-
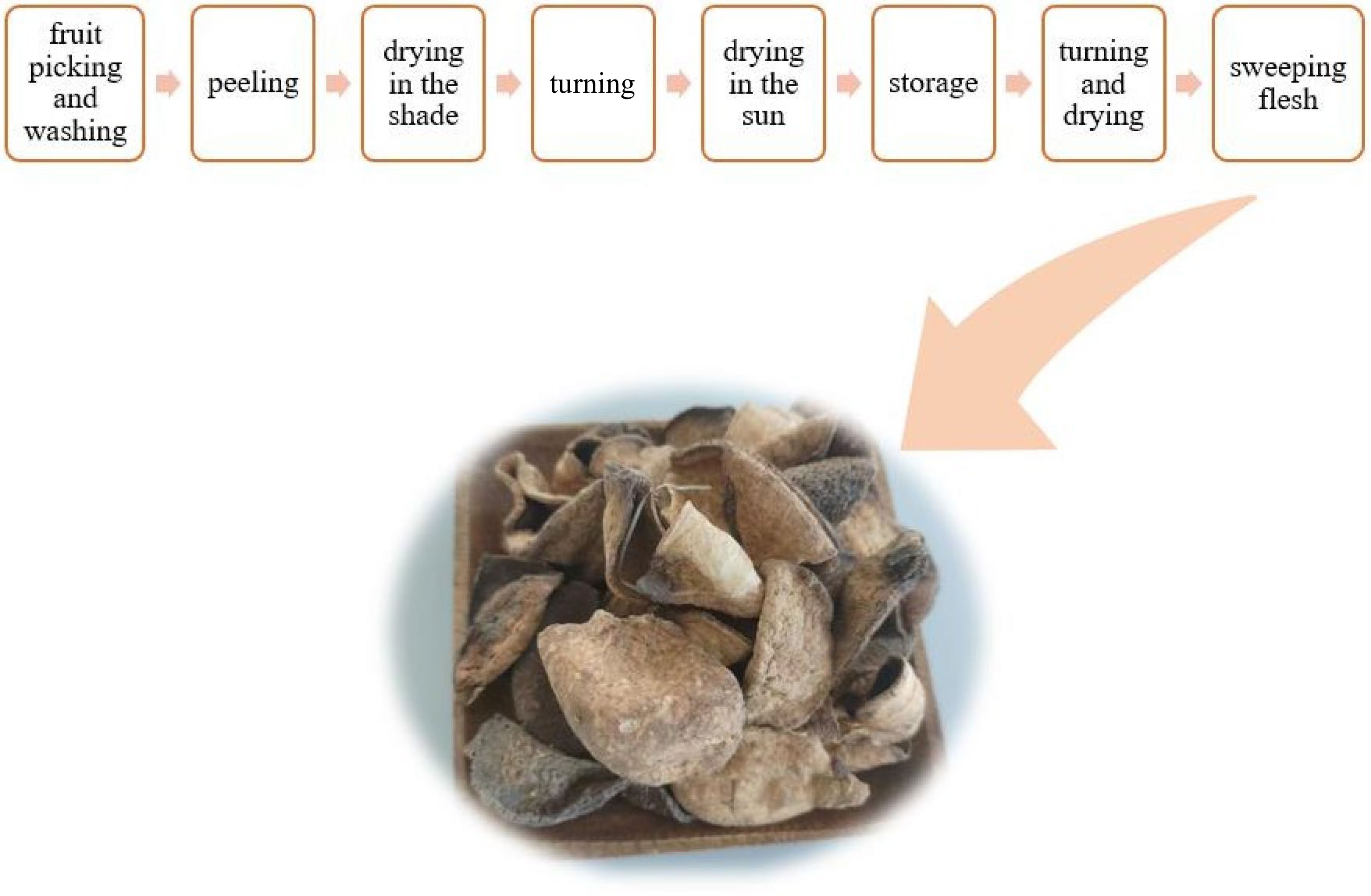
Chenpi (Tangerine peel) is made from ripe peels of citrus (Citrus reticulata Blanco), dried in the sun or low temperatures[1]. The production of dried Chenpi mainly includes fruit picking and washing, peeling, drying in the shade, turning, drying in the sun, storage, turning and drying, and flesh sweeping, as shown in Fig. 1. Chenpi is known for its 'chen' (in Chinese it means a long time), so the best Chenpi comes from being stored for a very long period of time. It should be stored at least for three years. Citrus are usually produced from Guangdong, Fujian, Sichuan, Guangxi and Zhejiang, etc, in China. Multiple kinds of citrus from different places can be made into Chenpi of various qualities. And Xinhui Chenpi is top in its family, Chenpi is usually hard and brittle with a fragrant-pungent, and bitter taste. It plays an important part in promoting digestion and curing-vomiting, coughs, and respiratory diseases[2−8].
The unique aroma of Chenpi is among the many criteria used to evaluate its quality. According to available literature, the aroma of food comes mainly from its volatile compounds, also called aroma-active compounds[9]. Much progress has been made in studying aroma-active compounds of Chenpi with the method of GC-MS. Duan et al.[10] used the GC-MS metabolomics approach combined with principal component analysis and orthogonal partial least squares discriminant analysis to show that the method effectively differentiated samples. Finally, 15 compounds such as methyl methanthranilate, α-sinensal, geranyl acetate, thymol were identified as chemical markers of Chenpi samples. With the method of GC-MS, Luo et al.[2] detected 98 compounds in Chenpi volatile oil and their main components were d-limonene, γ-pinene, α-pinene, linalool and myrcene.
The extraction of volatile compounds of Chenpi is a key technology for its quality analysis. At present, most studies on aroma-active compounds of Chenpi are based on shearing treatment[11] or extraction of volatile compounds with the help of solvent apparatus, and the main extraction methods involved are supercritical fluid extraction (SFE)[12], hydro-distillation (HD)[9], vacuum steam fraction (VSF)[13], solvent extraction (SE)[6,14] and simultaneous distillation extraction (SDE)[15], each of which has its own advantages and disadvantages. The main disadvantages of these methods including the solvent being contaminated and the time consuming nature of the complicated steps, as well as the variety of equipment needed. In this study, water vapor condensation reflux method was used to extract the aroma-active compounds, which is simple and easy to operate. This method resulted in an aqueous solution of aroma-active compounds (hereafter referred to as aroma water), which fully enriched the aroma-active compounds of Chenpi without solvent contamination.
In addition, GC-O was more often used to screen aroma-active compounds. Analyzing Chenpi oil by GC-O and AEDA, Dharmawan et al.[16] concluded that β-pinene, α-pinene, linalool, and 2-methoxy-3-(2-methylpropyl) pyrazine were characteristic aroma substances. Xiao et al.[17] used GC-O to analyze five citruses and concluded that substances such as nonanal, hexanal, linalool and limonene (OAVs ≥ 1) were the characteristic aroma compounds of orange juice samples.
This experiment collects Chenpi aroma water from Chenpi volatiles obtained by heating and hydro distillation. Then the aroma water was analyzed by HS-SPME-GC-MS, and its essential aroma substances were selected using the method of GC-O, AEDA results were expressed as flavor dilution factors (FD), and OAV was used to identify the unique contribution of each compound to the characteristic aroma. In addition, the essential aroma-active compounds of Chenpi varied with heating time during the aroma collecting process. Therefore, the release pattern of essential aroma substances of Chenpi was identified.
-
Xinhui Chenpi sample (WG20210023, WG20210045, and WG20210066) were obtained from Yunnan Tasly Deepure Biological Tea Group Co., Ltd. (Pu’er, Yunnan, China).
Reagents and chemicals
-
Furfural (PubChem CID: 7362; ≥ 99.5%), octanal (PubChem CID: 454; 99%), myrcene (PubChem CID: 31253; ≥ 90%), γ-terpinene (PubChem CID: 7461; 95%), thymol (PubChem CID: 6989; > 99%), (−)-carvone (PubChem CID: 379; ≥ 95%), cymenol (PubChem CID: 10364; 99%), 2-methoxy-4-vinylphenol (PubChem CID: 332; ≥ 98%), citronellyl acetate (PubChem CID: 9017; ≥ 96%), methyl methanthranilate (PubChem CID: 6826; 98%), geranyl acetate (PubChem CID: 7780; 96%), α-terpinene (PubChem CID: 7462; 95%), octanoic acid (PubChem CID: 379; ≥ 99.5%) and n-decanol (PubChem CID: 8174; 98%) were purchased from McLean Biochemical Technology Co., Ltd (Shanghai, China). Geraniol (PubChem CID: 637566; 99%), 1-nonanol (PubChem CID: 8175; 99.5%), 4-terpineol (PubChem CID: 11230; 98%), α-terpineol (PubChem CID: 17100; > 95%), β-ionone (PubChem CID: 638014; 97%), p-cymene (PubChem CID: 7463; ≥ 99.5%), 1-octanol (PubChem CID: 957; > 99.5%), n-decanoic acid (PubChem CID: 2969; > 99%) and β-ionone (PubChem CID: 638014; 97%) were purchased from Aladdin (Shanghai, China). nonanal (PubChem CID: 31289; ≥ 95%), linalool (PubChem CID: 6549; 98%) and d-limonene (PubChem CID: 440917; > 95%) were purchased from TCI (Shanghai, China). NaCl (analytical purity 99.5%) were purchased from Tianjin Zhiyuan Chemical Reagant Co. n-Alkanes solution (C8-C32) was employed to calculate the retention index (RI) of the detected components.
Extraction of the volatile compounds
-
Samples were treated using two different methods. Samples were crushed with a universal crusher (Tianjin Teste Instruments Co., Ltd, Tianjin, China) and passed through 30 mesh sieves before the experiment. The second method is as follows: 100 g of sample was weighed in a 2000-mL round bottom flask, 1,500 mL of hot water was added, soaked for 40 min (in order to shorten the extraction time, and the aroma substances are concentrated), then the samples were distilled by heating. The hydro-distillation method was used to recover condensed water (rich in volatile aroma-active components) and saved for HS-SPME pretreatment. The experiments were repeated in triplicate.
HS-SPME procedure
-
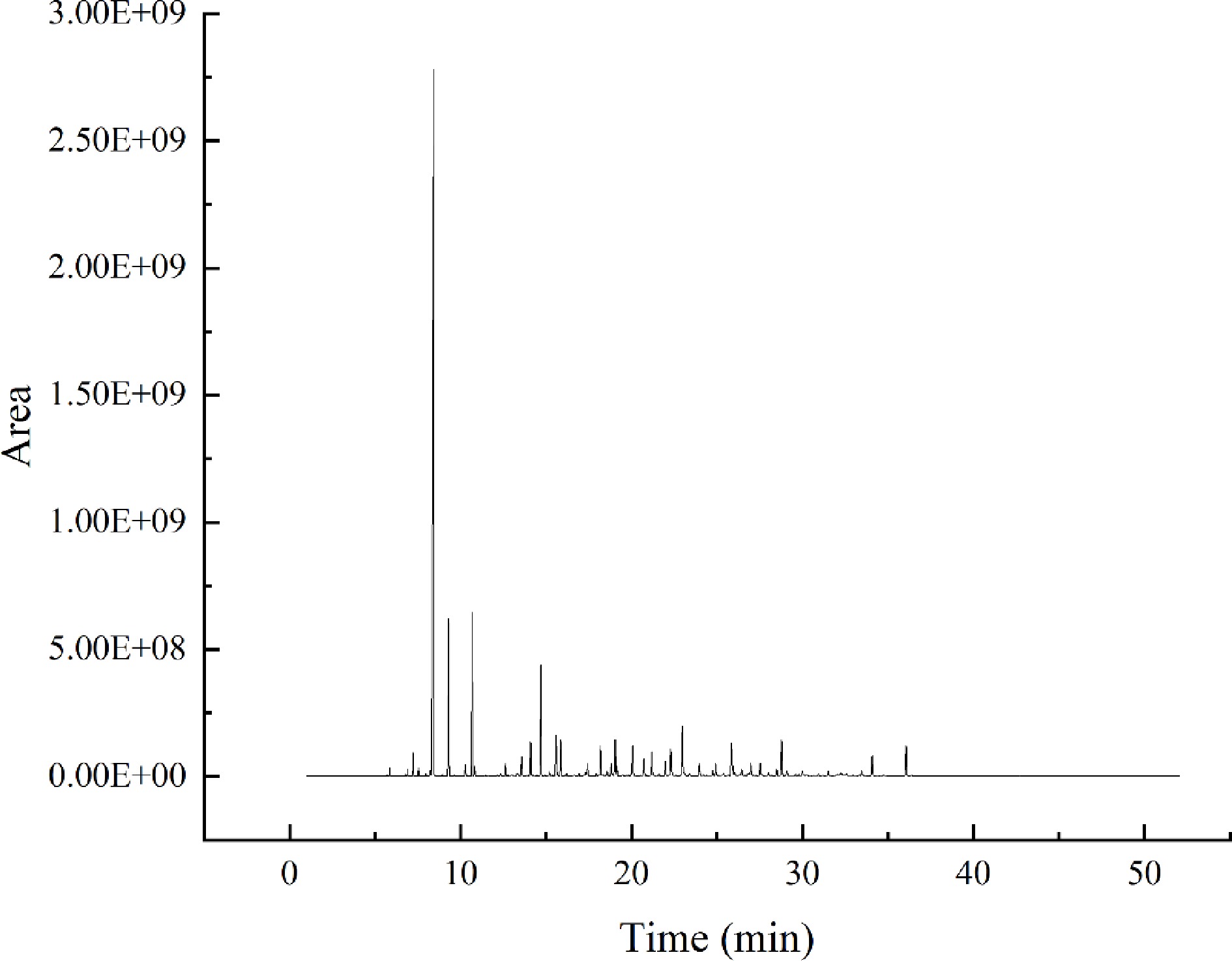
The analytical conditions of HS-SPME, GC-MS, and GC-O were adapted based on the proven method of Liu et al.[11]. The temperature rise procedure, shunt ratio, sample volume, and NaCl addition were optimized to establish the detection method, as shown in Fig. 2. The results showed good peak patterns and were suitable for this study.
New extraction heads were activated, as follows: the 50/30 μm DVB/CAR/PDMS extraction head was placed on the GC-MS inlet for about 30 min under the inlet temperature of 250 °C.
TriPlus RSH autosampler (Thermo Fisher Scientific, USA) coupled with 50/30 μm divinylbenzene/carboxen/Polydimethylsiloxane (DVB/CAR/PDMS) (purchased from Supelco) was used for HS-SPME analysis. Respectively, 1.0 g of sample crushing (add 5 mL of ultra-pure water), 5 mL aroma water sample and 1.5 g of NaCl were placed into a 20-mL headspace bottle. Extraction temperature 65 °C, extraction time 30 min, equilibrium temperature 65 °C, equilibrium time 10 min, desorption time 5 min.
GC-MS analyses and quantitation of aroma-active compounds
-
The equipment used was Rtx-5MS column (30 m × 0.25 mm internal diameter, 0.25 μm film thickness (Restek, Bellefonte, PA, USA), TRACE1300-ISQ gas chromatograph-mass spectrometer (Thermo Fisher Scientific, USA), Thermo Scientific Barnstead water purification system (Thermo Fisher Scientific, USA). The procedures of temperature increase are as follows: initial temperature 40 °C, ramp-up to 70 °C at 10 °C/min, ramp-up to 190 °C at 3 °C/min, ramp-up to 250 °C at 10 °C/min, hold for 3 min; flow rate: 1.0 mL/min; inlet temperature: 250 °C; injection volume: 1.0 μL; split ratio: 70:1; carrier gas: 99.999% high purity helium. EI ion source; ion source temperature: 250 °C; transfer line temperature: 250 °C; ionization energy: 70 eV; scan mode: full scan; mass scan range 40−500 m/z.
Quantitative analysis of volatile compounds was conducted using a standard curve that obtained for each compound. The stock solution composed of 24 aroma-active compounds was configured and diluted with ethanol to six gradients of 2, 2.5, 10/3, 5, 10 and 20 to produce the standard curve. Meanwhile, the retention indices and aroma characteristics of each aroma-active compound were also used to verify the qualitative results.
GC-O analyses
-
GC-O analysis was performed using a Thermo Trace 1300 gas chromatograph (Thermo Fisher Scientific Inc., USA) equipped with a flame ionization detector and a sniffing port (ODP2, Gerstel, Inc., Germany), Rtx-5MS column (30 m × 0.25 mm internal diameter, 0.25 µm film thickness; Restek, Bellefonte, PA, USA) was used for the separation, other equipment used was SGH-300 high-purity hydrogen generator, SGK-2L low-noise air pump (Beijing Zhongke Jirui Technology Co. Ltd., Beijing, China).
Experienced sensory evaluators completed GC-O sniffing experiments, three panelists were selected and trained based on GB/T 16291.1-2012 (Sensory analysis-general guidance for the selection, training, and monitoring of assessors)[18]. The time of odor appearance, end time, and odor description were recorded by the selected panelists.
Odor activity value (OAV)
-
OAV is often used to evaluate and screen the contribution of aroma-active compounds to the aroma formation of samples[19]. According to the calculation formula, OAV is the ratio of concentration of aroma-active compounds relative to their respective threshold in water, the compound concentration is the absolute concentration corrected by the GC-MS standard curve, and the odor threshold in water is obtained from previous literature[16,20−24].
Aroma extraction dilution analysis (AEDA)
-
Aroma water was diluted in gradient with water at 2n to obtain dilution multiples of 1:2, 1:4, 1:8, 1:16, etc. GC-O detection, based on sniffing until the aroma-active compound odor disappears, was determined as the FD factor of the compound.
Aroma-active compound release pattern
-
In order to study the content changes of aroma-active compounds of samples with heating time during the aroma enrichment process, segmented interception of aroma extracts was performed during the extraction of aroma substances, with each 50 mL segment being collected 20 times. The relative quantification was performed by adding 0.01 mL of n-decanol (0.13 mg/kg) to analyze the release pattern of aromatic compounds. The process was repeated three times.
-
The study analyzed crushed samples, as well as aroma water samples. The number of aroma-active compounds in the samples and their relative content was examined to select the best treatment method. As shown in Fig. 3, 24 aroma-active compounds were identified in the detected values of the two forms of samples, such as d-limonene, linalool, 2-methoxy-4-vinylphenol, furfural, and α-terpinene, etc.; aroma-active compounds specific to the heated enriched aroma-active water such as p-cymene, 1-octanol, 4-terpineol, etc. There were 19 kinds of aroma-active compounds particular to the directly crushed samples such as undecanal, citronellol, neral, etc. There were eight kinds of aromatic compounds.
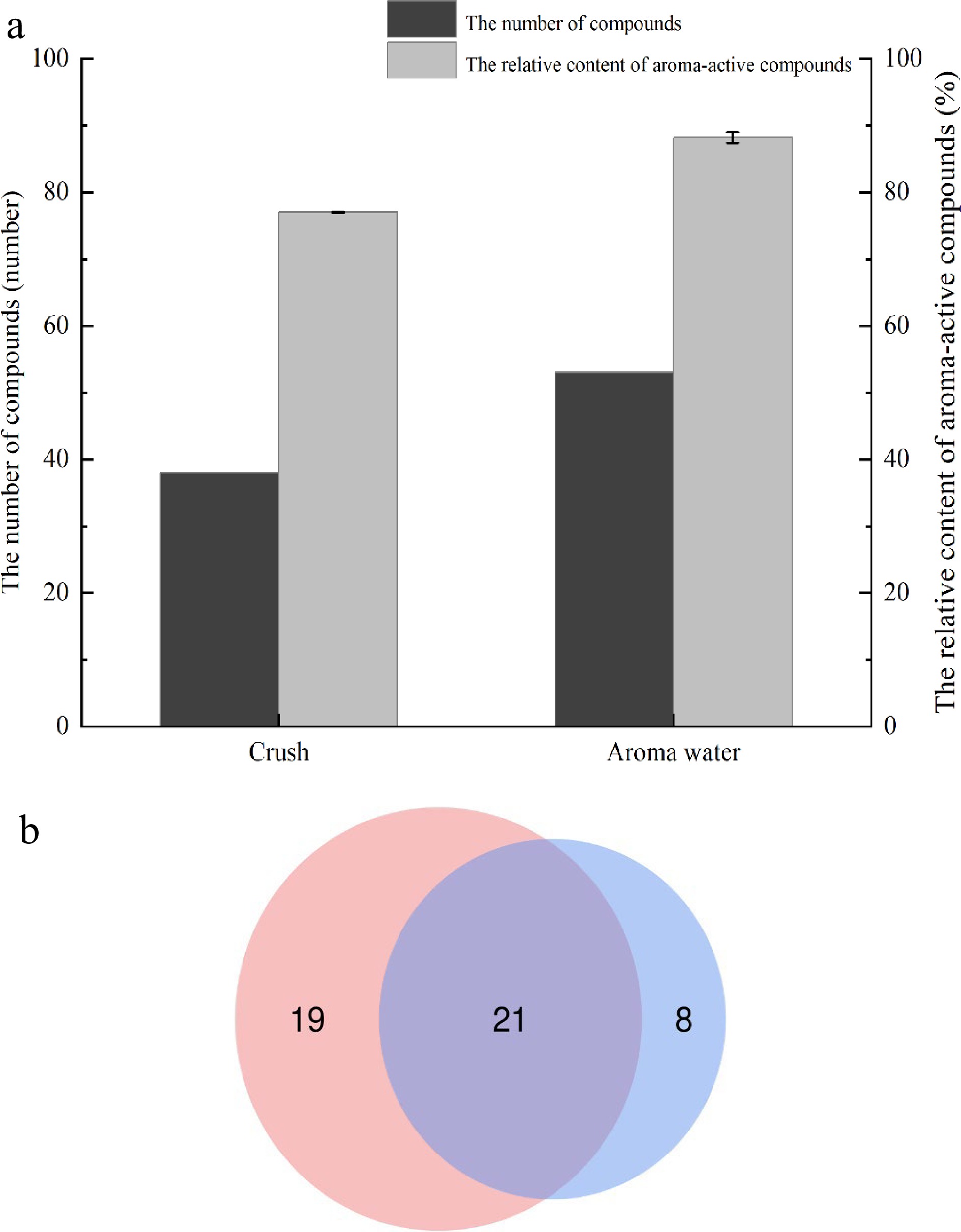
Figure 3.
Comparison of aroma-active compounds in two forms, (a) concentration and number; (b) the number of compounds.
The relative content of aroma-active compounds was collected using the normalization method. The sum of the relative content of aroma-active compounds in the water sample was greater than that of the crushed sample, and the aroma-active compounds were also more in the aroma water sample. In conclusion, the experimental feasibility of studying the aroma-active compounds of aroma water met expectations, and the aroma-active compounds of the Xinhui Chenpi samples could be characterized in this way.
Quantitative analysis of aroma-active compounds
-
The GC-MS qualitative analysis of the volatile compounds contained in the aroma water detected a total of 24 volatile aroma-active compounds with a matching index greater than 80% compared with the database, including six alcohols, four olefins, three aldehydes, two ketones, one acid, three phenols and three esters, as shown in Table 1. The corresponding standard curves were established for each compound with good linearity (equations of standard curves, where y is the area of the peak of an authentic standard, and x is the concentration of the authentic standard). Among them, d-limonene (3,291.64 mg/kg) showed highest in all compounds, followed by linalool (561.39 mg/kg), 4-terpineol (370.81 mg/kg), γ-terpinene (354.97 mg/kg), furfural (274.80 mg/kg), 2-methoxy-4-vinylphenol (253.67 mg/kg) and α-terpineol (250.54 mg/kg). According to the available literature, compounds such as d-limonene, γ-terpinene, linalool, and myrcene are essential components of the aroma composition of Chenpi[25−30], and substances such as 4-terpineol, furfural, 2-methoxy-4-vinylphenol, and p-cymene in the Chenpi aroma-active substances have been rarely reported previously. In the present study, octanal (78.71 mg/kg), thymol (38.18 mg/kg), cymenol (29.62 mg/kg), and myrcene (21.19 mg/kg) were the compounds that contributed significantly to the aroma of Xinhui Chenpi.
Table 1. The concentration of volatile compounds detected in aroma water samples.
No Time
(min)RI Compound Molecular formula Calibration curves R2 Linear range
(mg/kg)Content
(mg/kg)aRSD (%)b 1 4.12 840 Furfural C5H4O2 y = 2.08E+09x + 1.76E+06 R2 = 0.9965 92.34-4.62 274.80 17.64 2 7.42 983 Myrcene C10H16 y = 3.55E+11x – 1.25E+07 R2 = 0.9939 1.37-0.07 21.19 24.52 3 7.74 1002 Octanal C8H16O y = 3.05E+10x – 4.37E+06 R2 = 0.992 2.34-0.12 78.71 31.41 4 8.19 1009 α-Terpinene C10H16 y = 2.73E+11x – 6.85E+06 R2 = 0.9919 0.51-0.03 3.47 27.55 5 8.44 1017 p-Cymene C10H14 y = 4.27E+11x – 5.02+05 R2 = 0.9959 0.41-0.02 12.85 13.06 6 8.60 1023 d-Limonene C10H16 y = 1.06E+11x + 2.55E+08 R2 = 0.9916 127.98-6.40 3291.64 28.02 7 9.53 1054 γ-Terpinene C10H16 y = 1.50E+11x – 3.95E+07 R2 = 0.9941 12.92-0.65 354.97 26.94 8 9.87 1071 1-Octanol C8H18O y = 1.62E+11x + 6.65E+06 R2 = 0.9923 0.40-0.04 3.32 20.64 9 10.91 1101 Linalool C10H18O y = 1.37E+11x + 9.32E+08 R2 = 0.9902 33.41-3.34 561.39 17.00 10 11.08 1105 Nonanal C9H18O y = 1.61E+11x + 4.96E+06 R2 = 0.9915 0.38-0.04 6.45 18.43 11 13.58 1168 1-Nonanol C9H20O y = 1.05E+11x + 3.83E+07 R2 = 0.9942 1.38-0.14 10.10 21.04 12 13.90 1192 4-Terpineol C10H18O y = 9.17E+10x + 3.02E+08 R2 = 0.9909 13.14-0.66 370.81 25.28 13 14.40 1202 α-Terpineol C10H18O y = 1.08E+11x + 1.66E+08 R2 = 0.9908 19.11-1.91 250.54 30.68 14 14.70 1219 Octanoic acid C8H16O2 y = 2.00E+09x + 1.41E+07 R2 = 0.9946 - - 15 16.52 1291 (-)-Carvone C10H14O y = 2.07E+11x + 4.62E+07 R2 = 0.9909 092-0.09 6.18 24.79 16 16.92 1304 Geraniol C10H18O y = 2.00E+11x + 4.34E+07 R2 = 0.9901 1.39-0.14 16.99 28.71 17 18.51 1358 Thymol C10H14O y = 6.54E+11x + 6.06E+08 R2 = 0.9922 7.88-0.79 38.18 26.40 18 18.89 1371 Cymenol C10H14O y = 6.14E+11x + 7.10E+07 R2 = 0.9916 0.77-0.08 29.62 21.95 19 19.40 1340 2-Methoxy-4-vinylphenol C9H10O2 y = 3.41E+10x + 2.15E+08 R2 = 0.9911 31.33-3.92 253.67 30.06 20 21.00 1360 Citronellyl acetate C12H22O2 y = 8.53E+11x + 1.20E+07 R2 = 0.9907 0.14-0.01 1.68 26.58 21 21.56 1380 n-Decanoic acid C10H20O2 y = 3.55E+11x – 1.42E+07 R2 = 0.9924 0.48-0.05 7.17 31.25 22 22.22 1385 Geranyl acetate C12H20O2 y = 1.01E+12x + 6.69E+06 R2 = 0.9906 0.08-0.008 0.54 30.38 23 23.22 1453 Methyl methanthranilate C9H11NO2 y = 1.16E+11x + 7.55E+06 R2 = 0.9914 0.53-0.05 15.20 10.21 24 26.40 1486 β-Ionone C13H20O y = 5.98E+11x + 3.47E+06 R2 = 0.9994 0.06-0.007 1.94 23.68 a: The data of concentration is mean (n = 3 for aroma water sample). b: The RSD is standard deviation (n = 3 for aroma water sample). −: Indicates no detection results. GC-O verifies the aroma-active compounds in samples
-
The odor activity of 24 aroma-active compounds detected by GC-MS was characterized by GC-O combined with AEDA and expressed as FD dilution factor, which was used in combination with the OAV value of each aroma-active compounds to verify its contribution to the aroma-active compounds of Xinhui Chenpi[24,31,32]. As shown in Table 2, the FD dilution factors ranged from 2 to 8,192, the higher FD factors, the stronger the odor of the compounds, and the greater the contribution to the aroma of the sample. The compounds with the highest FD factors were linalool (8,192) with sweet, cymenol (8,192) with pungent and refreshing odors, 2-methoxy-4-vinylphenol (8,192) with pungent and flower odors, followed by β-ionone (4,096) with woody odor, 4-terpineol (2,048) with woody and loamy incense odors, α-terpineol (2,048) with woody and flower odors, (−)-carvone (2,048) with mint and spicy odors, geranyl acetate (2,048) with medicinal odor, n-decanoic acid (2,048) with flower odor. The aroma-active compounds of Xinhui Chenpi are mainly alcohols, olefins, esters and aldehydes, and other compounds such as ketones and phenols. Among them, alcohols and olefins accounted for the highest proportion, such as d-limonene and linalool, which mainly show the typical orange and sweet odor of Chenpi, are the most abundant compounds in Xinhui Chenpi, with FD factors of 32 and 8,192, respectively. Aldehydes also contributed to the aroma-active formation of Chenpi, such as octanal, which has a typical orange flavor with an FD factor of 512. Furfural has a nut odor, which has rarely been reported in previous studies of aroma-active compounds of Chenpi.
Table 2. The FD factor of aroma-active compounds.
No Compound Odor description* FD** 1 Furfural Nut 8 2 Myrcene Pungent 32 3 Octanal Orange flavor 512 4 α-Terpinene Wax, orange 16 5 p-Cymene Refreshing 8 6 d-Limonene Citrus 32 7 γ-Terpinene Woody 8 8 1-Octanol Oily, fruity 128 9 Linalool Flowers, sweet 8192 10 Nonanal Oily, sweet, orange 8 11 1-Nonanol Orange scent 2 12 4-Terpineol Woody, loamy incense 2048 13 α-Terpineol Flowers, woody 2048 14 Octanoic acid Fruity 32 15 (−)-Carvone Mint, spicy 2048 16 Geraniol Rose 512 17 Thymol Medicine 1024 18 Cymenol Pungent, refreshing 8192 19 2-Methoxy-4-vinylphenol Pungent, flowers 8192 20 Citronellyl acetate Flowers 16 21 n-Decanoic acid Flowers 2048 22 Geranyl acetate Medicine 2048 23 Methyl methanthranilate Orange, flowers 1024 24 β-Ionone Woody 4096 * Description of the sniffing results by the sensory evaluator (n = 3 for sensory evaluator).
** Maximum dilution of the aroma-active compound.To identify the contribution of aroma-active compounds to the aroma of Xinhui Chenpi, the OAV was calculated to verify that each compound's odor threshold was known from the literature. The literature has reported that an aroma-active compound OAV ≥ 1 indicates that the compound contributes to aroma formation[24,33,34]. As shown in Table 3, the compounds have been ranked in order of OAV from largest to smallest. Octanoic acid was not detected in the content; other than that, all 22 aroma-active compounds revealed had OAV > 1 in aroma water. The results showed that d-limonene had the highest OAV (24,027) in all compounds, followed by linalool (20,050), 2-methoxy-4-vinylphenol (13,351), geraniol (1,699), thymol (382), octanal (342), α-terpineol (291), β-ionone (231). The above results show that the most important aroma-active compounds were d-limonene and linalool in Chenpi. Notably, although the contents of geraniol (16.99 mg/kg) and β-ionone (1.94 mg/kg) were not very high, their OAVs were the highest, because the thresholds were low (0.01 and 0.0084 mg/kg).
Table 3. The results of OAVs calculation of aroma-active compounds.
No Compound Concentration
(mg/kg)Odor threshold in
water (mg/kg)OAV 6 d-Limonene 3291.64 0.14b 24026.58 9 Linalool 561.39 0.03a 20049.61 19 2-Methoxy-4-vinylphenol 253.67 0.02b 13351.10 16 Geraniol 16.99 0.01a 1699.27 17 Thymol 38.18 0.10b 381.80 3 Octanal 78.71 0.23b 342.23 13 α-Terpineol 250.54 0.86b 291.32 24 β-Ionone 1.94 0.01c 231.49 18 Cymenol 29.62 0.18b 164.56 8 1-Octanol 3.32 0.02b 144.30 15 (-)-Carvone 6.18 0.07a 92.26 1 Furfural 274.80 3.00c 91.60 10 Nonanal 6.45 0.10a 64.50 12 4-Terpineol 370.81 6.40a 57.94 21 n-Decanoic acid 7.17 0.13b 55.19 23 Methyl methanthranilate 15.20 0.35b 43.54 2 Myrcene 21.19 0.67a 31.63 11 1-Nonanol 10.10 1.00a 10.10 7 γ-Terpinene 354.97 55.00c 6.45 22 Geranyl acetate 0.54 0.15a 3.60 5 p-Cymene 12.85 7.20b 1.78 20 Citronellyl acetate 1.68 1.00b 1.68 4 α-Terpinene 3.47 2.40b 1.45 14 Octanoic acid − 0.86b − Odor thresholds in water found in the literature. a: Indicates reference[18]. b: Indicates reference[35]. c: Indicates reference[21]. −: Indicates no detection. Release pattern of aroma-active compounds
-
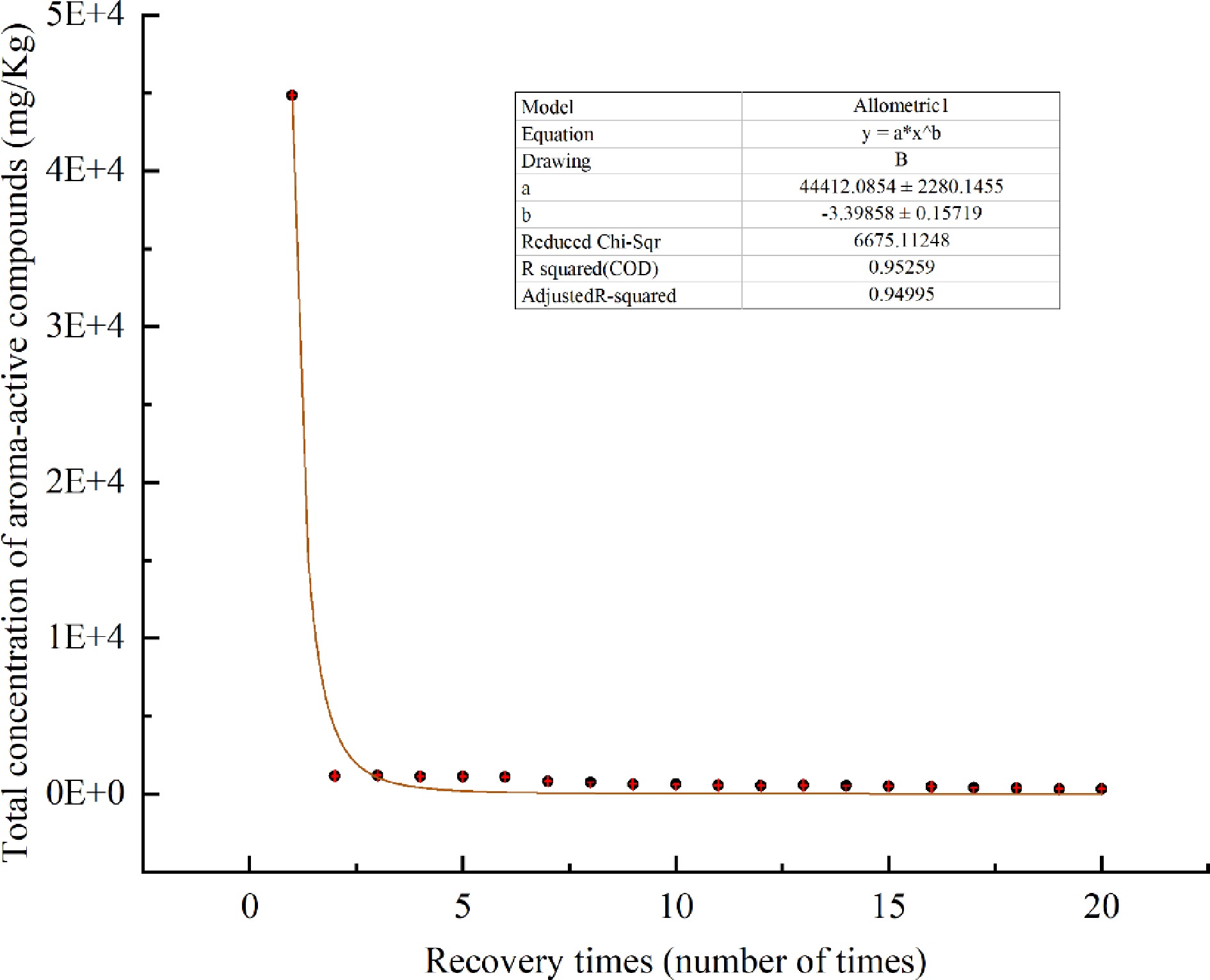
The changes of 24 aroma-active compounds with heating time when samples were heated to enrich the aroma-active compounds were averaged over three replicate values. As shown in Table 4 and Fig. 4, a decreasing pattern was seen, with a sharp decrease after the 1st time, and a slight increase from the 2nd to the 3rd time, followed by a slight decrease.
Table 4. Changes in the concentration of aroma-active compounds of Chenpi with extraction time.
Time (min) Compound Concentration (mg/Kg) 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 4.12 Furfural 4.07 2.46 2.49 2.48 2.59 2.69 2.71 2.70 2.53 2.41 2.24 2.32 2.63 2.58 2.54 2.66 2.70 2.61 2.51 2.21 7.42 Myrcene 1053.51 0.37 0.25 0.22 0.31 0.40 0.19 0.51 0.20 0.37 0.35 0.39 1.09 0.44 0.85 0.69 1.20 0.77 0.77 0.98 7.74 Octanal 69.18 4.19 6.48 5.87 6.80 5.30 6.97 5.64 5.25 5.56 5.53 5.08 6.29 4.98 5.10 4.96 5.89 7.61 5.99 4.75 8.19 α-Terpinene 114.06 0.56 0.59 0.36 0.52 0.51 0.44 0.56 0.50 0.52 0.56 0.42 0.75 0.59 0.78 0.72 1.10 0.97 0.88 1.11 8.44 p-Cymene 53.16 1.18 1.18 0.89 1.24 1.18 0.88 1.38 0.82 1.16 1.17 1.17 1.34 1.34 0.72 1.71 1.58 1.41 1.33 1.97 8.60 d-Limonene 36762.66 20.30 20.33 21.24 20.29 21.32 18.23 44.92 20.00 32.18 30.15 33.80 45.82 36.91 42.08 30.50 20.79 21.79 29.84 26.29 9.53 γ-Terpinene 3591.33 1.63 1.52 1.72 1.81 1.61 1.59 3.73 1.78 2.80 2.76 3.12 4.00 3.05 3.77 4.13 3.67 4.07 3.65 3.33 9.87 1-Octanol 20.45 2.86 2.86 1.72 1.87 1.81 1.42 1.08 1.17 1.02 0.88 1.11 0.75 0.58 0.47 0.39 0.45 0.55 0.42 - 10.91 Linalool 1080.19 396.41 408.52 350.55 346.02 352.47 270.33 240.68 232.30 211.12 194.25 173.38 180.08 161.11 140.28 138.62 121.05 112.08 61.12 59.41 11.08 Nonanal 58.69 3.08 3.30 3.00 3.72 3.61 3.49 2.96 2.91 3.43 3.59 3.29 3.56 3.28 3.50 3.56 3.54 3.34 3.04 3.39 13.58 1-Nonanol 16.30 7.72 7.62 7.31 6.96 6.80 6.62 5.51 4.71 3.90 3.23 3.29 3.28 3.23 3.29 3.36 2.98 3.00 2.97 2.79 13.90 4-Terpineol 498.61 147.36 144.59 136.17 133.69 123.46 117.43 108.56 112.45 110.63 104.22 104.54 103.89 103.29 102.11 106.10 89.27 84.65 50.56 79.04 14.40 α-Terpineol 394.39 134.60 127.01 114.39 113.81 100.99 79.61 68.79 70.19 67.84 61.17 56.70 56.03 56.12 55.62 55.00 43.90 41.62 41.01 40.69 14.70 Octanoic acid − 4.28 4.11 − − − − − − − − − − − − − − − − − 16.52 (−)-Carvone 30.11 15.90 14.41 13.32 11.47 9.65 6.81 5.31 4.92 4.14 3.34 2.86 2.87 2.10 1.84 1.55 1.04 0.98 0.91 0.78 16.92 Geraniol 24.16 18.26 17.67 15.09 14.85 13.24 10.37 7.84 8.16 7.78 6.63 5.70 5.60 5.28 4.82 4.95 3.85 3.43 2.84 2.66 18.51 Thymol 207.17 187.75 184.75 180.91 168.43 134.25 103.75 86.04 83.95 74.10 57.64 54.99 51.60 47.08 40.06 38.67 28.19 23.13 18.93 17.40 18.89 Cymenol 72.98 152.32 175.42 189.58 198.40 198.78 116.71 82.08 20.87 20.15 20.01 20.93 21.03 27.24 30.94 24.91 8.89 7.49 14.32 13.62 19.40 2-Methoxy-4-vinylphenol 22.97 23.21 31.70 62.03 68.22 71.49 54.06 53.21 51.45 50.46 50.34 50.10 49.72 48.85 48.53 49.54 49.70 44.43 44.72 39.54 21.00 Citronellyl acetate 21.93 5.32 5.30 5.20 5.55 5.26 3.65 3.40 1.73 1.52 1.48 1.41 1.73 1.67 1.48 1.44 1.56 1.46 1.86 2.09 21.56 n-Decanoic acid − 12.05 14.52 14.52 15.45 16.58 12.05 10.00 8.03 7.42 7.31 7.73 7.30 7.33 5.27 5.73 5.61 5.11 5.72 5.50 22.22 Geranyl acetate 7.13 1.12 1.16 1.29 1.16 1.29 1.20 1.18 1.18 1.16 1.16 1.15 1.16 1.04 1.16 1.17 1.09 1.09 1.18 1.13 23.22 Methyl methanthranilate − 5.12 6.93 5.65 5.02 4.89 4.84 4.82 3.10 2.82 2.52 2.16 1.98 1.68 1.54 1.49 1.43 1.05 0.89 0.73 26.40 β-Ionone 7.72 6.95 5.23 4.88 4.88 3.82 3.21 3.02 2.85 2.62 2.36 2.23 2.06 1.84 1.72 1.66 1.46 1.09 0.99 0.99 Total concentration 44839.78 1155.54 1187.93 1138.38 1132.86 1081.40 826.57 743.90 641.10 615.12 562.88 537.87 554.55 521.93 498.49 483.52 400.94 373.74 326.45 310.42 - indicates no detection. As shown in Table 4, the nine compounds including furfural, myrcene, octanal, α-terpinene, p-cymene, d-limonene, γ-terpinene, nonanal, and geranyl acetate showed a pattern of rapidly decreasing to a minimum and then being continuously maintained; the 10 compounds including 1-octanol, linalool, 1-nonanol, 4-terpineol, α-terpineol, (−)-carvone, geraniol, thymol, citronellyl acetate, β-ionone showed a gradually decreasing pattern; cymenol, 2-methoxy-4-vinylphenol, n-decanoic acid, and methyl methanthranilate showed a pattern of increasing first, reaching a maximum value, and then gradually decreasing; octanoic acid was detected only in the 2nd and 3rd times.
The alcohols and ketones in the aroma-active compounds of Xinhui Chenpi mainly showed a gradually decreasing pattern. For example, among the alcohols, linalool (1,080.19 mg/kg), 4-terpineol (498.61 mg/kg), α-terpineol (394.39 mg/kg) are of the highest proportion, linalool decreased from 1,080.19 mg/kg to 59.41 mg/kg; 4-terpineol decreased from 498.61 mg/kg to 79.04 mg/kg; α-terpineol decreased from 394.39 mg/kg to 40.69 mg/kg. This indicates that the alcoholic compounds of Xinhui Chenpi are persistent in aroma and slow in release.
The olefins and aldehydes mainly showed a pattern of rapidly decreasing at the lowest level and then being continuously maintained. Among the olefins, d-limonene (36,762.66 mg/kg), γ-terpinene (3,591.33 mg/kg), myrcene (1,053.51 mg/kg) and α-terpinene (114.06 mg/kg) are of the highest proportion, and d-limonene decreased from 36,762.66 mg/kg to 20.30 mg/kg in the 2nd and maintained since then; The γ-terpinene decreased from 3,591.33 mg/kg to 1.63 mg/kg in the 2nd and maintained since then; Myrcene decreased from 1,053.51 mg/kg to 0.37 mg/kg in the 2nd and maintained since then; α-terpinene decreased from 114.06 mg/kg to 0.56 mg/kg in the 2nd and maintained at a consequent level after that. It indicates that the olefins of Xinhui Chenpi are easily soluble and released extremely fast.
Other compounds, such as thymol gradually decreased from 207.17 mg/kg to 17.40 mg/kg; cymenol gradually increased from the initial 72.98 mg/kg, reaching a maximum value of 198.78 mg/kg at the 6th pass, and then started to decrease to 13.62 mg/kg; 2-methoxy-4-vinylphenol gradually increased from the initial 22.97 mg/kg to a maximum of 71.49 mg/kg at the 6th time and then started to decline to 39.54 mg/kg. This indicates that the aroma was persistent and remained in the release phase during the heating extraction.
Furfural differed from the other substances, with a small amount of 4.07 mg/kg initially, which decreased to 2.46 mg/kg in the 2nd and then continued to be maintained, indicating that Furfural is more difficult to volatilize than the other substances.
-
Aroma water obtained by heating and enrichment has obvious advantages. The aroma water of Xinhui Chenpi was comprehensively analyzed using the methods of GC-MS, GC-O and AEDA. It was concluded that linalool, d-limonene, 2-methoxy-4-vinylphenol, and α-terpineol were the four aroma-active compounds with high content and high contribution to the aroma formation, which were identified as the essential aroma substances of Xinhui Chenpi, and mainly presenting orange, sweet, spicy, woody and floral aromas. The heat release pattern was studied, and the results showed that the content of the aroma-active compounds generally showed a decreasing pattern with the heating time in the process of heating enrichment. In particular, the overall content decreased sharply after the 1st time, and it was speculated that this occurred because the Chenpi samples were soaked in heated water before being enriched by heating and distillation. However, this conclusion needs to be further verified. Although the overall content changed significantly after the 1st time, and combined with the respective emission pattern of 24 aroma-active compounds, it is difficult to volatilize all the aroma-active compounds in a short time, so this experiment recommends that the first 10 times of aroma water can be collected to achieve a shorter time and enrichment of the most aroma-active compounds.
This work was supported by Youth Top Talents Project in Tianjin Special Support Program, the Science and technology Project of Tianjin (Grant No. 21ZYCGSN00410), and the Science and technology Project of Tianjin (Grant No. 21YDTPJC00910). This work was supported by the National Natural Science Foundation of China [32102004]; the Project funded by China Postdoctoral Science Foundation [2021M701169, 2022T150206], the China Agriculture Research System of MOF and MARA; the Construction of World Big-leaf Tea Technology Innovation Center and Industrialization of Achievements [202102AE090038]; and the Scientific and Technological Talents and Platform Plan (Academician Expert Workstation) [202104AC100001-B01].
-
The authors declare that they have no conflict of interest.
- Copyright: © 2023 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Yang D, Wu X, Shi H, Zhang J, Wang C. 2022. Essential aroma substances and release pattern of Xinhui Chenpi. Beverage Plant Research 2:22 doi: 10.48130/BPR-2022-0022
Essential aroma substances and release pattern of Xinhui Chenpi
- Received: 12 September 2022
- Accepted: 16 November 2022
- Published online: 09 December 2022
Abstract: This study focuses on analyzing aqueous solutions of aroma-active compounds in Xinhui Chenpi distilled after being heated with headspace solid-phase microextraction and gas chromatography-mass spectrometry (HS-SPME-GC/MS). Feasibility of this method was also tested by comparison with crushed samples. The study also analyzed the aqueous solutions of the aroma-active compounds employing gas chromatography-olfactometry (GC-O) defense, aroma extraction dilution analysis (AEDA) and odor activity value (OAV) as well as their emission patterns. According to the study, there are 24 major aroma-active detected in the aqueous solution. Linalool, d-limonene, 2-methoxy-4-vinylphenol, and α-terpineol with sweet, spicy, and woody aroma contributed the most to the aroma-active compounds and were considered to be the essential aroma substances of Chenpi. When heated, the aroma of aromatic-active compounds rich in Chenpi volatilized rapidly and the release dropped dramatically in a short time. The substantial aroma-active compounds can be collected from the first ten released segments.