-
Fusarium wilt, caused by Fusarium oxysporum f. sp. cubense (Foc), is an economically important disease affecting banana. Wilting results from the restrictive movement of water in the vascular bundles[1]. However, part of the pathogenesis and invasion of plants by Foc can be attributed to toxic metabolites produced by the fungus[2]. Major toxins of Foc include fusaric acid and beauvericin, which act as virulence factors for infection by causing significant browning of vascular tissues and plant necrosis[3−5]. Fusaric acid (FA) is a non-specific toxin many fungal pathogens produce[6]. Based on previous studies, FA supports disease development through the induction of cell membrane early super polarization[3], H+ pumping, K+ leaking suppression[7], mineral chelation[8] and inhibition of plant defensive enzymes activity, leading to reduced cell viability[9], changes in membrane permeability and potential[7,10], and production of reactive oxygen species[11]. Previously, Brown et al. recorded that lack of FA production did not affect the virulence of F. oxysporum in cacti or F. verticillioides in maize seedlings[12]. In contrast, several studies reported the importance of FA production in the virulence of Foc in banana[2,5,13].
Meanwhile, beauvericin (BEA) is a secondary metabolite produced by several species from the Fusarium fujikuroi species complex[14] and the entomopathogenic fungus Beauveria bassiana[15]. Although FA and BEA are major toxins of Foc, the two are not considered mycotoxins with significant human, animal, food, and feed safety risks[2]. Still, some studies have documented physiologic disorders in experimental animals and human cell lines treated with FA[16] and BEA[17−19].
Phytotoxin insensitivity of plant cell lines can be used as a potential marker in breeding programs for early screening of resistance against plant pathogens[20,21]. The approach requires a significant correlation between toxin sensitivity and host susceptibility to the pathogens producing them[22]. Several studies have taken advantage of this via somaclonal cell selection - a technique that utilizes phytotoxins as selective agents for developing resistant clones against economically important diseases such as common scab of potato[23,24]. Production of Fusarium wilt-resistant date palm[22], carnation[25], abaca[26], and banana cv. 'Maca'[27] has also been reported by cell selection. Yet, little attention has been given to the phytotoxicity of Foc metabolites in banana.
This study determined the phytotoxicity of FA and BEA in leaf tissues, multiple bud clumps, tissue-cultured plantlets, and calli in-vitro. These results would aid in determining the appropriate concentration of the metabolites for somaclonal cell selection.
-
The test substances: fusaric acid (≥ 99% purity, CAS No. 536-69-6) and beauvericin (≥ 97% purity, CAS No. 28048-05-5), were obtained from Sigma-Aldrich. The toxins were dissolved in 0.05% methanol to obtain a 10 mM stock solution. The stock solution was then filter-sterilized by passing through Whatmann no.1 filter paper (125 mm size) (CAT no. 1001-125, GE Healthcare Life Sciences) and stored in a 4 °C refrigerator until use.
Phytotoxicity testing in different tissues of banana
-
Different concentrations of fusaric acid (FA) and beauvericin (BEA) in tissue-cultured Cavendish plantlets, multiple bud clumps (MBC), callus, and attached leaf tissues were determined. FA stock solution was further diluted to 0, 1, 5, 10, 20, 30, 50, 70, 100 μm concentrations by adding desired concentrations in culture medium. Meanwhile, 0, 1, 5, 10, and 20 μm BEA were tested. These concentrations were selected based on the previous findings of Li et al.[2] on the phytotoxicity of FA and BEA in tissue-cultured plantlets of banana.
For tissue-cultured plantlets tests, tissue-cultured 'Grand Nain' and 'Lakatan' were obtained from the National Plant Genetic Resources Laboratory (NPGRL), IPB, UPLB. We used 'Grand Nain' because it is commercially propagated in Mindanao, where the Foc TR4 is present. 'Lakatan' was also used since it is a local variety commonly grown in the Philippines, especially by small-scale growers. The tissue-cultured materials were cultured and maintained in the laboratory for micropropagation. The plantlets were propagated by cutting a 2-cm segment of tissue containing the pseudostem and apical meristems. The tissues were then cut in half and cultured in a multiplication-inducing medium which consisted of Murashige and Skoog medium[28] + 3 mg·L−1 BAP. Four segments were placed in each glass jar. Cultures were maintained in 14 h light with temperature ranging from 20 ± 5 °C. Cultures were routinely transferred in the same medium every two weeks to prevent browning. One-month-old tissue-cultured plantlets with roots were transferred in test tubes containing 10 ml liquid medium and treated with FA and BEA by pipetting desired concentration in the medium. Plantlets treated with sterile distilled water served as control. Cultures were maintained in 8 h light at ambient temperature (20 ± 5 °C).
For multiple bud clumps studies, the protocol of Matsumoto et al.[27] was used to produce multiple bud clumps. Rhizome tissues were cut from 'Grand Nain' and 'Lakatan' shoot cultures and were grown in MS medium containing BAP (5 mg·L−1) to initiate multiple bud clump (MBC) formations. Cultures were maintained under the same conditions as previously mentioned. One-month-old bud clumps were harvested from a multiple bud clump and transferred into test tubes containing 10 ml liquid medium. Five individual buds were placed in each tube and treated with FA and BEA as previously described.
Corm segments (approximately 2 cm in size) were cut from one-month-old shoot cultures of banana cv for callus studies. 'Grand Nain' and 'Lakatan' and inoculated on MS basal medium supplemented with 2,4-dichloro phenoxy acetic acid (1.0 mg·L−1) and myo-inositol (50 mg·L−1). Cultures were stored in dark conditions under a chamber covered with black cloth and incubated at ambient temperature (20 ± 5 °C). Individual calli were harvested eight weeks post-inoculation and transferred into test tubes containing 10 ml liquid medium. Five individual calli were placed in each tube and treated with FA and BEA as previously described.
For the attached leaf assay, tissue-cultured 'Grand Nain' and 'Lakatan' with multiple shoots were grown in basal MS medium for rooting. After one month, the rooted plantlets were acclimatized and transferred to plastic bags containing sterile soil in the nursery. The plantlets were grown for two months and watered daily. FA and BEA were diluted to the desired concentration by adding sterile distilled water in 1.5 ml Eppendorf tubes. The attached leaves were inoculated by pipetting 20 μl of solution in wounded (pricked using a syringe needle) tissues. The experiment was repeated twice.
Data collection and statistical analyses
-
The phytotoxicity of fusaric acid (FA) and beauvericin (BEA) in different banana tissues was assessed ten days post-inoculation. Wilting rates were measured ocularly based on the 10-point arbitrary scale produced. A one-way ANOVA was performed to identify significant differences between treatments using Statistical Tool for Agricultural Research (STAR Nebula) with a 95% confidence level.
-
Wilting of different tissues treated with fusaric acid (FA) and beauvericin (BEA) was observed ten days post-inoculation. Ten-point visual hedonic browning scales were developed based on varying degrees of wilting (Fig. 1a−c). The rating scales were used to measure the degree of phytotoxicity in tissue-cultured plantlets (TC plantlets), multiple bud clumps (MBC), and callus cultures.
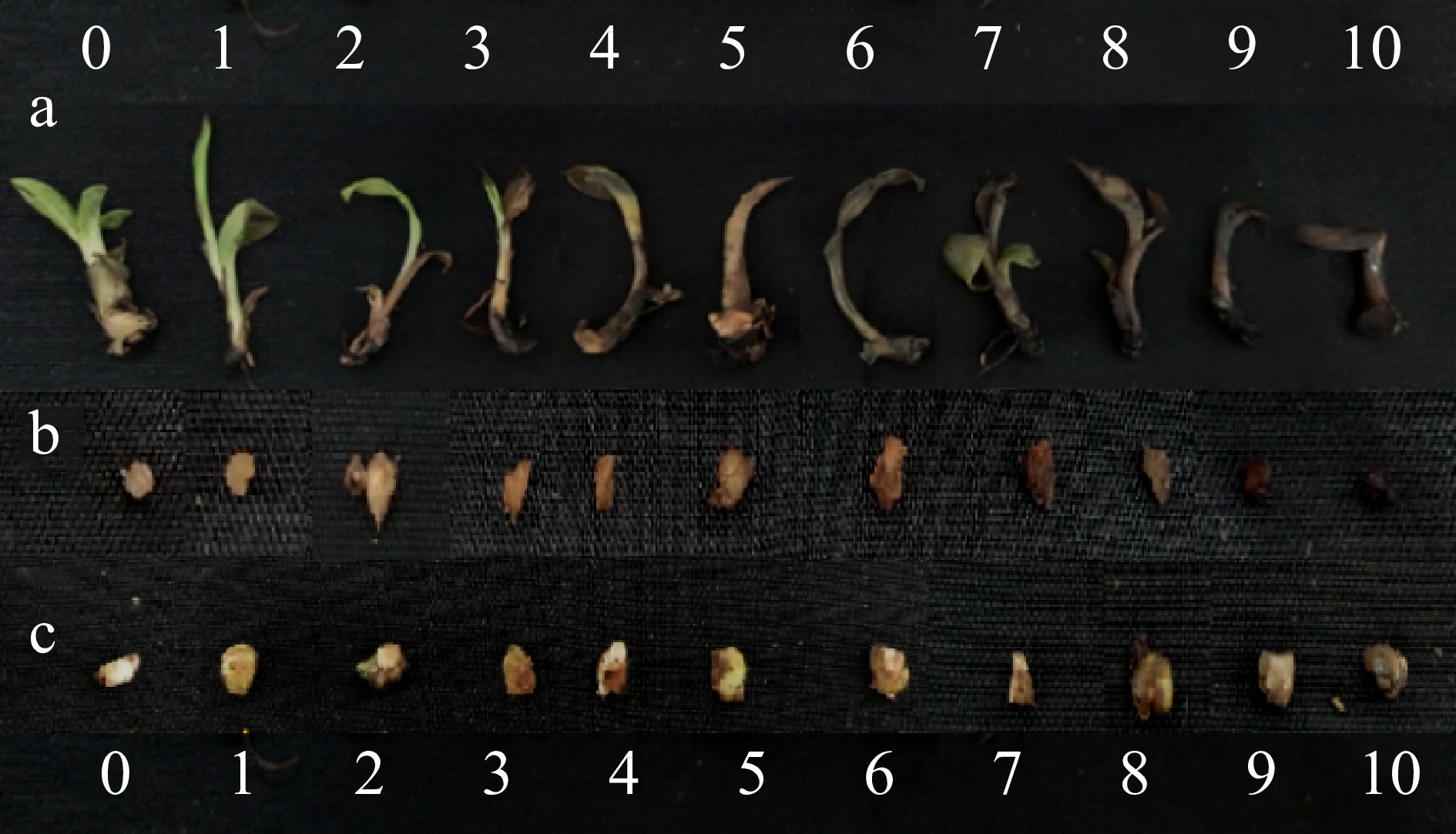

Figure 1.
Ten point rating scales produced in this study for measuring degree of wilting in (a) tissue cultured plantlets, (b) calli, and (c) multiple bud clumps, as caused by Fusarium toxins.
Overall, there is an increasing trend between the toxin concentration and the degree of browning observed (Figs 2−4). For both 'Grand Nain' and 'Lakatan,' the highest browning rates were observed in tissues treated with 100 μm FA and 20 μm BEA (Tables 1 & 2). A higher degree of browning was recorded in 'Lakatan' TC plantlets, MBC, and callus than in 'Grand Nain,' except in TC plantlets treated with 20 μm BEA, where a higher average browning rate score was observed in 'Grand Nain' (9) than in 'Lakatan' (Tables 1 & 2).
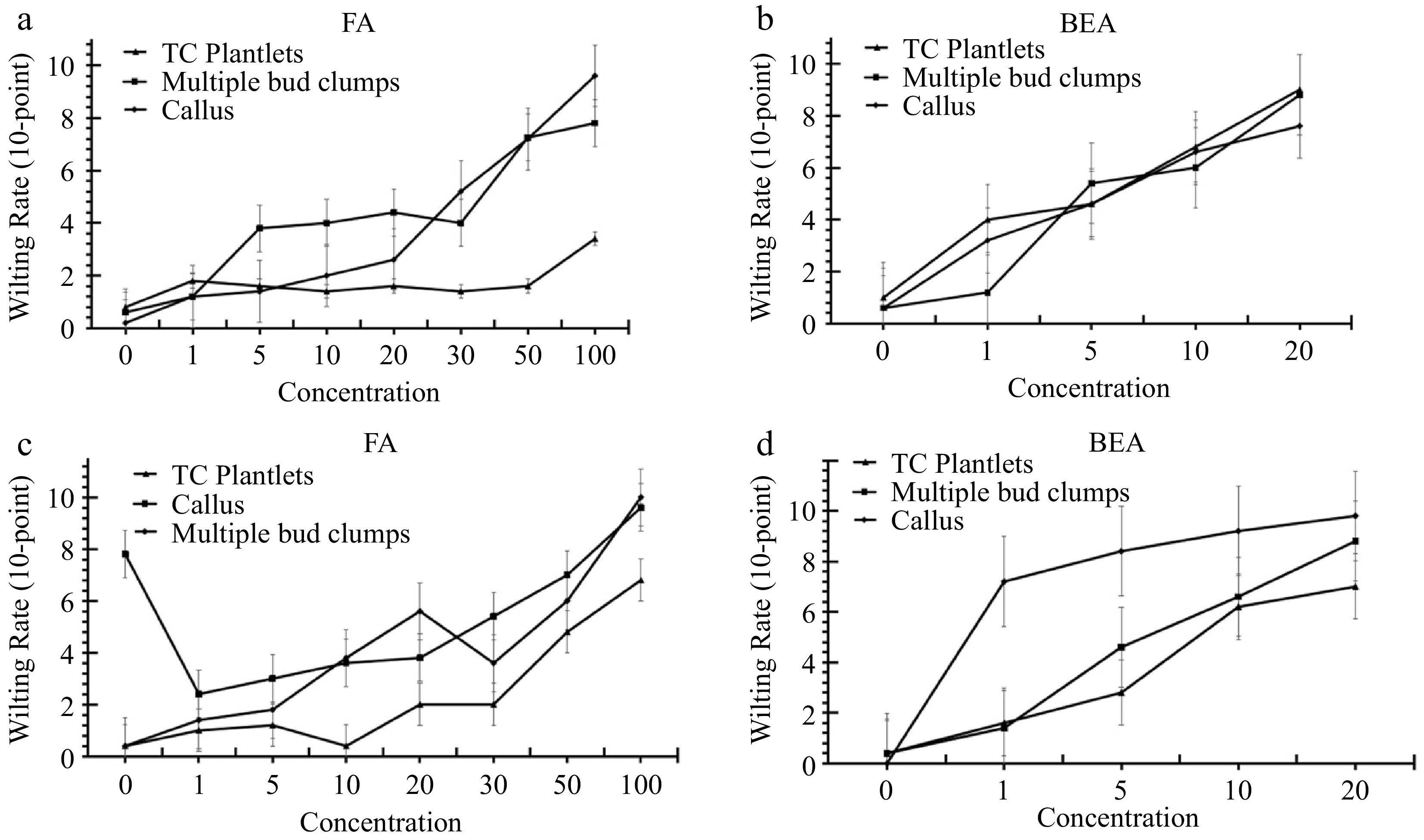
Figure 2.
Browning rates of tissue-cultured plantlets, calli, and multiple bud clumps of banana cvs. (a), (b) ‘Grand Nain’ and (c), (d) ‘Lakatan' treated with varying concentrations of FA and BEA at 10 d post-incubation.
Average browning rates of 'Grand Nain' TC plantlets, MBC, and callus treated with 100 μm FA were 3.4, 7.8, and 9.6, respectively, using the 10-point system (Table 1). Meanwhile, on average, the browning rates of tissue-cultured 'Lakatan' TC plantlets, multiple bud clumps, and calli treated with the same concentration were 6.8, 9.6, and 10 (Table 1). For phytotoxicity tests using BEA, 'Grand Nain' TC plantlets, MBC, and callus scored 9, 8.8, and 7.6, while the same tissues of 'Lakatan' scored 7.0, 9.8, and 8.8, respectively (Table 2).
Table 1. Wilting rates of different tissues of banana 10 d after fusaric acid treatment.
FA (µm) 'Grand Nain' 'Lakatan' TC plantlet MBC Callus TC plantlet MBC Callus Control 0.8 ± 0.37b 0.6 ± 0.24c 0.2 ± 0.20d 0.4 ± 0.24c 0.4 ± 0.24e 7.8 ± 0.37ab 1 1.8 ± 0.49ab 1.2 ± 0.20c 1.2 ± 0.20cd 1.0 ± 0.55c 1.4 ± 0.24e 2.4 ± 0.68e 5 1.6 ± 0.60ab 3.8 ± 0.86bc 1.4 ± 0.24cd 1.2 ± 0.20c 1.8 ± 0.37de 3.0 ± 0.45e 10 1.4 ± 0.24b 4.0 ± 1.48bc 2.0 ± 0.00cd 0.4 ± 0.24c 3.8 ± 0.58cd 3.6 ± 0.51de 20 1.6 ± 0.40ab 4.4 ± 1.50abc 2.6 ± 0.24c 2.0 ± 0.32c 5.6 ± 0.24bc 3.8 ± 0.58de 30 1.4 ± 0.40b 4.0 ± 0.00bc 5.2 ± 1.24b 2.0 ± 0.00c 3.6 ± 0.75cd 5.4 ± 0.68cd 50 1.6 ± 0.24ab 7.3 ± 0.56ab 7.2 ± 0.49b 4.8 ± 0.49b 6.0 ± 0.63b 7.0 ± 0.00bc 100 3.4 ± 0.24a 7.8 ± 0.92a 9.6 ± 0.24a 6.8 ± 0.80a 10.0 ± 0.00a 9.6 ± 0.24a Values represent the mean ± SE of five replicates Means in a column with the same letter are not significantly different (p > 0.05). Table 2. Wilting rates of different tissues of banana 10 d after beauvericin treatment.
BEA
(µm)'Grand Nain' 'Lakatan' TC plantlet MBC Callus TC plantlet MBC Callus Control 1.0 ± 0.32d 0.6 ± 0.40c 0.6 ± 0.24d 0.4 ± 0.24d 0.0 ± 0.55e 0.4 ± 0.00d 1 4.0 ± 0.63c 1.2 ± 0.20c 3.2 ± 0.37c 1.6 ± 0.24c 7.2 ± 0.55d 1.4 ± 0.84c 5 4.6 ± 0.75c 5.4 ± 0.40b 4.6 ± 0.51b 2.8 ± 0.20b 8.4 ± 0.55c 4.6 ± 0.55b 10 6.8 ± 0.37b 6.0 ± 0.32b 6.6 ± 0.24a 6.2 ± 0.37a 9.2 ± 0.55b 6.6 ± 0.45a 20 9.0 ± 0.45a 8.8 ± 0.37a 7.6 ± 0.24a 7.0 ± 0.45a 9.8 ± 0.84a 8.8 ± 0.45a Values represent the mean ± SE of five replicates Means in a column with the same letter are not significantly different (p > 0.05). Treatment with FA and BEA significantly affected the browning rate compared to the control treatment (Tables 1 & 2). For 'Grand Nain,' the browning rate of TC plantlets and MBC significantly increased at 100 μm and 50 μm concentrations, respectively. At the same time, a significant increase in browning can be observed in callus cultures in as low as 20 μm FA (Fig. 3). FA also caused a significant browning of 'Lakatan' TC plantlets and MBC at 50 and 10 μm concentrations, respectively (Fig. 4). Additionally, 1 μm FA sufficiently caused significant browning in 'Lakatan' callus (Table 2, Fig. 4). BEA, at 1 μm concentration, already caused significant browning for both 'Grand Nain' and 'Lakatan' genotypes (Figs 3 & 4), except in 'Grand Nain' MBC where 5 μm BEA significantly affected the browning rate (Fig. 3). For both 'Grand Nain' and 'Lakatan' attached leaf tissues assay, browning around the inoculated sites were observed with 10 μm and 20 μm FA and BEA (Figs 3 & 4). But more severe symptoms were observed in the 'Lakatan' genotype than in 'Grand Nain' (Figs 3 & 4).
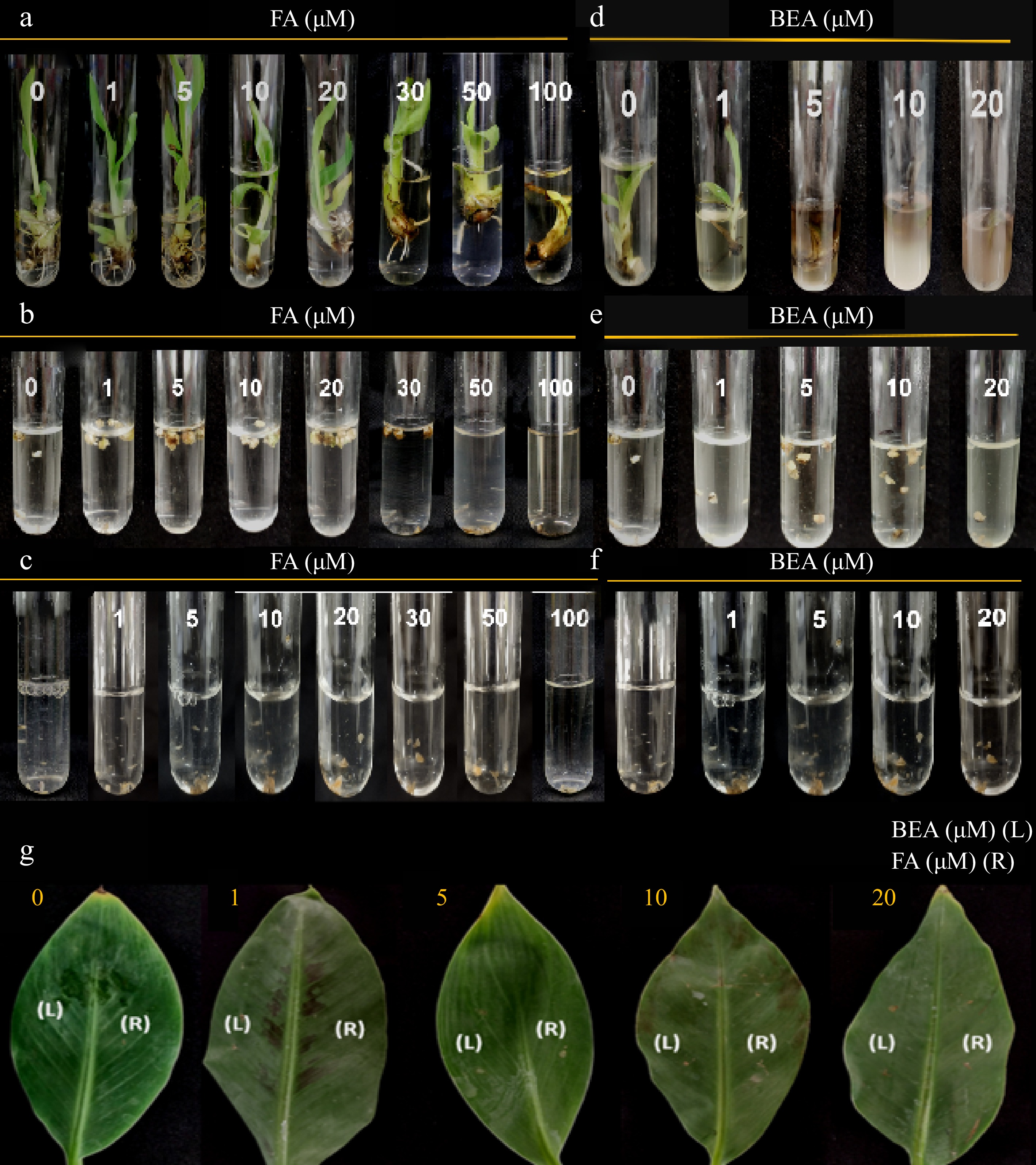

Figure 3.
Toxic effect of FA and BEA in different tissues of banana cv. 'Grand Nain'. Varying concentrations were tested in (a), (d) one-month old tissue-cultured plantlets, (b), (e) multiple buds clumps, (c), (f) callus and (g) attached leaf tissues of two-month old greenhouse plantlets were used. Symptoms were assessed at 10 d post-inoculation.
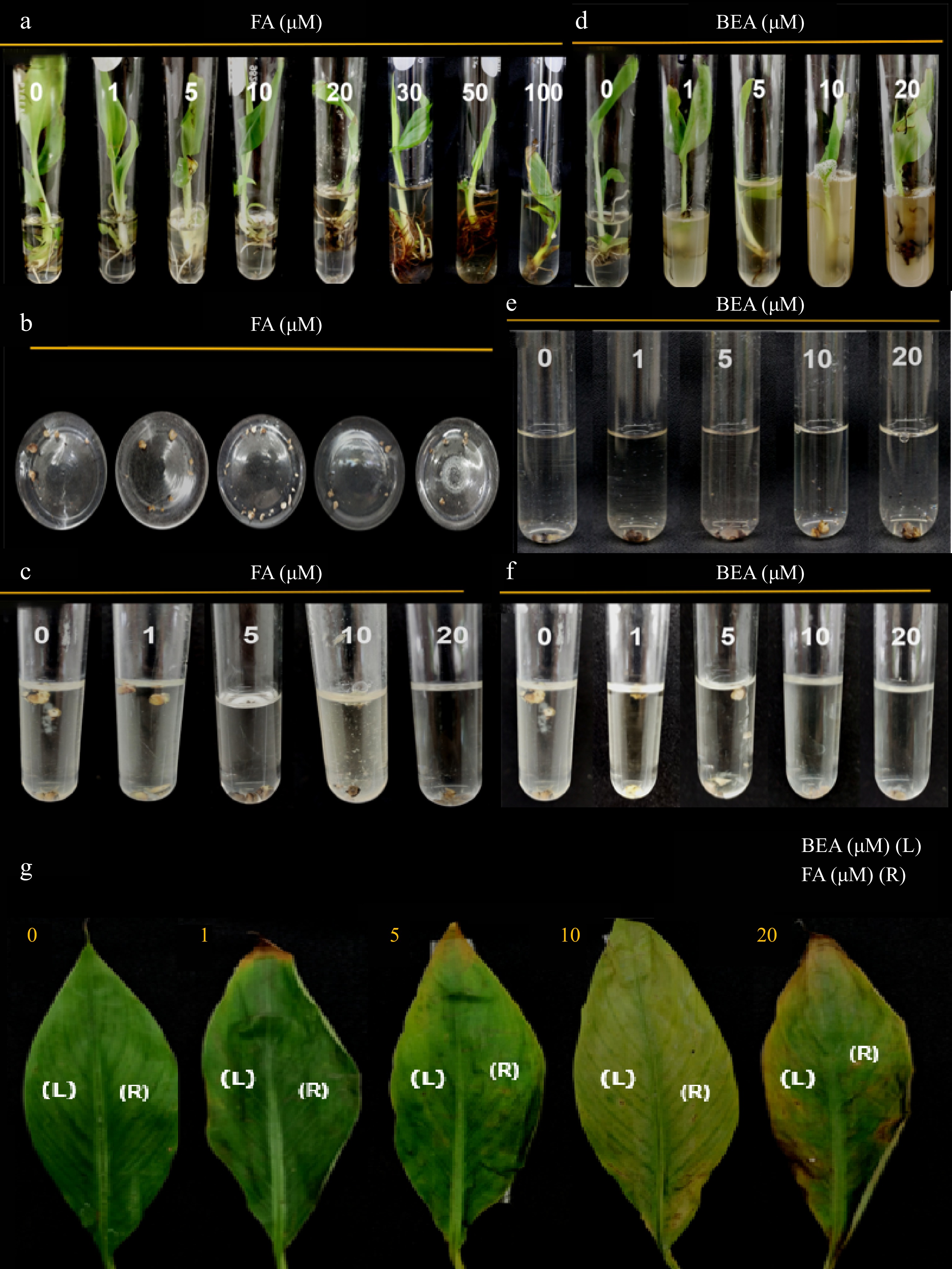
Figure 4.
Toxic effect of FA and BEA in different tissues of banana cv. 'Lakatan'. (a), (d) Varying concentrations were tested in one-month old tissue-cultured plantlets, (b), (e) multiple bud clumps, (c), (f) callus and (g) attached leaf tissues of two-month old greenhouse plantlets were used. Symptoms were assessed at 10 d post-inoculation.
-
This study demonstrated the phytotoxic effect of varying fusaric acid (FA) and beauvericin (BEA) concentrations in banana 'Grand Nain' and 'Lakatan'. The results were consistent with the previous findings of Li et al.[2], where 100 µm FA and 20 µm BEA sufficiently caused toxicity in tissue-cultured bananas. However, a lower concentration of BEA (1 µm) was found to cause wilting in this study. The arbitrary 10-point visual hedonic scales used in this study could be used to measure the phytotoxicity of Fusarium toxins in the test tissues of banana. A higher degree of vascular browning was recorded in the test tissues of 'Lakatan' treated with FA and BEA than in the 'Grand Nain' genotype. This suggests that banana sensitivity to phytotoxins differs from one genotype to another. In a study conducted by Molina et al., they recorded relatively higher susceptibility and disease incidence of 'Lakatan' in the field than in the 'Grand Nain' genotype[29]. Thus, the sensitivity rate of banana tissues to phytotoxins (FA and BEA) in-vitro may be correlated with its susceptibility to Fusarium wilt disease in the field. The degree of browning rates of test plants: tissue-cultured plantlets (TC plantlets), multiple bud clumps (MBC), callus, and attached leaf tissues to FA and BEA varied from one another. Higher susceptibility of callus and MBC than TC plantlets and attached leaf tissues were observed, which indicated higher sensitivity of the tissues to the phytotoxins. This confirms callus and MBC as suitable materials for screening resistant cell lines via somatic cell selection due to heightened toxin sensitivity, although the reason behind this is still unclear. A lower browning rate of attached leaf tissues than in in-vitro test plants demonstrated that toxin sensitivity was more pronounced in vitro than in planta. This may be attributed to several factors, including the environment and the type of tissues[2].
-
Fusaric acid (FA) and beauvericin (BEA) are toxic to banana 'Grand Nain' and 'Lakatan' through significant vascular browning. The arbitrary 10-point visual hedonic scales developed in this study for tissue-cultured plantlets (TC plantlets), multiple bud clumps (MBC), and callus tissues could be used to measure the phytotoxicity of Fusarium toxins. The higher degree of vascular browning observed in 'Lakatan' banana treated with FA and BEA toxins than in 'Grand Nain' suggested genotype-dependent toxin sensitivity and subsequent susceptibility of banana to Fusarium wilt disease, although further data are needed to support this. Data obtained here would aid in determining the effective concentration for toxin-based cell selection using callus and MBC cultures.
This study was supported by the Department of Agriculture- Bureau of Agricultural Research (DA-BAR), Philippines. We thank Fe dela Cueva, Edzel Evallo, Diana Rose Biglete, Monica Fronda, Yron Retuta, Rodel Pia, Pamela Quintos, May Eljera, Loida Pascual, Flora R. Cuevas, Rochelle Delgado, and Eugene Parañaque for the technical assistance and support. Special thanks to the Fruit Crops Section and the National Plant Genetic Resources Laboratory of the Institute of Plant Breeding, College of Agriculture and Food Science for providing the banana cultures and plantlets.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2023 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Cruz MA, Alcasid C, Millado CS, Balendres MA. 2023. Toxicity of fusaric acid and beauvericin in tissue-cultured banana 'Grand Nain' and 'Lakatan'. Technology in Horticulture 3:15 doi: 10.48130/TIH-2023-0015
Toxicity of fusaric acid and beauvericin in tissue-cultured banana 'Grand Nain' and 'Lakatan'
- Received: 23 April 2023
- Accepted: 18 May 2023
- Published online: 04 September 2023
Abstract: Fusarium oxysporum forma specialis cubense (Foc) produces toxins known to contribute to virulence and infection in the host. Fusaric acid (FA) and beauvericin (BEA) are major toxins contributing to Foc virulence in the host plant. Recent advancements allow the production of disease-resistant crops via cell selection, a process that involves resistance screening of somaclones using phytotoxin exposure. Determination of the appropriate concentration is an important step for the toxin-based selection of disease-resistant plants. In this study, the toxicity of FA (0, 5, 10, 20, 30, 50, and 100 μm) and BEA (0, 5, 10, and 20 μm) in different tissues of banana cv. ‘Grand Nain’ and 'Lakatan' were investigated. Overall results indicated a positive relationship between the toxin concentration and plant sensitivity, as indicated by the degree of vascular browning rate. Results demonstrated that lower concentrations of BEA are required for phytotoxicity than in FA. Furthermore, a higher degree of vascular browning was recorded in the test tissues of ‘Lakatan’ treated with FA and BEA than in the ‘Grand Nain’ genotype suggesting genotype-dependent sensitivity of banana to phytotoxins. To our knowledge, this study is the first to investigate the phytotoxicity of FA and BEA in callus cultures of banana ‘Lakatan.’
-
Key words:
- Phytotoxins /
- Musa sp /
- Banana /
- Fusarium oxysporum /
- Fusarium wilt












