-
To meet growing energy demands, it is estimated that 22.3 million acres of arable cropland will need to be allocated to biofuel production by the year 2030[1,2]. Perennial forage grasses grown in marginal lands are an attractive source of sustainable energy, and as such, they have been extensively studied as promising second-generation bioenergy crops[1]. These second-generation biofuel feedstocks, such as switchgrass, contain large amounts of lignocellulosic biomass that can provide an inexpensive and abundant source of renewable energy[3].
Lignocellulosic feedstock material is comprised of three major components: lignin, matrix polysaccharides, and cellulose. In conjunction with minor components, such as minerals and proteins, these molecules function together to form the structural base of the plant cell wall[4]. The concentrations of lignin, matrix polysaccharides, and cellulose vary among plant species[4]. Grasses typically contain 25%−40% cellulose, 35%−50% hemicellulose, and 10-30% lignin[5].
Switchgrass is a C4 perennial grass that used to be commonly found growing across the vast prairie region of North America. There are three ecotypes of switchgrass, lowland, upland, and coastal, that differ in their habitat preference[6]. Lowlands and coastals are typically found growing across the warm southern plains of the United States whereas uplands tend to grow across the northern prairies into the southern parts of Canada[7]. Morphologically, lowland ecotypes have thicker stems, wider leaves, and taller tillers than their upland counterparts[7]; whereas coastals have thin, but tall stems. The two better studied ecotypes vary significantly in overall biomass production. Lowland varieties have been shown to produce on average 12.9 Mg·ha−1 of biomass per year, while the upland varieties have been shown to produce on average 8.7 Mg·ha−1 of biomass per year[8]. Currently, several commercial varieties of switchgrass have been released that are suitable for large-scale sustainable biomass production, including lowland varieties 'Alamo' and 'Kanlow', as well as upland cultivars 'Cave-In-Rock' and 'Summer'[9,10].
For switchgrass to be fully utilized as a bioenergy crop, the quality of the lignocellulosic component of the biomass must be improved. Significant effort has been put into identifying elite switchgrass germplasm from already existing cultivars and into developing the best management practices for optimal biomass output[11,12]. In addition, traditional breeding methods have been employed to enhance certain characteristics of switchgrass feedstock, including biomass production and forage digestibility[13,14]. Considering the time constraints of current breeding practices, it takes approximately ten years to develop a new switchgrass cultivar with enhanced characteristics using traditional methods[15].
Recently, genetic engineering practices have been used to create transgenic switchgrass lines with altered cell wall compositions. Since lignin is a limiting factor in the use of lignocellulosic biomass for bioethanol production, several studies in switchgrass have used RNAi technology to knock down genes coding for key enzymes in the lignin biosynthesis pathway, including 4-coumarate:coenzyme A ligase (4CL)[16], cinnamyl alcohol dehydrogenase (CAD)[17,18], and caffeic acid O-methyltransferase (COMT)[19]. Xu et al. found that in comparison to the wild-type plants, transgenic switchgrass lines with reduced 4CL activity had a 22% reduction in overall lignin and released 57.2% more fermentable sugar with dilute acid pretreatment[20]. Alternatively, two independent studies found that down-regulating CAD in switchgrass results in 23% less lignin and cutin[18] or 14%−22% less lignin[17], respectively. Finally, down-regulation of the COMT gene in switchgrass produced up to 38% more ethanol using current biomass fermentation practices[19].
An alternative to directly targeting components of the lignin pathway is to manipulate the master regulator that plays a role in regulation of cell wall composition. Several transcription factors have been identified as key regulators of cell wall biosynthesis[21−25]. The Arabidopsis SHNE family belongs to the APETALA2/ Ethylene Responsive Factor (AP2/ERF) transcription factor family that consists of three members (AtSHN1, AtSHN2, and AtSHN3)[26]. Arabidopsis shn mutants have aberrant deposition of epicuticular wax and altered flower morphology[26,27]. AtSHN1 and its orthologues can regulate wax deposition and drought tolerance in plants[26,28−31]. AtSHN2 and its orthologues function as key regulators of cutin, polysaccharides, and lignin deposition[32−34]. Overexpression of AtSHN2 in rice resulted in transgenic plants with a 34% increase in cellulose content and a 45% decrease in lignin[32]. However, unlike AtSHN1 and AtSHN2, the biological function of AtSHN3 has not been intensively characterized.
Despite its importance as a promising bioenergy crop, only a handful of studies in switchgrass have aimed to identify transcriptional control mechanisms underlying cell wall deposition[35−37]. In this study, we created transgenic switchgrass plants overexpressing the AtSHN3 cDNA sequence from Arabidopsis. The transgenic switchgrass lines consistently displayed stunted growth, but alterations in several matrix polysaccharide monomers varied between AtSHN3 transgenic lines and the wild-type plants. Additionally, we report that overexpressing AtSHN3 in switchgrass compromised rust disease resistance. The results of this study provide insights into the biological functions of AtSHN3 that may negatively regulate the rust disease resistance in switchgrass.
-
Following Agrobacterium transformation of somatic embryogenic switchgrass callus, a total of 49 potential ZmUbi10pro: AtSHN3-overexpressing switchgrass plants were regenerated and transplanted into soil. These 49 plants were derived from seven independent transformation events. Four plants, representing four independent transformation events, were selected for further analysis. DNA samples for all four transgenic lines, as well as the wild-type HR8 control, were analyzed by Southern blot. Southern blot analysis showed that three of the four selected lines contained multiple transgene insertions (Fig. 1). SHN4-1 contained three copies of the transgene, whereas SHN5-2 and SHN7-2 contained two copies of the transgene. SHN6-3 was the only line with a single insertion copy of the transgene.
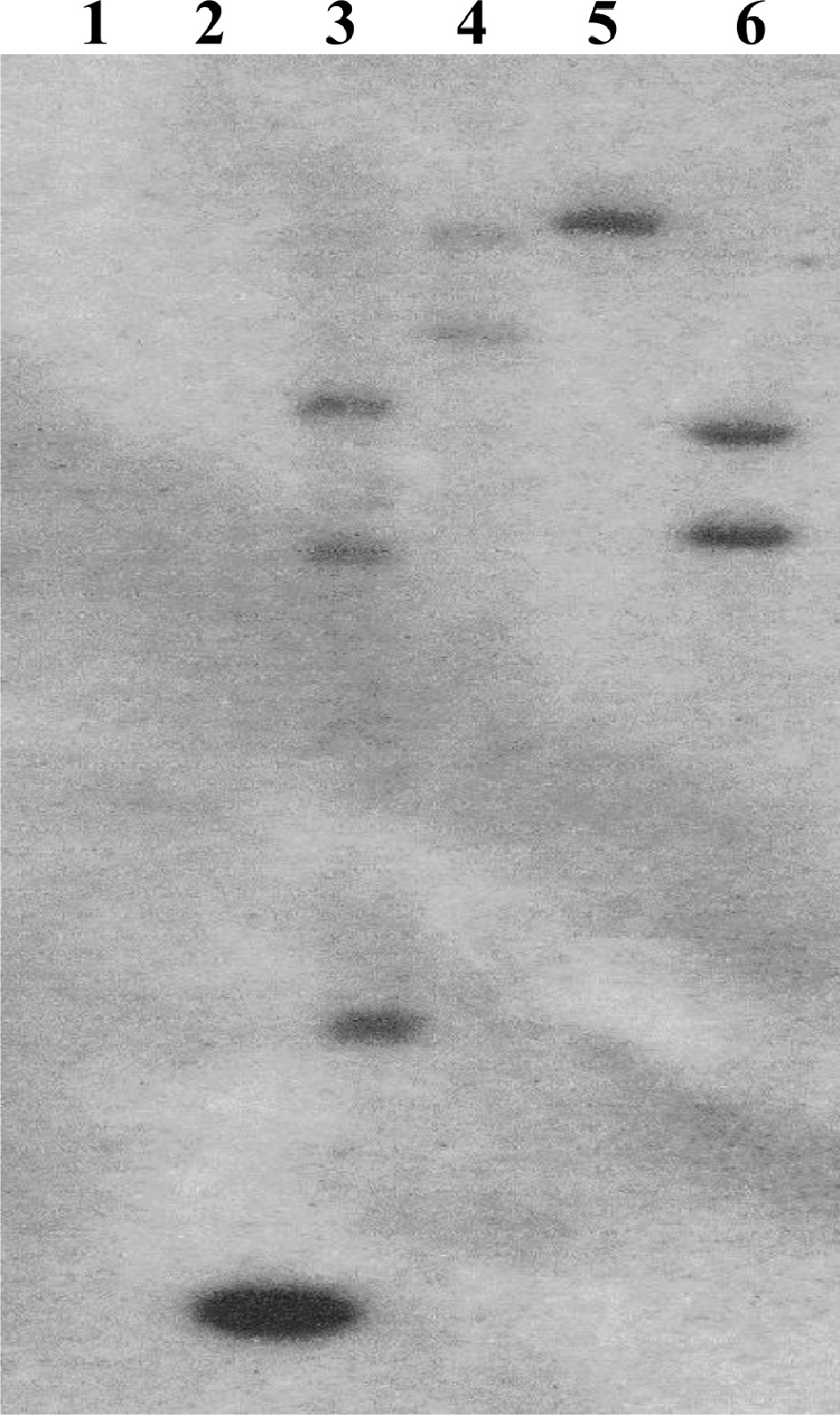
Figure 1.
Southern blot confirmation of transgene insertion. A portion of the hygromycin selection gene was used as a probe. (1) HR8 negative control, (2) 1 kb positive standard, (3) SHN4-1, (4) SHN5-2, (5) SHN6-3, (6) SHN7-2.
AtSHN3-overexpressing transgenic switchgrass are phenotypically different from wild-type plants
-
Growth and development were compared between greenhouse-grown transgenic AtSHN3-overexpressing plants and the wild-type HR8 control plants after three months. Two of the AtSHN3-overexpressing transgenic plants, SHN4-1 and SHN7-2, appeared shorter than the HR8 control (Fig. 2). Several agronomic traits were measured for all plants with three replicates to evaluate the degree of stunting. These included the number of tillers, tiller height, leaf length, leaf width, stem size, and overall biomass. The number of tillers produced was not statistically different (p > 0.01) between the transgenic lines and the wild-type plants (Table 1). All of the plants in this study possessed between 6 and 12 tillers per line. Tiller height measurements revealed that the SHN4-1 and the SHN7-2 plants were significantly shorter (p < 0.01) than the wild-type plants (Table 1).
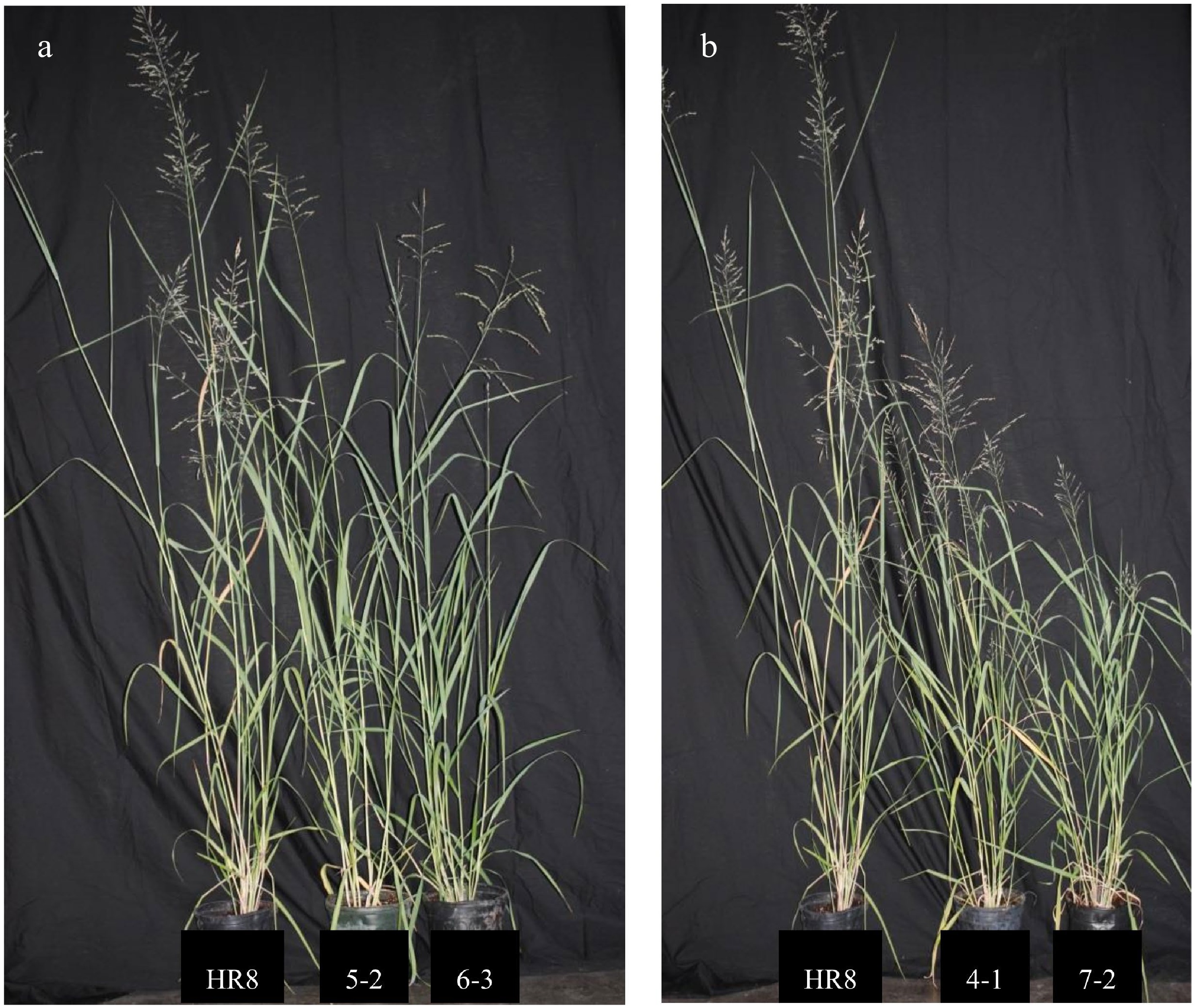

Figure 2.
AtSHN3-overexpressing transgenic switchgrass lines are smaller than wild-type plants. (a) HR8 control plant (left) in comparison to SHN 5-2 (middle) and SHN 6-3 (right). (b) HR8 control plant (left) in comparison to SHN 4-1 (middle) and SHN 7-2 (right).
Table 1. Comparison of agronomic trait measurements for AtSHN3-overexpressing transgenic switchgrass and HR8 control plants. Trait means were not statistically significantly different unless stated, i.e., p > 0.01.
T0 plants Tiller number Tiller height (cm) Flag leaf length (cm) Flag leaf width (mm) Stem width (mm) Biomass (kg)* HR8 12 126.04 33.3 9.33 4.09 0.096 SHN4-1 10.5 85.25** 25.27 7.23** 3.29 0.069 SHN5-2 6.8 126.50 32.17 10.13 4.35 0.087 SHN6-3 9.2 104.59 23.47 8.01 3.93 0.066 SHN7-2 10.6 75.46** 24.77 7.99 3.58 0.073 * = biomass of plant fresh weight; ** = statistically different at p < 0.01. Despite the difference in overall height, the flag leaf lengths of all transgenic lines were not statistically distinguishable from the control plants (p > 0.01, Table 1). SHN4-1 plants had a significantly smaller leaf width (p < 0.01) compared to the HR8 control (Table 1). Both the transgenic AtSHN3-overexpressing lines and the HR8 control plants had similar stem sizes (Table 1). An indicator of change in cell wall composition is the abnormal lengthening of internode stem segments[38]. In this study, we found that the second internode from the base of the plant was shorter for the SHN4-1 and SHN7-2 plants (Fig. 2). Surprisingly, despite their stunted growth, the AtSHN3-overexpressing lines produced comparable biomass to the wild-type plants under greenhouse conditions (Table 1).
Transgene copy insertion number is associated with AtSHN3 gene expression in transgenic switchgrass plants
-
Since the SHN4-1, SHN5-2, and SHN7-2 plants have multiple copies of the transgene (Fig. 1), qPCR was performed to determine if there was a correlation between the transgene copy number and AtSHN3 gene expression. In comparison to SHN6-3, which has a single copy of the transgene, we found that AtSHN3 gene expression increased with increasing transgene copy numbers. SHN4-1 has at least three copies of transgenes (Fig. 1), and it exhibited the highest transgene expression. SHN5-2 and SHN7-2 both have two copies of transgenes. However, their expression was not statistically different from that of SHN6-3 (Fig. 3). SHN4-1 was the shortest among all of the transgenic lines. Thus, differences in the expression levels of AtSHN3 may be contributing to the stunted growth phenotype observed in SHN4-1 switchgrass plants.
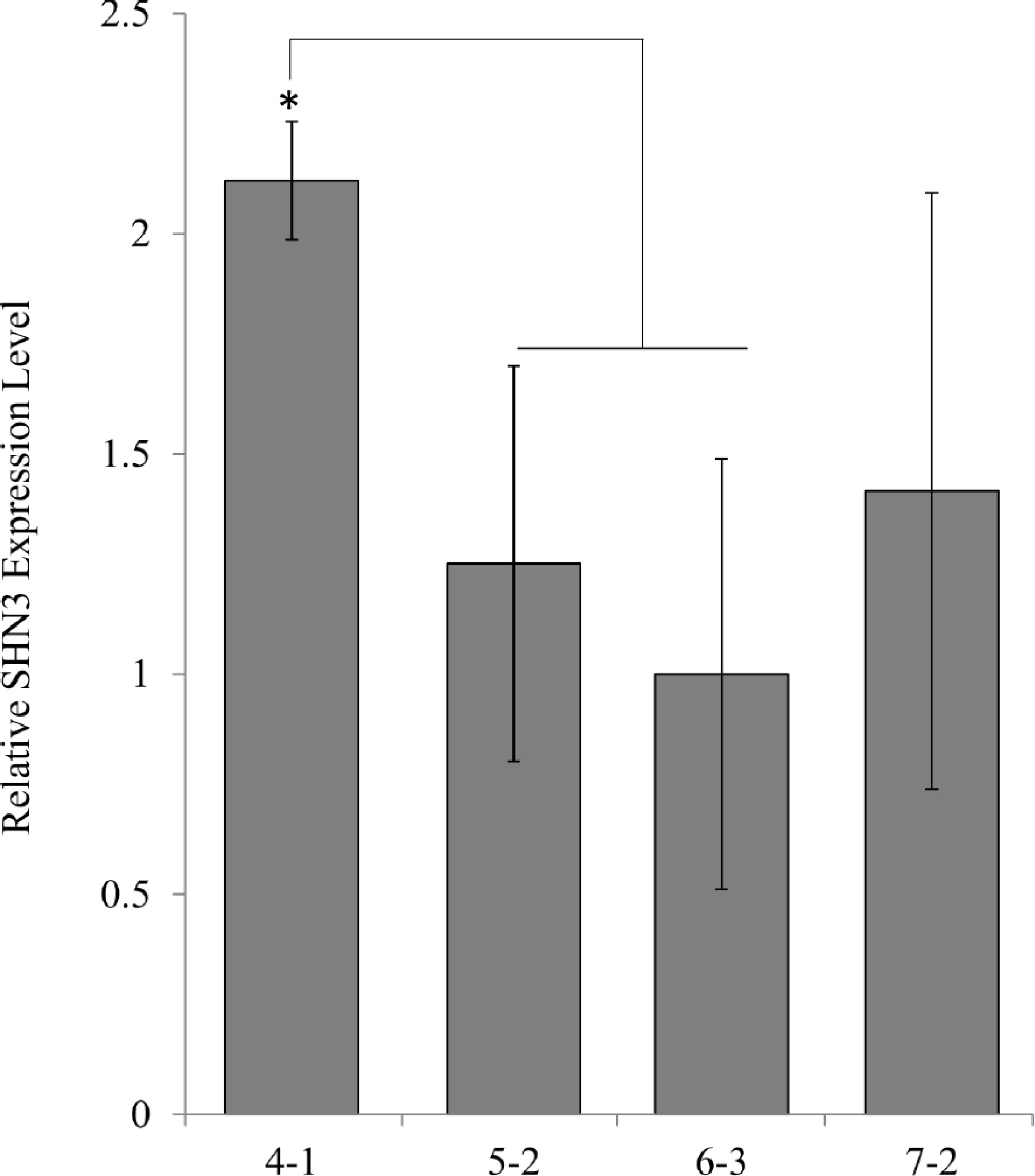
Figure 3.
qPCR analysis of transgene expression levels in AtSHN3 transgenic plants. Expression levels were normalized to the values obtained for SHN3 6-3, which contains one transgene insertion. N = 3, the error bars are standard deviations.
AtSHN3-overexpressing plants have similar lignin and cellulose content to wild-type plants
-
Phloroglucinol staining of I2 sections of transgenic and wild-type switchgrass stems suggested that overexpression of AtSHN3 in switchgrass might alter lignin and cellulose content (Fig. 4). Therefore, I2 stem fragments were subjected to quantify the amount of acid-soluble and acid-insoluble lignin and overall lignin content via sulfuric acid hydrolysis assays. However, the acid-soluble and acid-insoluble lignin contents were not statistically significantly altered between the wild-type and transgenics (Table 2).
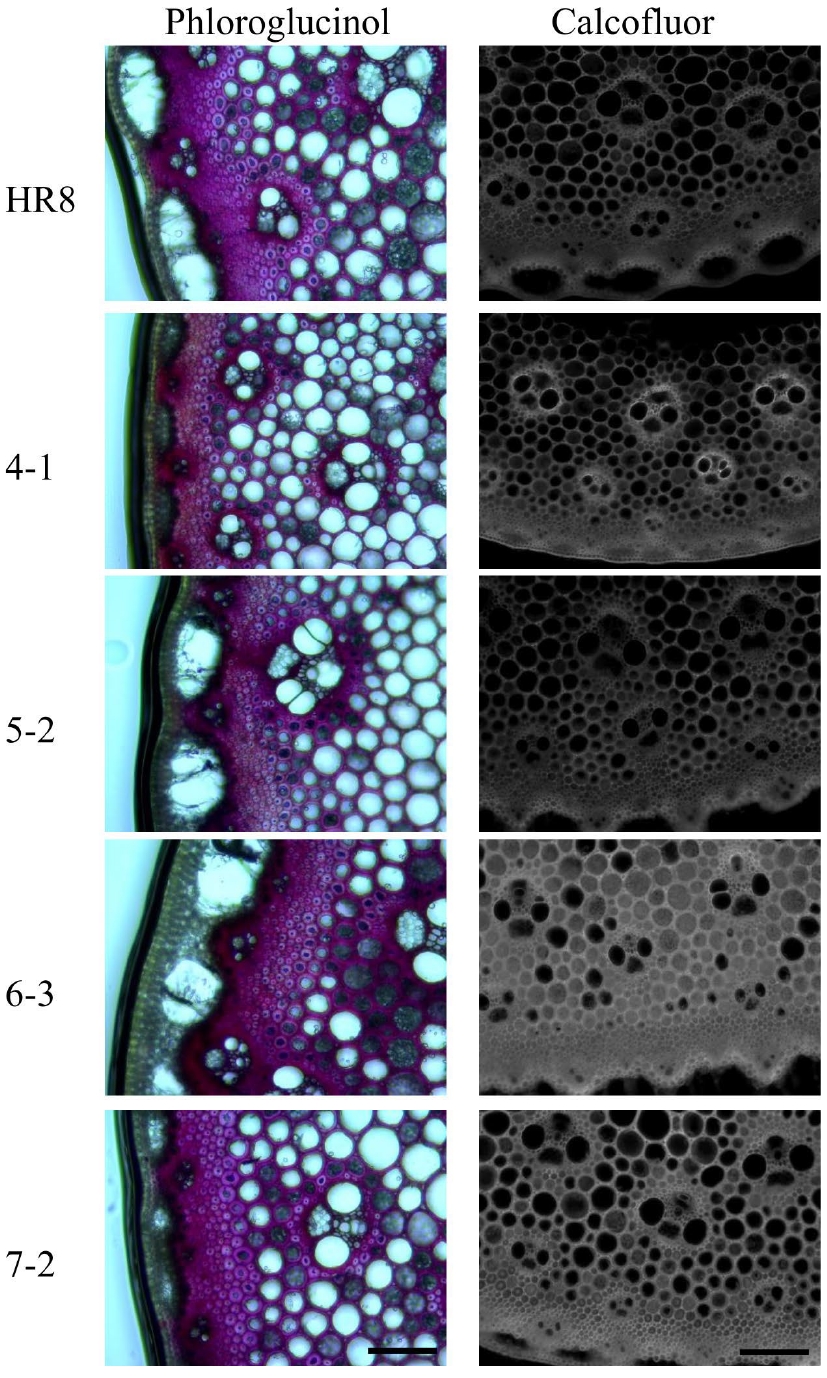
Figure 4.
Phloroglucinol and calcofluor staining of I2 stem sections of wild-type and AtSHN3 transgenics. The tiller segment sections of wild-type and AtSHN3 transgenics were stained with either Phloroglucinol or calcofluor white, and observed under a microscope. Lignin stained with Phloroglucinol is in cherry pink color and the cellulose stained with calcofluor white is showing fluorescence under UV light. All experiments were performed at least twice with similar results.
Table 2. Acid-soluble lignin and acid-insoluble lignin measurement for the AtSHN3-overexpressing transgenic plants and the wild-type control. N = 3, error represents standard deviation.
Switchgrass line % Acid soluble lignin % Acid insoluble lignin % Total lignin HR8 15.5 ± 1.1 2.2 ± 0.2 17.7 ± 1.0 SHN4-1 13.1 ± 0.0 2.1 ± 0.2 15.2 ± 0.4 SHN5-2 15.8 ± 1.1 2.5 ± 0.1 18.3 ± 1.1 SHN6-3 15.1 ± 0.3 2.1 ± 0.4 17.2 ± 0.7 SHN7-2 14.2 ± 0.7 2.1 ± 0.2 16.4 ± 0.9 The cellulosic glucose content of I2R3 stem segments was also measured to determine if the transgenic lines had an increase in cellulose. However, there is no statistically difference between the wild-type and AtSHN3 transgenic plants at p < 0.01 level (Fig. 5). We further analyzed the SHN transgenics of matrix polysaccharide monomers, including arabinose, galactose, glucose, xylose, galacturonic acid, and glucuronic acid. Interestingly, we detected there were significant changes in a few of these hemicellulose sugars between the AtSHN3-overexpressing lines and the HR8 wild-type plants (Fig. 6). For example, SHN7-2 transgenic internodes had 31% more arabinose and 90% more xylose than HR8 control plants (p < 0.01). Also, SHN5-2 transgenic plants had 43% less matrix polysaccharide glucose than the wild-type HR8 plants (p < 0.01).
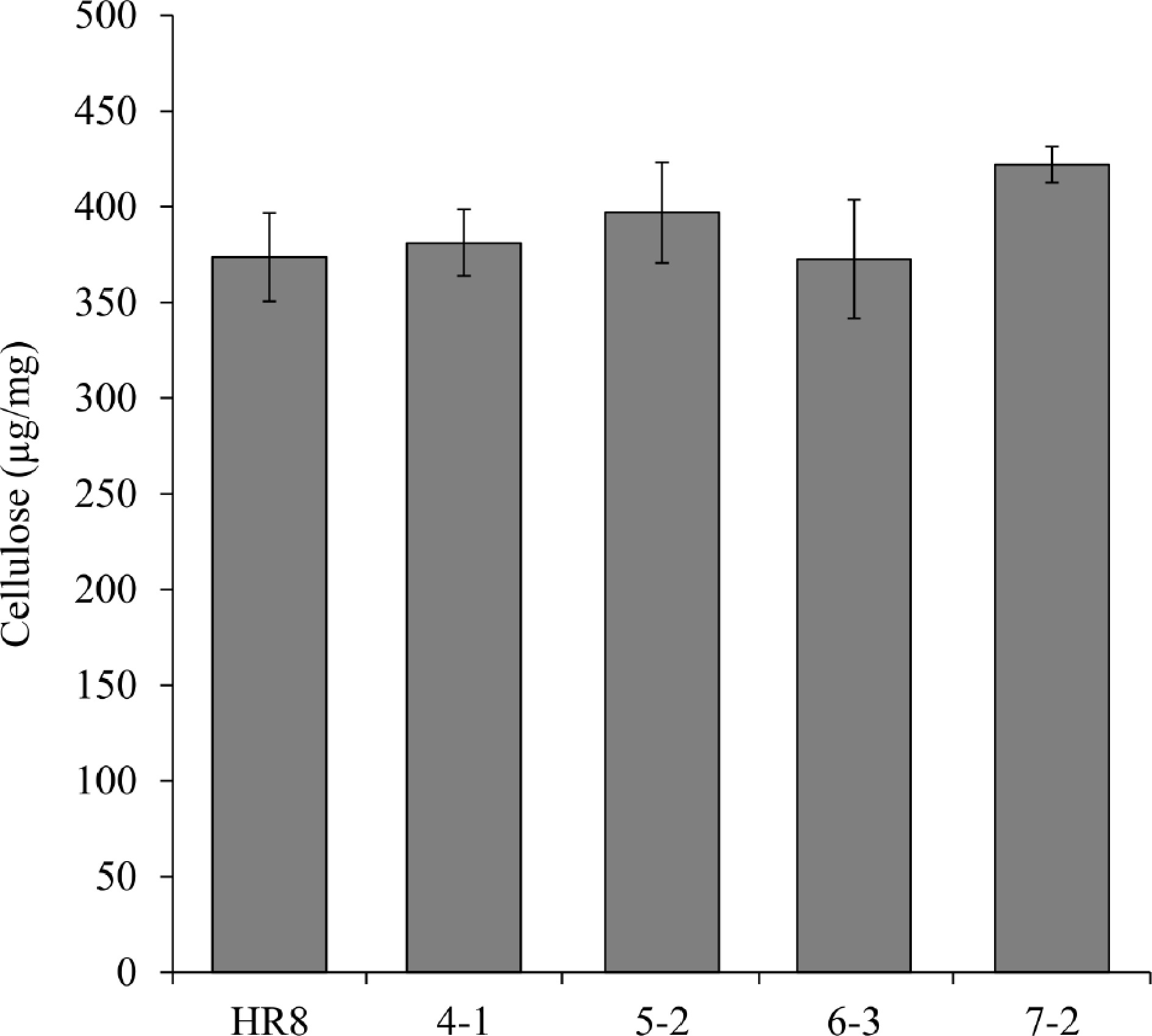
Figure 5.
Measurement of cellulose content between AtSHN3-overexpressing transgenic switchgrass and HR8 wild-type. The cellulose content of the transgenic plants was not statistically different from the wild-type (p < 0.01). N = 3, the error bars are standard deviation.
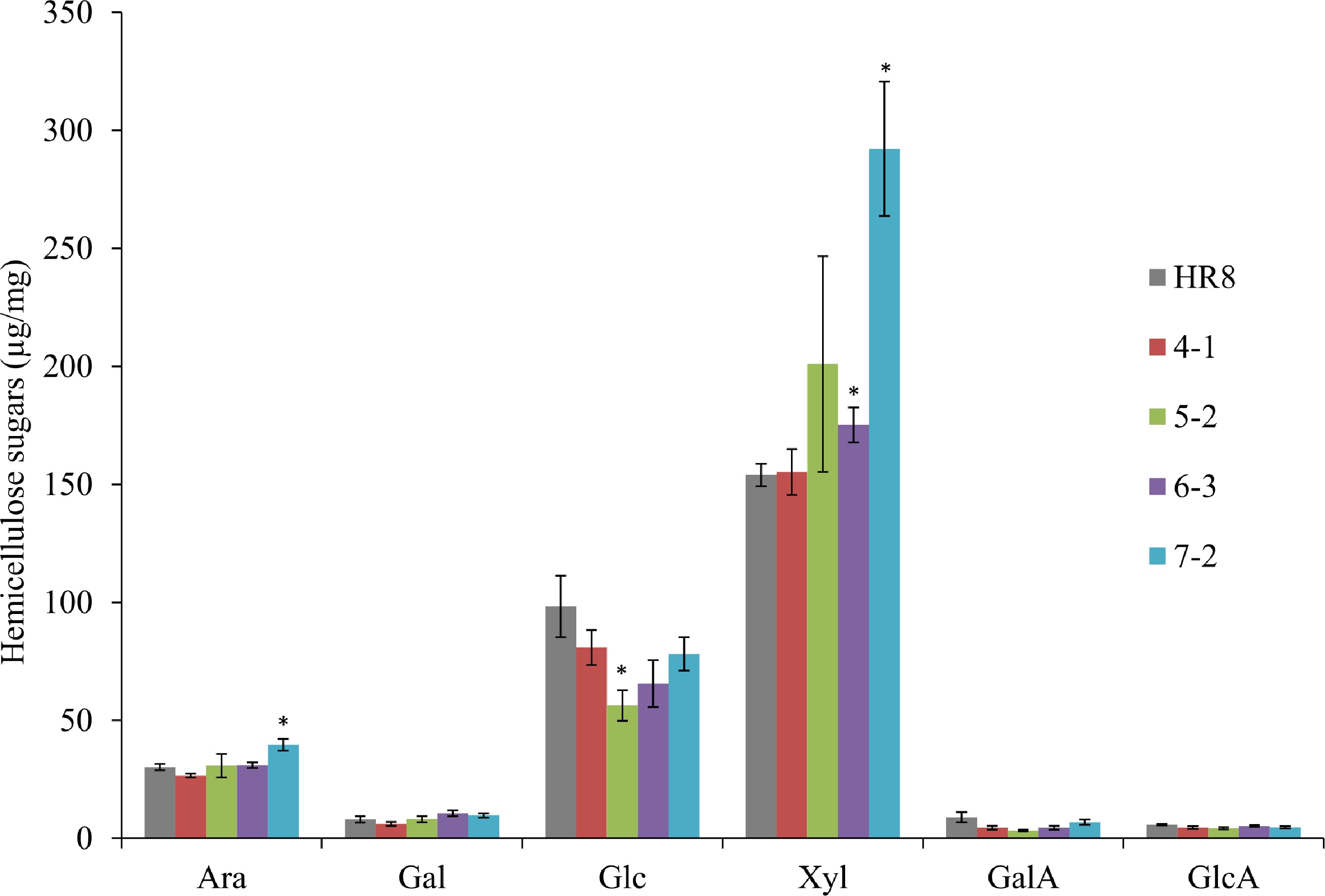
Figure 6.
Matrix polysaccharide sugars in AtSHN3-overexpressing transgenic switchgrass and HR8 wild-type. SHN7-2 transgenic plants were found to have 31% more arabinose and 90% more xylose than HR8 control plants (p < 0.01). SHN5-2 transgenic plants were found to have 43% less glucose than the wild-type HR8 plants (p < 0.01). N = 3, the error bars are standard deviations. The asterisks are the indicators of significant differences between wild-type and transgenics. Arabinose (Ara), Galactose (Gal), Glucose (Glc), Xylose (Xyl), Galacturonic acid (GalA) and Glucuronic acid (GlcA).
Taken together, the overexpression of AtSHN3 does not significantly change the lignin and cellulose contents in switchgrass cell wall biomass; instead, it can alter the deposition of hemicellulose sugars in switchgrass.
Overexpression of AtSHN3 in switchgrass does not alter the stiffness or mechanical strength of switchgrass stems
-
A change in cell wall composition could alter the strength of the stem, which helps the plant maintain an upright growth habit and to withstand abiotic stress such as wind. Storage modulus tests were conducted to measure if the altered hemicellulose contents of the AtSHN3 transgenics could also affect the stiffness of the AtSHN3-overexpressing stems. The test was performed by applying an oscillating stress to the sample and measuring the responding strength. Our results suggest there is no significant difference between the AtSHN3 transgenics lines and the wild-type control plants at the p <0.01 level (Table 3).
Table 3. Average storage modulus derived from stress sweeps at 25ºC for AtSHN3-overexpressing transgenic plants and HR8 wild-type control. The number of repetitions for this experiment is n = 2 for all biological samples.
Storage modulus G’ (Pa) HR8 SHN4-1 SHN5-2 SHN6-3 SHN7-2 Strain 2.1E8 2.2E8 2.2E8 1.5E8 1.8E8 Standard deviation 1.8E7 1.1E8 2.9E7 1.5E7 2.6E7 p-value 0.06 0.89 0.85 0.34 In addition to the stiffness, we also tested if changing the hemicellulose content of cell walls affected the overall mechanical strength of the transgenic switchgrass stems. To accomplish this, fracture tests were performed on switchgrass stem sections by continuously increasing levels of torsion force applied to the stem sections until the stems broke. From the fracture tests, two parameters correlated to the overall mechanical strength of the stem: 1) the slope of the linear region, which reflects the stiffness of the stem, and 2) the breaking point, which correlates to the strength of the stem. The results from both the linear region and breaking point analyses suggest that there was no significant difference between the transgenics and the wild-type control plants (Table 4).
Table 4. Initial linear strength measurement and shear stress at breaking point for the AtSHN3-overexpressing transgenic lines and the HR8 control. The number of repetitions used for this analysis was n = 5.
Measurement HR8 SHN4-1 SHN5-2 SHN6-3 SHN7-2 Initial linear strength (Pa) 2.0E6 1.7E6 1.3E6 1.7E6 1.6E6 Standard deviation 9.3E5 3.1E5 5.6E5 7.7E5 3.9E5 p-value 0.53 0.24 0.65 0.47 Shear stress at breaking
point (Pa)1.9E7 1.6E7 1.7E7 1.4E7 1.8E7 Standard deviation 5.7E6 2.5E6 3.5E6 4.3E6 8.1E5 p-value 0.39 0.57 0.27 0.61 AtSHN3 over-expressing switchgrass plants are more susceptible than wild-type plants to switchgrass rust
-
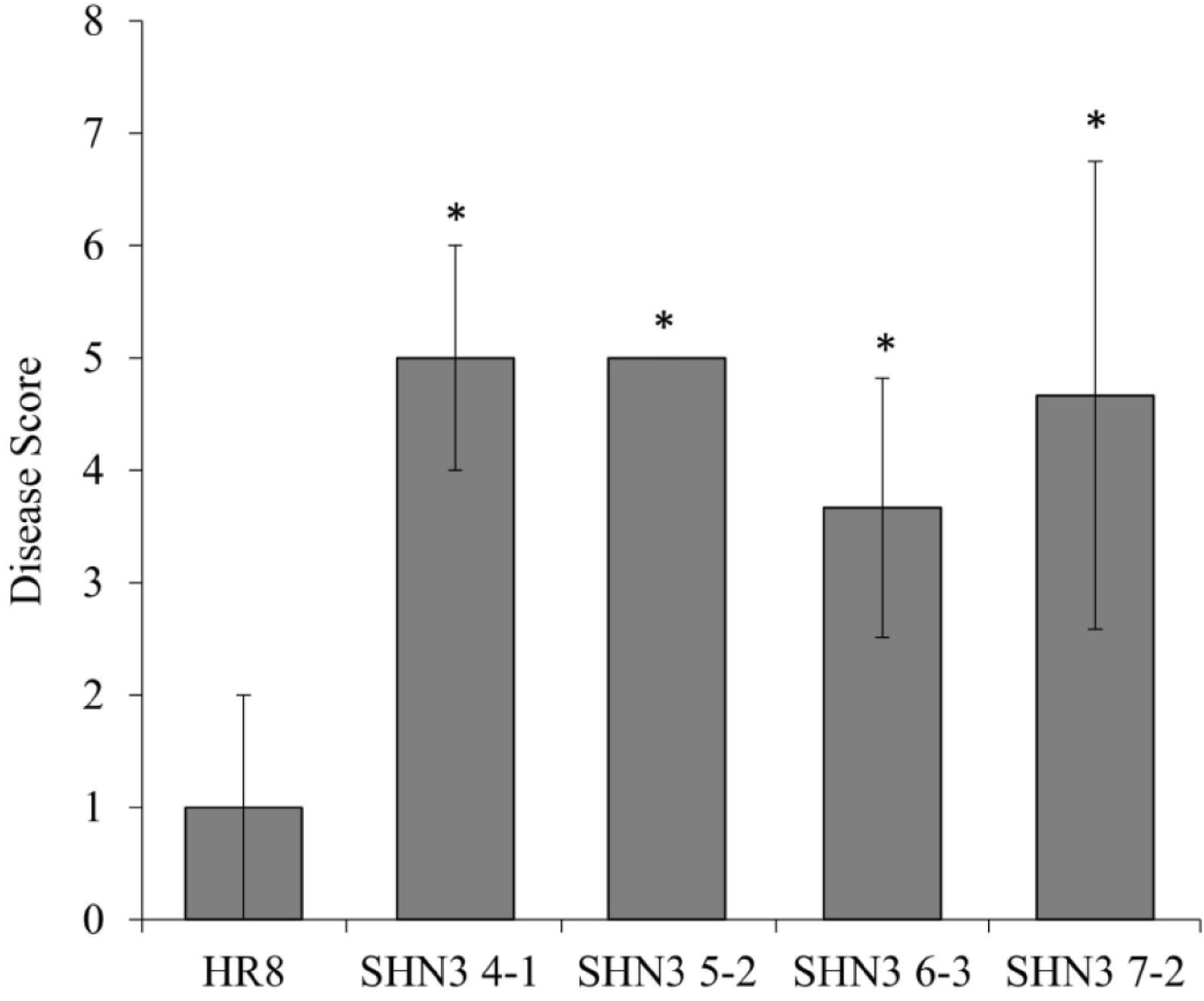
The plant cell wall is the first physical barrier encountered by plant pathogens upon initiation of infection[39]. Since three of these AtSHN3-overexpressing plants (SHN5-2, SHN 6-3, and SHN7-2) have altered hemicellulose content in the cell wall biomass, we further investigated whether or not the SHN3 transgenic plants were more or less susceptible to a rust fungal pathogen. After inoculating both the transgenic lines and the wild-type control with Puccinia emaculata urediniospores, we found that all AtSHN3-overexpressing plants were more susceptible to rust than the HR8 control plants (Fig. 7).
-
Switchgrass is a promising bioenergy crop, and switchgrass cultivars that contain reduced levels of lignin and increased cellulose are desirable for cost-effective and efficient bioethanol production. It is possible to coordinate the activation and repression of these two cell wall components through genetic manipulation of specific master regulators[32]. A previous report suggests that the overexpression of Arabidopsis transcription factor AtSHN2 in rice could increase cellulose and decrease lignin contents of cell wall biomass[32]. AtSHN2 belongs to a small gene family with three members (AtSHN1, 2, and 3) that vary in their developmental and tissue-specific gene expression patterns[26]. AtSHN3 has the broadest expression pattern that is active in almost all plant organs[26]. Despite its proven role in wax accumulation, the other biological functions of AtSHN3 genes have not yet been explored. It is also unclear if AtSHN3, similar to AtSHN2, functions as a master regulator of lignin and cellulose biosynthesis in monocots.
In this study, the Arabidopsis SHN3 cDNA was cloned and transformed into switchgrass. While others have reported a glossy phenotype of the leaf surface upon overexpression of SHN genes[26,40], this phenotype was not observed in any of the transgenic switchgrass plants created in this study. AtSHN3-overexpressing switchgrass plants, however, exhibited stunted growth in comparison to wild-type plants (Fig. 2). Transgenic tomato plants over-expressing SlSHN3, the tomato ortholog of AtSHN3, also displayed stunted growth[40]. Interestingly, the stunted growth phenotype of tomato plants was more severe in SlSHN3-overexpressing plants than in SlSHN1-overexpressing plants[41]. This suggests that while the SHN-family proteins may have similar functions, their tissue-specific expression patterns are essential for proper cell wall development.
The SHN genes regulate wax deposition on both leaf and fruit cuticle surface[26,27]. In addition, members of the SHN family function in cell elongation and secondary cell wall thickening[27,32]. AtSHN2-overexpression can reduce lignin and increase cellulose in transgenic rice plants[32]. The overexpression of AtSHN3 in switchgrass stems might alter the lignin and cellulose contents based on the phloroglucinol and calcofluor staining (Fig. 4). However, the quantitive measurements of cellulose content did not reveal a statistically significant difference between the wild-type and transgenic plants (Fig. 5, Table 2). The phloroglucinol and calcofluor staining methods are relatively easy and cost-effective, but the data interpretation is more subjective, therefore, it is always valuable to obtain quantitative measurement as described in this study. In the future, it will be worthy to re-quantify the cellulose content by using more sample replicates that may reduce the experimental variations. Interestingly, the SHN7-2 plants possessed significantly more arabinose and xylose in their matrix polysaccharide compared to wild-type control plants (HR8). The SHN5-2 line, however, had a significant reduction in the amount of glucose present in its matrix polysaccharide, which is likely due to a reduction in amorphous cellulose or mixed linkage glucan. Further studies are needed to determine if the changes in the hemicellulose content in these lines result in enhanced bioethanol production.
Changes in the cell wall compositions of plant stems could compromise the plant's ability to withstand extracellular forces associated with abiotic forces such as wind and rain that cause lodging. For instance, brittle stalk (bk2) mutants of maize contain less cellulose and more lignin, have compromised the mechanical strength of stems, and are easily broken with minimal applied force[42]. In this study, storage modulus and fracture tests were performed on transgenic and wild-type switchgrass lines to assess the stem stiffness and mechanical strength, respectively. These tests were recently developed and optimized for plant biology research[43]. Storage modulus tests can evaluate the stiffness of the stem by analyzing the % strain output as it correlates to a specific stress. Fracture tests utilize tensile-torsion force to apply stress to a sample, and the % strain is measured during the linear region and at the sample breaking point[43]. Our results suggest that there is no statistical difference in terms of stem stiffness and mechanical strength between the wild-type and transgenic plants (Tables 3 & 4). Therefore, the altered hemicellulose content in the AtSHN3 transgenic plants does not significantly reduce the stem stiffness and mechanical strength.
Overexpression of SHN1 genes in transgenic Arabidopsis, tomato, and rice plants conferred greater tolerance to water restriction compared to wild-type plants[26,32,41]. This could be attributed to the accumulation of excess epicuticular waxes on the leaf surface, which contributes to the glossy leaf phenotype or to the reduced numbers of stomata in transgenic plants[41]. More studies will need to be conducted to determine if AtSHN3 can promote the accumulation of epicuticular waxes that may enhance drought tolerance in switchgrass.
The first physical barrier that foliar pathogens encounter is the plant cell wall. Thus, we investigated if AtSHN3 over-expression affected the disease resistance response of switchgrass rust. All AtSHN3 transgenic switchgrass lines were significantly more susceptible to the rust pathogen than the wild-type controls (Fig. 7). Because there are no consistent patterns of the altered polysaccharide monomers in the AtSHN3 transgenic plants, the variation of polysaccharide monomers cannot explain the AtSHN3-mediated disease susceptibility. A previous report suggests that overexpression of SlSHN3 in tomato leaves allowed the leaves to uptake toluidine blue, suggesting that SlSHN3-overexpressing plants contained a more permeable cuticle than the wild-type[40]. It is possible that overexpression of AtSHN3 in switchgrass could also increase permeable cuticle, which might explain the disease susceptibility phenotype of the AtSHN3 transgenic plants. Thus, it will be worth measuring the permeable cuticle in the AtSHN3 transgenic plants in the future.
-
Although Arabidopsis AtSHN3 shares high homology with AtSHN2 and AtSHN1, overexpression of AtSHN3 in switchgrass does not significantly alter the stem lignin and cellulose contents in transgenic switchgrass. Therefore, AtSHN3 may not have a similar function as AtSHN1 and AtSHN2 when they are overexpressed in a monocot plant species. In the future, a comprehensive analysis of all three switchgrass-specific SHN members will be necessary to understand their biological roles in switchgrass.
-
A plasmid containing the AtSHN3 (TAIR accession: U51209, AT5G25390) cDNA was obtained from TAIR-ABRC. The AtSHN3 open reading frame was amplified using a 50 µl PCR reaction with the following components: 25 µl High-Fidelity iProof master mix (Bio-Rad, Hercules, CA, USA), 10 µl plasmid DNA, 10 µl ddH2O, 2.5 µl 10 µM forward primer (5'-CACCGAATTCATGGTACATTCGAAGAAGTTCC-3'), and 2.5 µL 10 µM reverse primer (5'-CGTCTGCAGGACCTGTGCAATGGATCCAGATC-3'). The PCR reaction was run with an initial denaturation step at 98 °C for 3 min, followed by 30 cycles of denaturation at 98 °C for 30 s, annealing at 57 °C for 45 s, and extension at 72 °C for 1 min, and then completed with a final extension at 72 °C for 7 min. Successful amplification of the PCR product was visualized using a 0.8% agarose gel, and the PCR product was purified using a QIAquick Gel Extraction kit (QIAGEN Sciences Inc, Germantown, MD, USA).
Insertion of AtSHN3 into the pVT1629 Gateway-compatible binary vector
-
The purified AtSHN3 PCR product was cloned into the pENTR/D-TOPO vector (Invitrogen, Waltham, MA, USA). The AtSHN3 gene sequence was confirmed by DNA sequencing at the core facility at Virginia Tech. By using a Gateway LR® cloning kit (Invitrogen Inc), the AtSHN3 DNA fragment was subcloned into the pVT1629 destination vector that carries a maize Ubi10 promoter[16]. The final construct, pVT1629-AtSHN3, was conjugated into Agrobacterium tumefacient strain AGL1.
Switchgrass callus formation and Agrobacterium-mediated transformation
-
The method for Agrobacterium-mediated transformation of switchgrass followed that previously described[20,44]. In brief, mature seeds of the HR8 genotype of the switchgrass cv. Alamo was dehusked with 60% sulfuric acid and sterilized with 50% bleach. The sterilized seeds were transferred to callus induction mediums. After 4−6 weeks, embryogenic calli were subcultured onto the callus induction mediums containing 20 g·l−1 proline. Ten days before transformation, embryogenic calli were subcultured again onto callus induction mediums containing proline and 200 µM acetosyringone. After two rounds of culture on selection mediums, the actively growing calli were subcultured to regeneration mediums. Following regeneration and root formation, regenerated plantlets were transplanted into pots containing MiracleGro Moisture Control soil and maintained in a greenhouse at Virginia Tech.
Phenotypic characterization of SHN3 transgenic switchgrass lines
-
In the middle of July 2015, individual E2 to E3 stage tillers from all transgenic SHN3 switchgrass lines, along with the HR8 control, were clonally propagated by splitting a single tiller and re-planted in gallon-size pots containing Miracle-Gro® Moisture Control potting mix. The plants were maintained in a greenhouse at a 16 h photoperiod with supplemental lighting used as needed. After three months of growth, the overall height, flag leaf length, flag leaf width, and I2 stem width of four R3 stage tillers were measured for three biological replicates of each transgenic line as well as the wild-type control. Finally, all plants were harvested at ground level and weighed to determine fresh biomass yield.
DNA extraction and confirmation of transgenic switchgrass plants
-
Leaves of putative transgenic and wild-type switchgrass plants were collected and immediately frozen in liquid nitrogen. Genomic DNA was extracted using a modified 2× CTAB protocol as previously described[45]. The quality and quantity of the DNA was assessed using agarose gels and a Nanodrop-D1000 (Nanodrop, Wilmington, DE, USA). The switchgrass DNA was then sent to Lofstrand Labs Ltd (Gaithersburg, MD, USA) for Southern blot analysis. Briefly, a total of 10 µg of genomic DNA was restriction enzyme digested with HindIII. DNA fragments were separated using gel electrophoresis and probed with a portion of the hygromycin selection gene to detect transgene insertion[16].
RNA extractions and qPCR analysis
-
Flag leaves of R3 stage switchgrass tillers were collected from greenhouse-grown switchgrass plants and immediately frozen in liquid nitrogen. The tissue was stored at −80 °C until further analysis. Tissue samples were collected for three biological replicates of both the transgenic and wild-type plants. Total RNA was extracted using the TRIzol reagent (Invitrogen, Grand Island, NY, USA) according to the manufacturer's protocol. The quality and quantity of the RNA was assessed using a Nanodrop-D1000 (Nanodrop, Wilmington, DE, USA).
The relative expression AtSHEN3 in transgenic plants was analyzed by qPCR with primers pPCRfor, 5'-TCTCTTGAAGAGAAGAGTGT-3', and qPCRrev, 5'-ACGGTGTCTGGTCTTTACAG-3'. The switchgrass ELF1a gene was used as the reference gene (5'-TCAGGATGTGTACAAGATTGGTG-3' and 5'-GCCTGTCAATCTTGGTAATAAGC-3'). First strand cDNA was synthesized using a DyNAmo cDNA synthesis kit (Thermo Fisher, Waltham, MA, USA). Quantitative Real Time-PCR (qPCR) was performed using an Applied Biosystems Power SYBR Green PCR Master Mix (Grand Island, NY, USA). The PCR reactions were performed on an Applied Biosystems 7300 Real-Time PCR machine with the following conditions: 1) an initial denaturation and enzyme activation step at 95 °C for 10 min and 2) 40 cycles of denaturation (95 °C for 30 s), annealing (60 °C for 30 s), and extension (72 °C for 1 min and 30 s). After the reactions had completed, the threshold was manually set to 3.0, and the data was exported for analysis.
Stem sectioning and histochemical staining for lignin and cellulose
-
The second internode (I2) of R3 stage tillers was selected for histochemical staining. I2 was characterized as the first full-length stem segment, located between the first and second distinguishable nodes, from the base of the plant. The I2 segments of transgenic and wild-type plants were cut into 40 µm sections using a microtome. The lignin and polysaccharide content of the transgenic switchgrass plants was visualized using Weisner (phloroglucinol) reactions and calcofluor staining, respectively. The protocols for these reactions were performed as previously described[46]. The Weisner stained stem sections were visualized using a Zeiss compound light microscope and the calcofluor stained stem sections were visualized using a fluorescence Zeiss AxioImager.M1 microscope mounted with a Zeiss AxioCam MRm (Carl Zeiss Microscopy Inc, Oberkochen, Germany).
Measurement of acid-soluble and acid-insoluble lignin
-
I2 segments of R3 stage tillers for three biological replicates of each transgenic line, as well as the HR8 control, were dried in an oven at 48 °C and then ground into a coarse powder using a coffee grinder. Acid-soluble and insoluble lignin content were determined using the procedure established by the National Renewable Energy Laboratory[47]. In brief, 300 mg of ground switchgrass samples were added to a pressure tube along with 3 mL of 72% sulfuric acid to hydrolyze the tissue. The tubes were incubated at 30 °C for 1 h with manual stirring every 5 min. Following incubation, 84 mL of deionized water was added to each tube to dilute the sulfuric acid to a concentration of 4%. The tubes were then autoclaved at 121 °C for 1 h. Next, the tubes were cooled to room temperature, and the mixture was vacuum-filtered through a porcelain crucible. The filtrate, which contained the acid-soluble lignin, was collected and diluted to a volume sufficient to obtain a UV absorption value of 0.7−1.0 at 205 nm. The acid-insoluble reside, which remained in the porcelain crucible, was dried in an oven at 105 °C overnight and then weighed to determine the acid-insoluble lignin content.
Measurement of cellulose and hemicellulose
-
A second set of I2 samples of R3 stage tillers for three biological replicates of control and transgenic plants were also dried in an oven at 48 °C. The samples were ground with a SPEX 2010 GenoGrinder (SPEX SamplePrep, Metuchen, NJ, USA) at 1,500 rpm. The fine powder was then made into alcohol insoluble residue (AIR) and de-starched as described previously[48]. The de-starched AIR was used for cellulose and hemicellulose assays. Hemicellulose monosaccharides were released by 4M TFA treatment for 2 h and then measured by HPLC. The pellets after TFA treatment were used for an anthrone cellulose assay as described previously[48]. Briefly, pellets were hydrolyzed by 72% sulfuric acid to release cellulosic glucose. The cellulosic glucose was quantified by a colorimetric reaction with an anthrone reagent and read on a plate reader at OD625nm.
Solvent-submersion tensile-torsion measurements to determine stem stiffness and mechanical strength
-
Fresh I2 stem segments of R3 stage switchgrass tillers were subjected to solvent-submersion tensile-torsion analysis using an AR G2 rheometer (TA Instruments, New Castle, DE, USA). The I2 segments were cut into 2 cm long fragments and then split longitudinally into four different sections. The samples were then fully saturated with ethylene glycol and stored at room temperature for future analysis. On the day of analysis, the samples were secured with tension clamps using 15cNxm torque and 1N static tensile force.
All of the testing steps were operated at a frequency of 0.5 Hz and a stress setting of 50,000 Pa. Storage modulus analysis, which is a reflection of stem stiffness, was conducted at room temperature by equilibrating the samples at 25 ºC for 5 min and then running the stress sweep. At least three observations were recorded for each sample type. Ultimate fracture tests of ethylene glycol saturated stem samples were conducted in tensile-torsion mode at room temperature. The specimens were clamped at both ends with slight tensile force (1N) to hold the sample vertically straight. Fracture tests were performed under continuous flow conditions with shear stress increasing from 1E5 Pa to 1E8 Pa. The tests were performed four times per sample. Data acquisition was performed in linear mode with a total collection time of 33 min and a total point set at 300. Tests were concluded once the specimens failed.
Rust disease assays of AtSHN3 transgenic plants
-
The AtSHN3-overexpressing transgenic lines and the HR8 control plant were clonally split into three biological replicates. Each biological replicate was planted in a pot containing MiracleGro Moisture Control soil and grown in the greenhouse under a 16 h photoperiod. Freshly collected Puccinia emaculata urediniospores were mixed 1:10 with talcum powder and hand inoculated on the first fully expanded leaf of E2 stage tillers. The plants were placed in a chamber with a humidifier and kept under 100% humidity for 16 h. Ten days post-inoculation, the severity of rust disease was scored according to the scale established by Gustafson et al.[49]
Statistical analyses
-
All statistical analyses were performed using Student's ANOVA-tests with a significance level of 0.01, chosen to compensate for multiple testing.
The project was supported by USDA-NIFA Grant Number 2011-67009-30133 (B. Zhao). The project was also partially supported by a Virginia Tech CALS integrative grant, a seed grant of the Institute for Critical Technology and Applied Science at Virginia Tech, and Virginia Agricultural Experiment Station (VA135872) to B. Zhao, and by a USDA South Central Sungrant to L. Bartley.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2023 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Frazier TP, Lin F, Wang G, Norris A, Toro C, et al. 2023. Overexpression of the Arabidopsis SHN3 transcription factor compromises the rust disease resistance of transgenic switchgrass plants. Grass Research 3:4 doi: 10.48130/GR-2023-0004
Overexpression of the Arabidopsis SHN3 transcription factor compromises the rust disease resistance of transgenic switchgrass plants
- Received: 28 July 2022
- Accepted: 15 February 2023
- Published online: 30 March 2023
Abstract: Switchgrass can generate large amounts of renewable biomass and hence is one of the most promising bioenergy crops. Improving the quality of switchgrass lignocellulosic biomass will enable its utilization for biofuels. Arabidopsis SHINE family of transcription factor SHN2 was previously identified as a master regulator of cell wall deposition in transgenic rice. However, it is unclear if the Arabidopsis SHN genes also have a similar biological function in switchgrass. Here, we generated transgenic switchgrass overexpressing the Arabidopsis SHN3 transcription factor. Compared with the wild-type, AtSHN3-overexpressing switchgrass plants were stunted in their growth. There were no significant differences in terms of lignin and cellulose content between the SHN transgenics and wild-type switchgrass plants. However, two AtSHN3 transgenic lines SHN7-2 and SHN5-2, displayed significant changes in several matrix polysaccharide monomers. Overexpression of AtSHN3 in switchgrass did not alter the stem mechanical strength when subjected to tensile-torsion analysis. Interestingly, the AtSHN3-overexpressing transgenic lines were more susceptible to switchgrass rust (Puccinia emaculata) than wild-type plants. Therefore, AtSHN3 may have a negative role in regulating disease resistance in switchgrass.
-
Key words:
- Overexpression /
- Arabidopsis /
- SHN3 /
- transcription factor /
- affects /
- cell