqueous phase of the cell wall is replaced by lignin during stalk lignification[24]. As shown in Fig. 1c, the water content of elephant grass stalks was significantly higher at the S2 stage than at the other three growth stages. Subsequently, the crushed elephant grass stalks were investigated in detail using a SEM. After crushing, the lignocellulosic structure in the S2 stalks was almost destroyed, resulting in a rough surface and a loose structure. However, from the S6 stage onwards, lignocellulose is damaged to a lesser extent, and a distinct tubular structure can be observed. This is attributed to the increased demand for structural support at the later stage of plant growth, the gradual maturation of the cell wall, and the accumulation of lignin, which increases the rigidity and strength of the cell wall[25]. This is consistent with the results of Fig. 1b for fresh weight of elephant grass, which was significantly lower for S2 than the other three growth stages, indicating that S2 was the least lignified. S10 had the highest stalks fresh weight, indicating the highest degree of lignification. Under consistent conditions, the increase of stalk strength mitigates the extent of lignocellulose disruption.
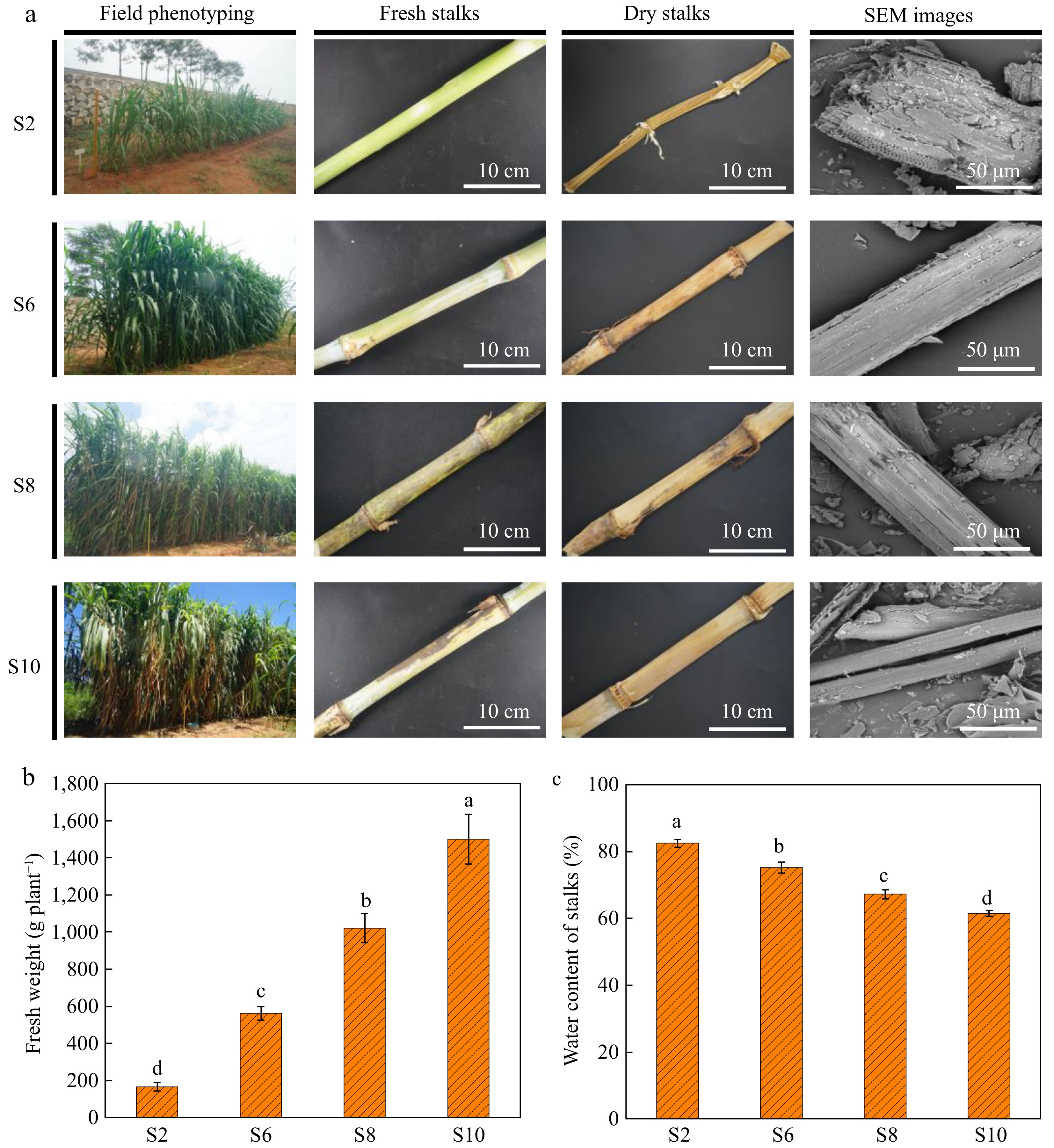
Figure 1.
(a) Field phenotype and stalk morphology of elephant grass. S2, S6, S8, and S10 represent the growth stages of elephant grass in the 2nd, 6th, 8th and 10th months, respectively. SEM (Scanning Electron Microscope) images depict samples of elephant grass stalks at various growth stages. (b) Fresh weight of elephant grass at different growth stages. (c) Water content of elephant grass stalks at different growth stages. Data are means ± SE of three biological replicates. Different letters indicate significant differences (p < 0.05).
Analysis of chemical composition
-
The detergent fiber analyses were initially proposed for forages[11, 26]. Neutral detergent fiber and acid detergent fiber provided estimates of cell wall components, including cellulose, hemicellulose and lignin[27]. They are important indicators to measure the quality cellulosic biomass. As shown in Fig. 2, S8 (69.075%) had the highest neutral detergent fiber content while S2 (44.985%) had the lowest, with no significant difference between S6 (66.926%), S8, and S10 (67.320%). The acid detergent fiber content of S10 (52.330%) was the highest, and likewise there was no significant difference among the three stages of S6 (49.351%), S8 (51.169%), and S10.
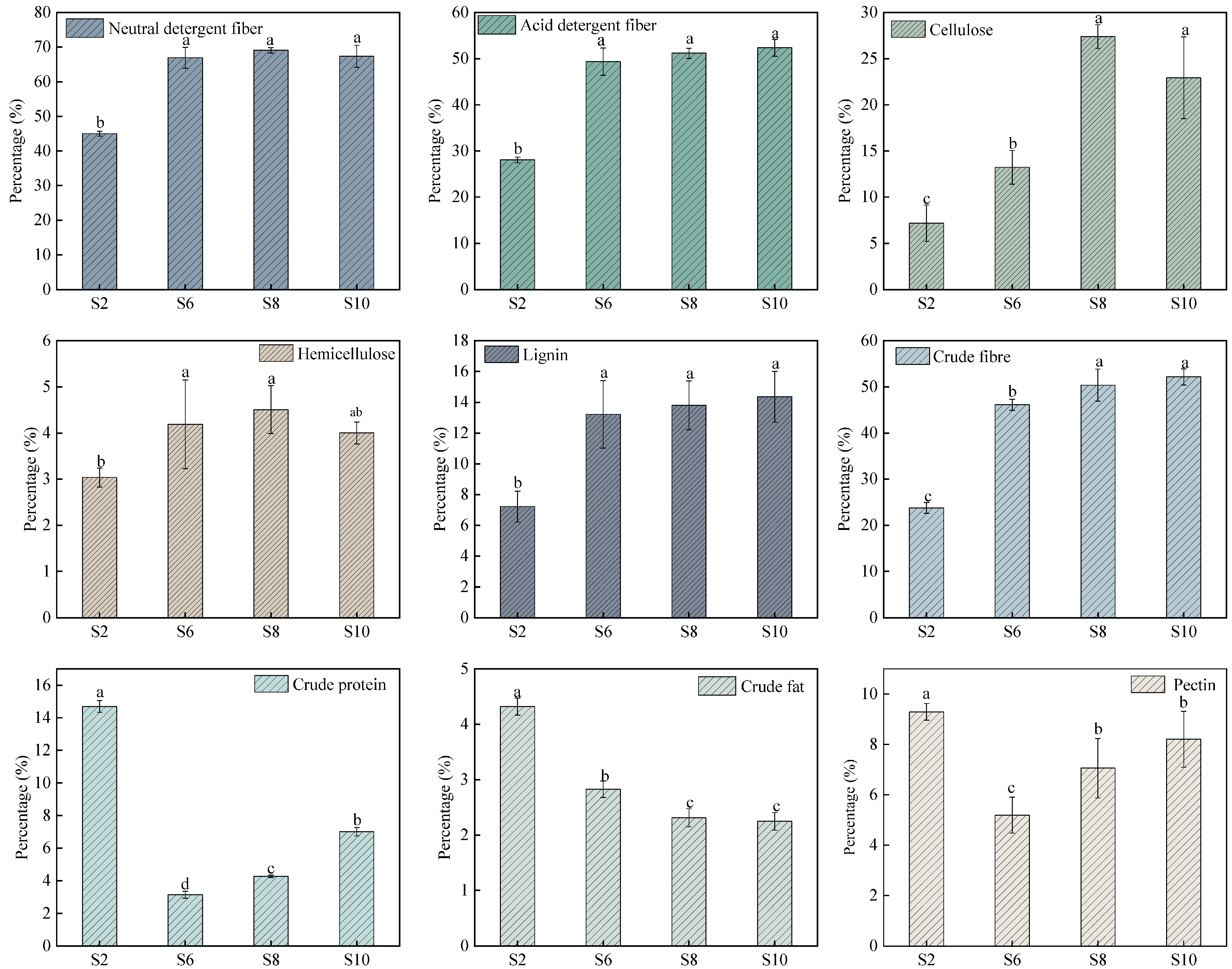
Figure 2.
Changes in the chemical composition content of elephant grass stalks at different growth stages. S2, S6, S8, and S10 represent the growth stages of elephant grass in the 2nd, 6th, 8th and 10th months, respectively. Data are means ± SE of three biological replicates. Different letters indicate significant differences (p < 0.05).
Cellulose and hemicellulose in the cell walls of perennial herbaceous plants act as the major reservoirs of structural carbohydrates, which can be converted into bioenergy. In addition, cellulose and lignin are crucial components affecting biomass conversion, so their content is a significant evaluation index for elephant grass as an energy grass[10]. As shown in Fig. 2, there were significant differences in the cellulose content of elephant grass stalks at different growth stages. The highest cellulose content of 27.386% was found in S8, which was significantly higher than S2 (7.182%) and S6 (13.243%). S10 (14.352%) had the highest lignin content, while S2 (7.224%) had the lowest, with no significant difference between S6 (13.219%), S8 (13.801%), and S10. S8 (4.506%) had the highest hemicellulose content, which was significantly higher than that of S2 (3.037%).
The cell wall is a cellulose-hemicellulose network embedded in the pectin matrix. As a result, pectin may mask cellulose and hemicellulose and prevent their utilization. However, pectin is a minor component of grass cell walls so elephant grass has much less pectin than dicotyledonous plants. In addition, like starch, pectin is mostly water-soluble and relatively easy to degrade compared to other components of the cell wall[28]. There were significant differences in pectin content in elephant grass stalks at different growth stages, with S2 (9.296%) having the highest significantly higher content than S6 (5.191%), S8 (7.059%), and S10 (8.210%), with no significant difference between S8 and S10. Cellulose produced using Acetobacter xylinus had fibers that were more malleable in the presence of pectin. Interestingly, these properties of cellulose remained even when pectin was removed, suggesting that pectin may contribute to the deposition of cellulose within the cell wall, acting as a spacer and preventing the formation of large aggregates in the original fibers[29]. It is suggested that the presence of small amount of pectin helps in the preparation of well dispersed cellulose nanofibrils.
Crude protein content showed an initial decrease followed by an increase, with S2 (14.700%) having the highest content, which was significantly higher than S6 (3.138%), S8 (4.265%), and S10 (7.013%), with S6 having the lowest content. The crude fat content showed a significant decreasing trend, with S2 (4.322%) being the highest, which was significantly higher than S6 (2.830%), S8 (2.313%), and S10 (2.249%), while S10 exhibited the lowest content. The crude fiber content in S10 (52.151%) reached the highest value, significantly higher than S2 (23.774%) and S6 (46.113%).
In conclusion, in the late growth stage of elephant grass, the nutrient content is low and the content of cellulose and other substances is high, which reduces the nutritional value of the forage grass. Therefore, elephant grass can be used as a raw material for isolating cellulose at this stage.
Recovery of crude cellulose
-
Further pretreatment of elephant grass stalks with deep eutectic solvent was used to isolate crude cellulose, and the effect of chemical content in the feedstock on the performance of isolated crude cellulose was investigated. As shown in Fig. 3, the solid recovery of crude cellulose showed an increasing trend. With the growth of elephant grass, the hardness of stalks increases, and the hydrogen bond forces between lignocellulose are enhanced. Therefore, under the same pretreatment conditions, the destructive power of deep eutectic solvent to the hydrogen bond inside lignocellulose is reduced, resulting in a poor lignin removal effect, so the crude cellulose isolated from S10 had the highest solid recovery. In addition, the composition analysis showed that S10 had the highest lignin content (Fig. 2). Therefore, it can be inferred that the lignin content of the crude cellulose isolated from S10 may be more than that of all other crude cellulose samples. High lignin content is not helpful for cellulose nanofibrillation. The low solid recovery rate of crude cellulose isolated from S2 indicates that the utilization rate of the elephant grass is low, and the use of stalks at this stage to isolated cellulose is not conducive to improving the economic benefits of elephant grass.
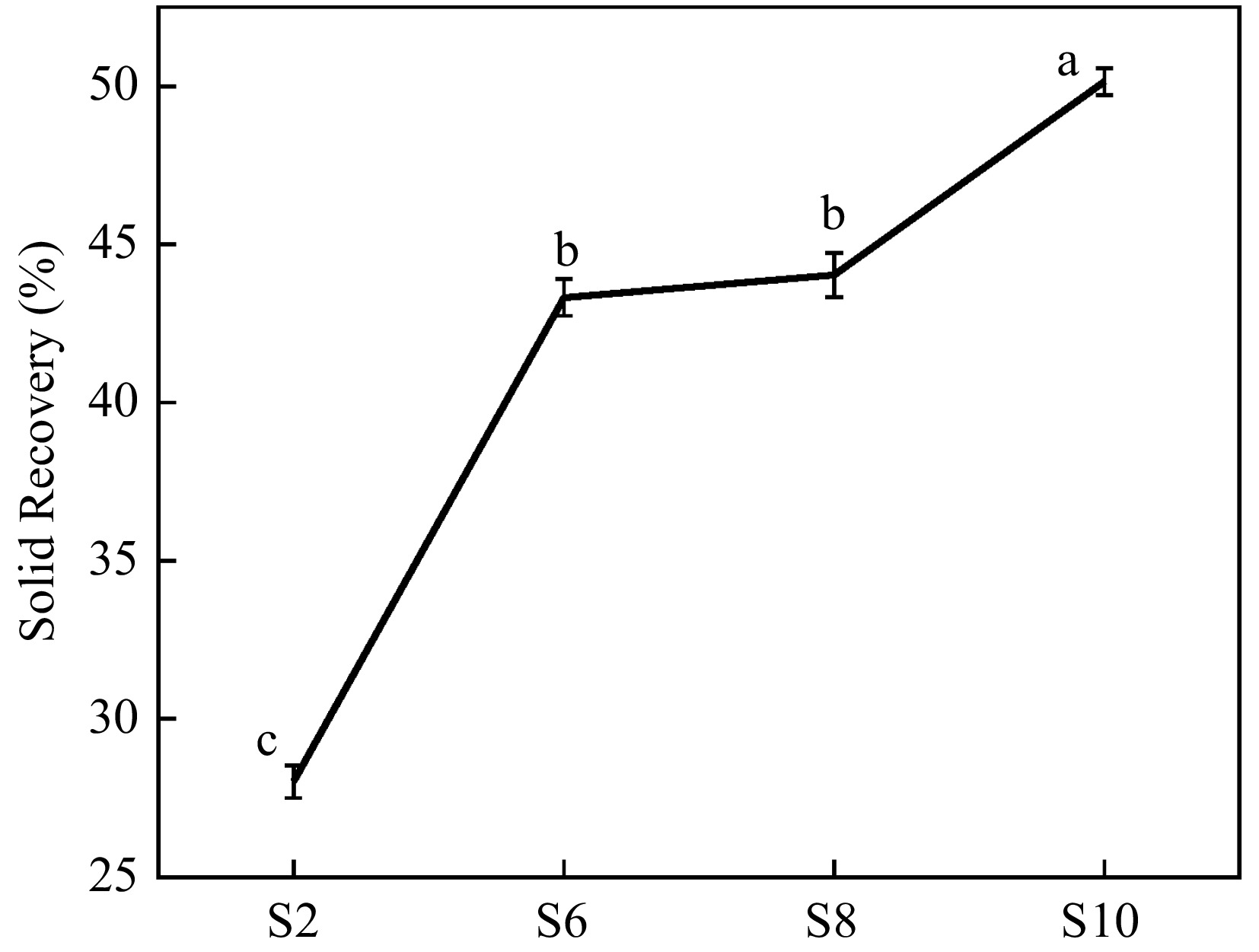
Figure 3.
The solid recovery of crude cellulose. S2, S6, S8, and S10 represent the growth stages of elephant grass in the 2nd, 6th, 8th and 10th months, respectively. Data are means ± SE of three biological replicates. Different letters indicate significant differences (p < 0.05).
SEM characterization of crude cellulose
-
The surface morphology of isolated crude cellulose from different growth stages was analyzed using SEM (Fig. 4). After further deep eutectic solvent pretreatment, the lignocellulose was broken more severely, and the tubular structure of lignocellulose was no longer observed at all in S2, with an increased surface area. In contrast, S6, S8, and S10 show obvious tubular structure, which can also be attributed to higher content of lignin, increasing the rigidity of the cell wall. Compared to S8, S6, and S10 have rougher surfaces. Previous studies have shown that the smooth surface of isolated cellulose after deep eutectic solvent pretreatment arises from the large removal of amorphous cellulose regions such as lignin and hemicellulose[8]. So, it can be inferred that the crude cellulose isolated from S6 and S10 have higher hemicellulose and lignin content.
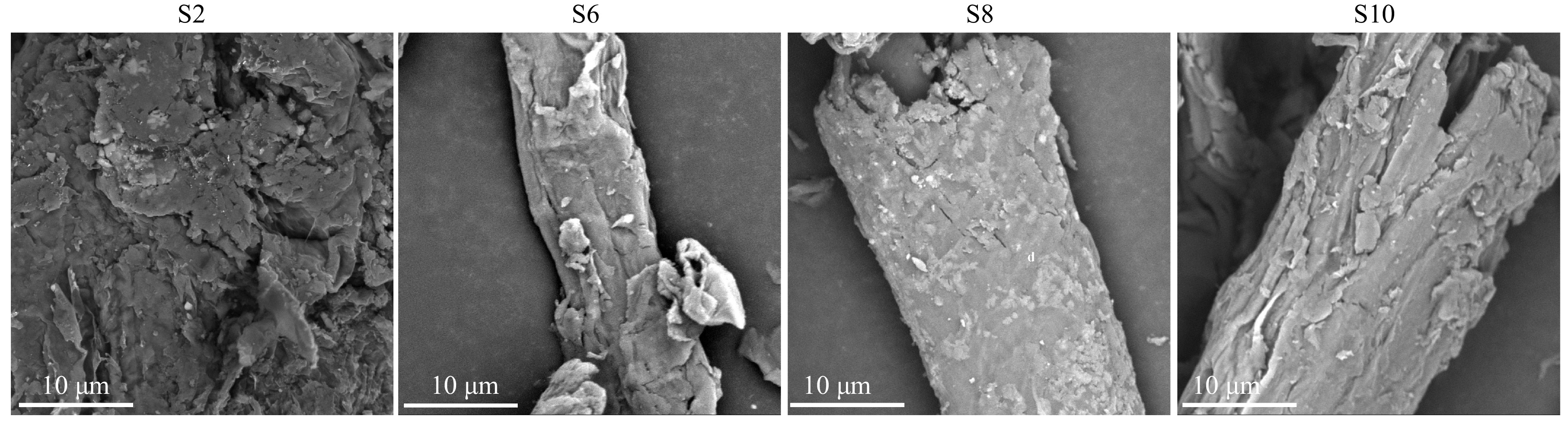

Figure 4.
Scanning Electron Microscope (SEM) image of the isolated crude cellulose from elephant grass stalks. S2, S6, S8, and S10 represent the growth stages of elephant grass in the 2nd, 6th, 8th and 10th months, respectively.
Chemical structure analysis of crude cellulose
-
FTIR can be used to determine the chemical composition in a sample. As shown in Fig. 5, the chemical structures of the isolated crude cellulose are essentially the same. The absorption peaks at 1,517 cm–1 and 1,264 cm–1 were attributed to the C=C (aromatic ring) stretching and C-O stretching of lignin, respectively[8]. The peak (1,517 cm−1) intensity of S2 was the weakest, with slight increases in S6, S8, and S10. The change of peak intensity may be caused by the difference of lignin content in raw materials. It is worth noting that S8 has slightly higher peak strength at 1,427 cm−1 (CH2 bending of pyranose ring), which represents the crystalline region of cellulose[30], and therefore it can be inferred that the cellulose content of S8 is slightly higher than that of the other samples. Interestingly, S2 has the highest peak intensity near 899 cm–1 (β-glycosidic linkage between glucose units). This can be attributed to the higher content of amorphous regions of cellulose in the crude cellulose from S2, the distribution of cellulose β-linkages provides information on the location of amorphous cellulose[30]. This is consistent with the results of the crystallinity analysis below, where S2 has the lowest crystallinity (Fig. 5).
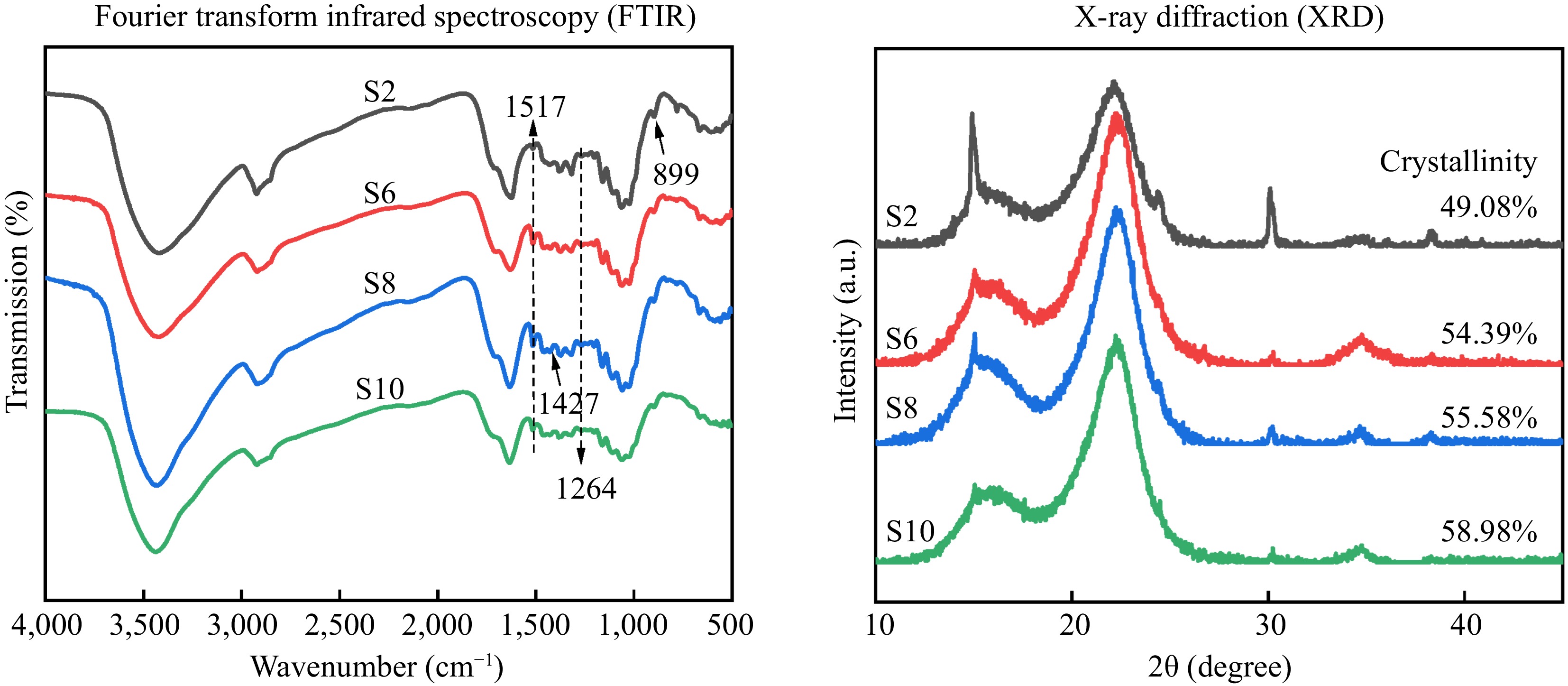
Figure 5.
Chemical structure of crude cellulose. S2, S6, S8, and S10 represent the growth stages of elephant grass in the 2nd, 6th, 8th and 10th months, respectively.
The crystal structure and crystallinity of the isolated crude cellulose were analyzed by XRD, and the ratio of amorphous and crystalline regions in the sample could be estimated from the crystallinity (Fig. 5). The XRD pattern of crude cellulose shows two distinct diffraction peaks around 18° and 22°. The crystallinity of the isolated crude cellulose increased with the growth of elephant grass. Previous studies on bamboo and cotton cellulose have shown that the crystallinity of cellulose increases with plant maturity[31, 32], which was consistent with this study. The development of intermolecular hydrogen bonds contributes strongly to the crystallization of cellulose[32]. S2 has the lowest crystallinity (49.08%). On the one hand, this may be due to the fact that S2 stalks have the lowest cellulose content and a high percentage of amorphous regions of cellulose; on the other hand, S2 stalks have a low strength and weak intermolecular hydrogen bonding forces, which make cellulose more easily degraded during the pretreatment process. The crystallinity of S6, S8, and S10 is more than 50%. This may be due to the removal of some lignin and hemicellulose after deep eutectic solvent pretreatment. Previous studies show that the increase in crystallinity was due to the deep eutectic solvent pretreatment removing some amorphous regions of cellulose, such as lignin and hemicellulose[8].
Thermal stability analysis
-
As shown in Fig. 6, the thermal stability of the isolated crude cellulose was improved to varying degrees due to the removal of some hemicellulose and lignin after deep eutectic solvent pretreatment. The mass loss between 200 °C and the maximum decomposition temperature (Tmax) is caused by the cleavage of glycosidic bonds, and hemicellulose is less thermally stable than lignin and cellulose, so hemicellulose is decomposed first[33]. It can be seen from the derivate weight loss curves of crude cellulose that all samples have a slight weight loss peak near 219 °C, which is caused by the degradation of hemicellulose. The intensity of this peak gradually decreased with the growth of elephant grass and almost disappeared in S8 and S10, showing a more homogeneous thermal degradation behavior, suggesting that the content of hemicellulose was decreased after pretreatment. From the derivate weight loss curve of elephant grass stalks (black line), it can be seen that S8 has the highest Tmax and therefore the highest thermal stability, which was attributed to the higher cellulose content of S8 stalks (Fig. 2). Similarly, among the crude cellulose samples (red line), S8 had the highest thermal stability, further demonstrating the higher cellulose content of the crude cellulose isolated from S8. Lignin is more thermally stable than carbohydrates. After 400 °C, the lignin begins to decompose and char, eventually converting to char with an amorphous structure[34]. It was found that among all crude cellulose samples, S8 (red line) had the lowest coke residue at 700 °C, indicating that the crude cellulose isolated from S8 had the lowest lignin content.
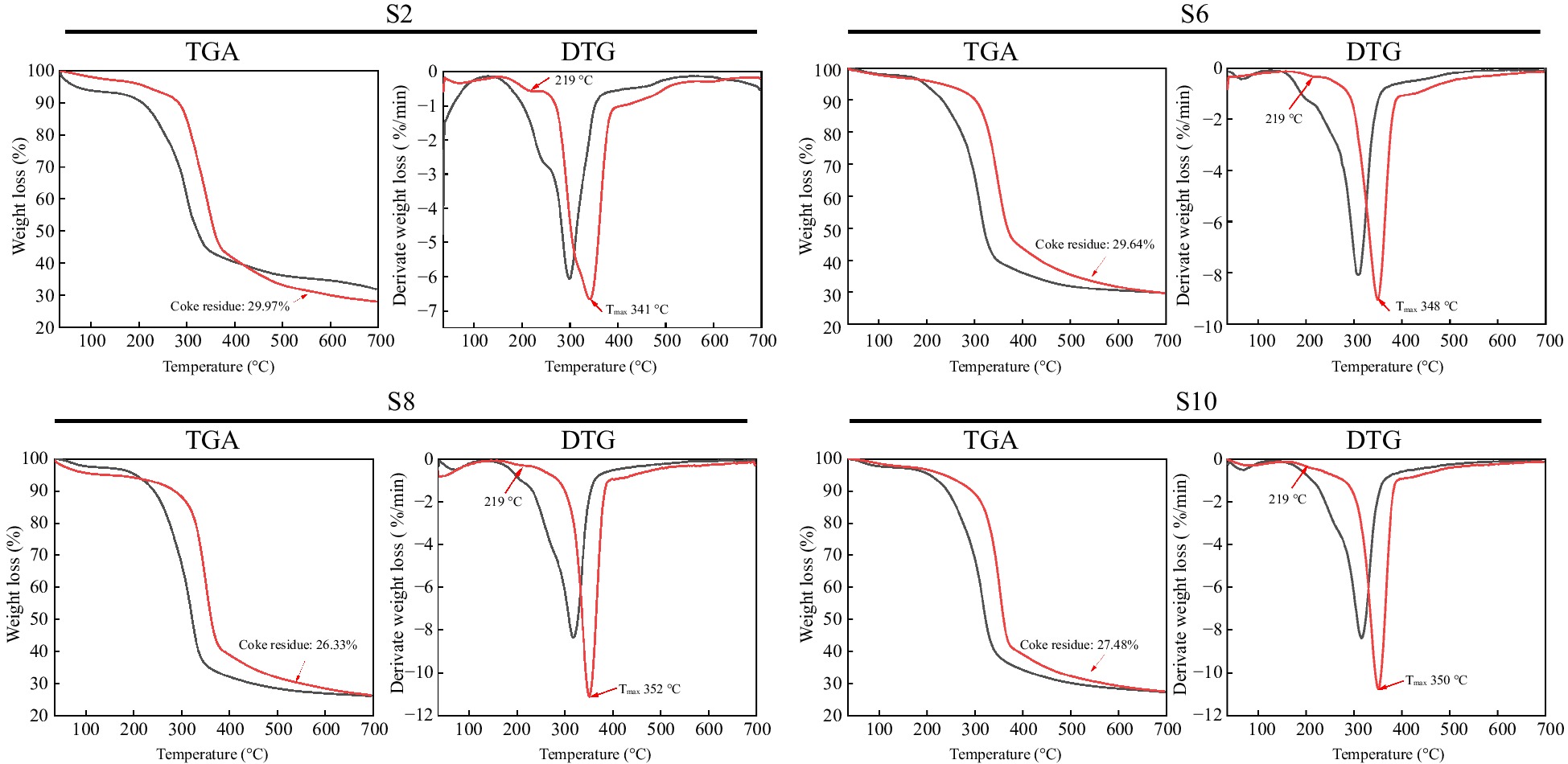
Figure 6.
Thermal gravimetric analysis (TGA) and derivative thermogravimetric analysis (DTG) of elephant grass stalks and crude cellulose isolated from them. The black curve represents elephant grass stalks, while the red curve represents isolated crude cellulose from the stalks. S2, S6, S8, and S10 represent the growth stages of elephant grass in the 2nd, 6th, 8th, and 10th months, respectively.
These results show that the crude cellulose isolated from S8 has the highest thermal stability due to its high cellulose content and low content of amorphous regions such as lignin and hemicellulose. This is consistent with the results of SEM and FTIR analysis. The low lignin and hemicellulose content favors cellulose nanofibrillation, and the higher thermal stability also favors the improved properties of the prepared cellulose nanofibres, which proves that S8 is the optimal raw material for the preparation of cellulose nanofibrils.
-
The effect of elephant grass stalks at different growth stages on the isolated crude cellulose and the possibility of producing cellulose nanofibers was investigated. Raw materials with high cellulose content allowed the isolation of crude cellulose after pretreatment that still maintains a high cellulose content and has excellent thermal stability. In addition, the crystallinity of crude cellulose increased with the maturation of elephant grass stalks. These properties are conducive to improving the properties of the prepared nanocellulose. In this study, the optimal raw material for the preparation of elephant grass cellulose nanofibers was determined, which laid the foundation for the preparation of elephant grass nanocellulose and provided a theoretical basis for the high-value utilization of elephant grass.
The research was financially supported by the Central Public-interest Scientific Institution Basal Research Fund for CATAS (No. 1630032022023) and the China Agriculture Research System of MOF and MARA (CARS-34).
-
The authors confirm contribution to the paper as follows: study conception and design: Huang R, Liu P; data collection: Yuan J, Huang R; analysis and interpretation of results: Yuan J; draft manuscript preparation: Yuan J, Liu P, Liu G. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article.
-
The authors declare that they have no conflict of interest.
-
Received 24 January 2024; Accepted 25 March 2024; Published online 7 May 2024
-
Elephant grass stalks harvested in the 8th month of the growth stage were the optimal raw material for the production of nanocellulose.
The crude cellulose isolated from elephant grass stalks has high thermal stability.
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press on behalf of Hainan University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Yuan J, Liu G, Liu P, Huang R. 2024. Comprehensive assessment of elephant grass (Pennisetum purpureum) stalks at different growth stages as raw materials for nanocellulose production. Tropical Plants 3: e013 doi: 10.48130/tp-0024-0013
Comprehensive assessment of elephant grass (Pennisetum purpureum) stalks at different growth stages as raw materials for nanocellulose production
- Received: 24 January 2024
- Revised: 14 March 2024
- Accepted: 25 March 2024
- Published online: 07 May 2024
Abstract: Elephant grass, a tropical grass plant, serves dual purposes as forage and as a biomass energy source. This study aimed to identify the optimal raw materials for the production of nanocellulose from elephant grass. Elephant grass stalks harvested during the 2nd, 6th, 8th, and 10th months of growth were pretreated with a deep eutectic solvent (containing choline chloride and oxalic acid in a 1:1 molar ratio) for 3 h at 60 °C, to isolate crude cellulose. The results showed that there were significant differences in the content of chemical components in elephant grass stalks at different growth stages. The cellulose content of elephant grass stalks was highest in the 8th month, reaching 27.4%. Meanwhile, there was no significant difference in lignin content among elephant grass stalks harvested in the 6th, 8th, and 10th months. The isolated crude cellulose from elephant grass stalks harvested in the 8th month exhibited excellent thermal stability, high crystallinity, and a high cellulose content. Thus, the stalks harvested in the 8th month were identified as the optimal raw material for preparing nanocellulose. This study lays the foundation for the production of nanocellulose from elephant grass and provides a theoretical basis for its high-value-added utilization.
-
Key words:
- Elephant grass /
- Energy crop /
- Nanocellulose