-
Functional foods have taken their place in the global market, due to the relationship between health and diet. Probiotics and prebiotics are the most used additives to produce functional foods[1, 2]. Various studies have shown the positive effects of probiotics against different diseases, such as the alleviation of lactose intolerance[3], developed resistance to infectious diseases[4], and a decrease in serum cholesterol concentration[5].
Dairy products are thought to be good vehicles for delivering probiotics because they have intrinsic features that encourage probiotic growth and keep them viable[6]. Probiotic dairy products constitute the main group in the functional food sector because of their high consumption levels. Furthermore, innovative functional products are expected to be developed in the market as they offer consumers an alternative choice. Functional dairy desserts can be a suitable option for individuals of different ages to receive probiotics in their daily diet[7]. The viability of probiotics must be preserved during the process and storage conditions of foods. Some studies consider 6−9 log CFU/mL or CFU/g as the minimum starting level for probiotics to survive in adverse conditions and reach the gut in sufficient quantities to exert their beneficial effects[8].
Microencapsulation is a promising strategy for increasing probiotic viability in a range of foods. Proper coating materials and drying procedures should be chosen to enhance the survivability of bacteria during the process, storage, and exposure to the digestive system[9]. Microencapsulation provides higher protection for the probiotics, and a lower decrease in viability is observed in the simulated gastrointestinal system[10]. The entire delivery system for food applications must be made of food-grade materials. Polysaccharides and proteins are common wall materials for microencapsulation[11]. Whey proteins contain a variety of inherent functional features that enable them to function as effective encapsulants, including the capacity to stabilize emulsions and produce gel matrices. They can be used to encapsulate a variety of food substances, both hydrophilic and hydrophobic, as well as probiotic microbes[12]. The different characteristics of polysaccharides can be used to produce microcapsules and protect the microencapsulated probiotics from harsh environments, including electrostatic interaction, ion-induced gelation, enteric dissolution, and so on. Various traditional polysaccharides such as pectin, alginate, inulin, and chitosan are now widely used as wall materials; however, with the increasing demand for new applications for probiotic microcapsules, these limited types of polysaccharides can only satisfy[13]. Gum arabic as a polysaccharide is widely used in the food and pharmaceutical industries as a stabilizer, thickening agent, and emulsifier[14]. According to the American Food and Drug Administration, gum arabic is one of the safest dietary fibers[15].
This study involves investigating microencapsulation of L. plantarum DSM 1954 by using the water-in-oil emulsion technique with gum arabic-whey protein concentrate complex and an examination of viability levels of microencapsulated and non-microencapsulated L. plantarum in the cocoa pudding and the main quality parameters and sensorial properties during the storage for 21 d at 4 °C. Additionally, the survival of L. plantarum was evaluated under a simulated gastrointestinal tract.
-
The commercial strain of Lactiplantibacillus plantarum DSM 1954 was obtained from the German Collection of Microorganisms and Cell Cultures GmbH (Germany). Whey protein concentrate (WPC) (contains 80% protein) and gum arabic (E 414, food additive) were provided from Alfasol, Turkey. Sucrose, corn starch, pasteurized whole milk, and cocoa were obtained from a local market.
Methods
Bacterial strain and culture preparations
-
L. plantarum stock culture was inoculated 1% (approximately 11−12 log CFU/mL) into 5 mL MRS (de Man, Rogosa and Sharpe, Merck, Germany) broth and incubated at 37 °C for 24 h under anaerobic conditions. The culture was then subcultured into 100 mL of MRS broth and incubated under the same conditions for 18 h. Then, the cells were harvested by centrifugation[16].
Preparation of microcapsules
-
Microencapsulation of L. plantarum was performed using a water-in-oil emulsion method[16]. Gum arabic (9% w/v) and WPC (9% w/v) were dissolved separately in distilled water for 3 h[17]. Afterward, WPC was denatured in a water bath at 80 °C for 30 min. Then, the denatured WPC was cooled to room temperature. Gum arabic and WPC solutions were mixed. The primary water-in-oil emulsions were formed by emulsifying an inner aqueous phase made up of gum arabic and WPC complex containing L. plantarum into an oil phase containing 1% soy lecithin (Alfasol, Turkey) as an emulsifier. The primary emulsion was homogenized with an Ultra Turrax homogenizer for 5 min (Ultra Turrax T25, Janke and Kunkel, IKA Labortechnik, Germany). The emulsions were then homogenized again in 0.1 M CaCl2 solution (Applichem, Germany) for 2 min. After that, these slurries were shaken with an orbital shaker (IKA Labortechnik KS125 basic, Germany) to harden the microcapsules. The hardened microcapsules were separated from the solution by centrifugation. The microcapsules were then lyophilized by a freeze-dryer (Lablanco Freezone 18, Kansas, USA) at –55 °C under a 0.050 mBar vacuum for 48 h. The microcapsules obtained were preserved at 4 °C for further analysis.
Physicochemical properties of microencapsulated L. plantarum
-
Color measurement of microcapsules was evaluated using the Konica Minolta colorimeter (CR 410, Konica Minolta, Tokyo, Japan).
The water activity of microcapsules after freeze-drying was measured at the temperature of 25 °C by the Hygrolab C1 water activity counter (Rotronic, Bassersdorf, Switzerland).
The moisture content of microcapsules was determined after drying at 105 °C for 24 h[18]. The moisture content (%) was calculated by using the equation:
Moisture content (%) = [(Wwet – Wdry)/(Wwet)] × 100
Where Wwet is wet sample weight and Wdry is dry sample weight.
The pH value was determined by using a digital pH meter (Hanna Instruments, HI 2211 pH/ORP Meter, USA). Microencapsulated cells were dissolved in distilled water (1:10) for the pH measurement.
The bulk density was evaluated by adding 2 g of the microencapsulated cells into a 10 mL measuring cylinder and the cylinder was tapped by holding it on a vortex vibrator for 2 min. The tapped bulk density was calculated by using an equation[19].
Bulk density (kg/m) = m/v
Where m is the mass of the microencapsulated cells (kg), and v is the volume loaded in the cylinder (m3).
Enumeration of microencapsulated L. plantarum and microencapsulation efficiency
-
The microencapsulated L. plantarum viability was determined after the microencapsulation process on days 0, 7, 14, and 21. Microencapsulated samples (1 g) were dissolved in 9 mL sterile peptone water and shaken vigorously. The homogenized solution was serially diluted with peptone solution and plated by the pour plate technique using MRS agar. Viable cell counts were determined after 48 h incubation under anaerobic conditions using an anaerobic kit (Thermo Scientific, Oxoid AnaeroGen, UK) at 37 °C, and viable cells were expressed as log CFU/g. The encapsulation efficiency was calculated as follows:
Microencapsulation efficiency (%) = (N/N0) × 100%
where N0 is the cell count before microencapsulation (log CFU/g), and N is the cell count after the microencapsulation process (log CFU/g).
Morphology of microencapsulated cells
-
The morphology of microencapsulated cells was observed using a scanning electron microscope (SEM) (FEI QUANTA 250 FEG). Microencapsulated samples were coated with a conductive gold layer, before the observation. The microencapsulated cells were examined at an accelerating voltage of 5 kV and magnification of 10,000 and 20,000 times.
Preparation of the pudding samples
-
For cocoa pudding, 10% sucrose, 4.2% corn starch, 2% cocoa, and 0.2% gum arabic were added slowly to 83.6% pasteurized milk at 40 °C. The mixture was mixed with a magnetic stirrer. It was heated to 85 °C for 20 min and then stirred at the same temperature for 5 min[20]. When the temperature of the pudding dropped to 45 °C, the cocoa pudding was divided into three batches. Probiotic cultures were added and mixed homogeneously with a sterile mixer.
There are;
• CP, control cocoa pudding samples without L. plantarum
• MP, cocoa pudding samples including microencapsulated L. plantarum
• FP, cocoa pudding samples including free cell (non-microencapsulated) of L. plantarum
All cocoa pudding samples were packaged in separate plastic containers, refrigerated, and stored at 4 ± 1 °C for up to 21 d.
Determination of storage stability of pudding
Viability of L. plantarum in the cocoa pudding
-
The viability of L. plantarum in pudding samples (MP and FP) were evaluated during the 0, 7, 14, and 21 d. Ten g of the pudding was mixed with 90 mL of sterile peptone water and vigorously shaken. Samples were serially diluted with peptone water and plated using MRS agar to determine the count of L. plantarum, VRB agar (Violet Red Bile Agar, Merck, Germany) to detect the Escherichia coli contamination, and PDA (Potato Dextrose Agar, Merck, Germany) to observe mold and yeast growth. Following 48 h of anaerobic incubation for MRS agar and VRBA, colonies on plates were counted. PDA was incubated in aerobic conditions for 5 d.
pH determination
-
The pH of all pudding samples (CP, MP, and FP) was measured on the 0, 7, 14, and 21 d. A pH meter (Hanna Instruments HI 2211, USA) was used for pH measurement. The pH value of the samples was determined by dipping the pH probe in the homogenized pudding sample.
Simulated in vitro gastrointestinal digestion
-
The experiment was performed on cocoa pudding samples supplemented with microencapsulated and non-microencapsulated L. plantarum, following the INFOGEST protocol with some modifications[21]. Three digestion fluids were prepared: salivary, gastric, and intestinal. Simulated salivary fluid (SSF) was prepared with 15.1 mM KCl, 13.6 mM NaHCO3, 3.7 mM KH2PO4, 0.15 mM MgCl2(H2O)6, 1.5 mM CaCl2, 0.06 mM (NH4)2CO3. The SSF was adjusted to pH 7.0 at 37 °C. Pudding samples were mixed and incubated at 37 °C for 2 min. Microencapsulated and non-microencapsulated cells within the SSF were also used as control. Simulated gastric fluid (SGF) was prepared with 6.9 mM KCl, 25 mM NaHCO3, 0.9 mM KH2PO4, 47.2 mM NaCl, 0.1 mM MgCl2(H2O)6, 0.15 mM CaCl2, 0.5 mM (NH4)2CO3 and 1,600 U/mL pepsin (Sigma-Aldrich, P7000) and adjusted to pH 3.0. After SSF, an oral bolus was added to the SGF. Samples were mixed with an overhead shaker (Heidolph Reax 2, Schwabach, Germany) and incubated at 37 °C for 120 min. Simulated intestinal fluid (SIF) was prepared with 6.8 mM KCl, 85 mM NaHCO3, 0.8 mM KH2PO4, 38.4 mM NaCl, 0.6 mM CaCl2, 0.33 mM MgCl2(H2O)6, 0.02 mM bile salts (Sigma-Aldrich, B8631) and 160 U/mL pancreatin (Sigma-Aldrich, P1750). The gastric chime was mixed with SIF and adjusted pH to 7.0. Samples were mixed with an overhead shaker and incubated at 37 °C for 120 min.
Sensory analysis
-
Sensory analysis was performed after the production of the cocoa pudding and the panel consisted of 25 untrained individuals. An acceptance test of qualifications (appearance, smell, texture, taste, consistency in mouth, and overall acceptance) using a 5-point hedonic scale (1 = very bad and 5 = excellent) was carried out[22]. Panelists evaluated three different cocoa pudding samples at one time. Each of the pudding samples was encoded with a 3-digit arbitrary number and presented properly to the panelists.
Statistical analysis
-
All experiments were performed in parallel. Data analysis was carried out using Minitab 19.0 software (Minitab Inc., State College, PA, USA). The results were expressed as mean ± standard deviations. Variance analysis and Tukey’s tests were used to demonstrate the differences between the pudding samples.
-
L. plantarum was microencapsulated with gum arabic and WPC complex by using the water-in-oil emulsion method. Before microencapsulation, the initial cell load was 12 log CFU/mL. During the 21-d storage, 86.66% microencapsulation efficiency and 10.33 log CFU/g of cell viability were observed (Fig. 1). According to the results obtained, it was observed that the viability of the microencapsulated L. plantarum increased on the 7th d during storage and the difference in viability change from the first day to the last day was observed as 0.27 log CFU/g.
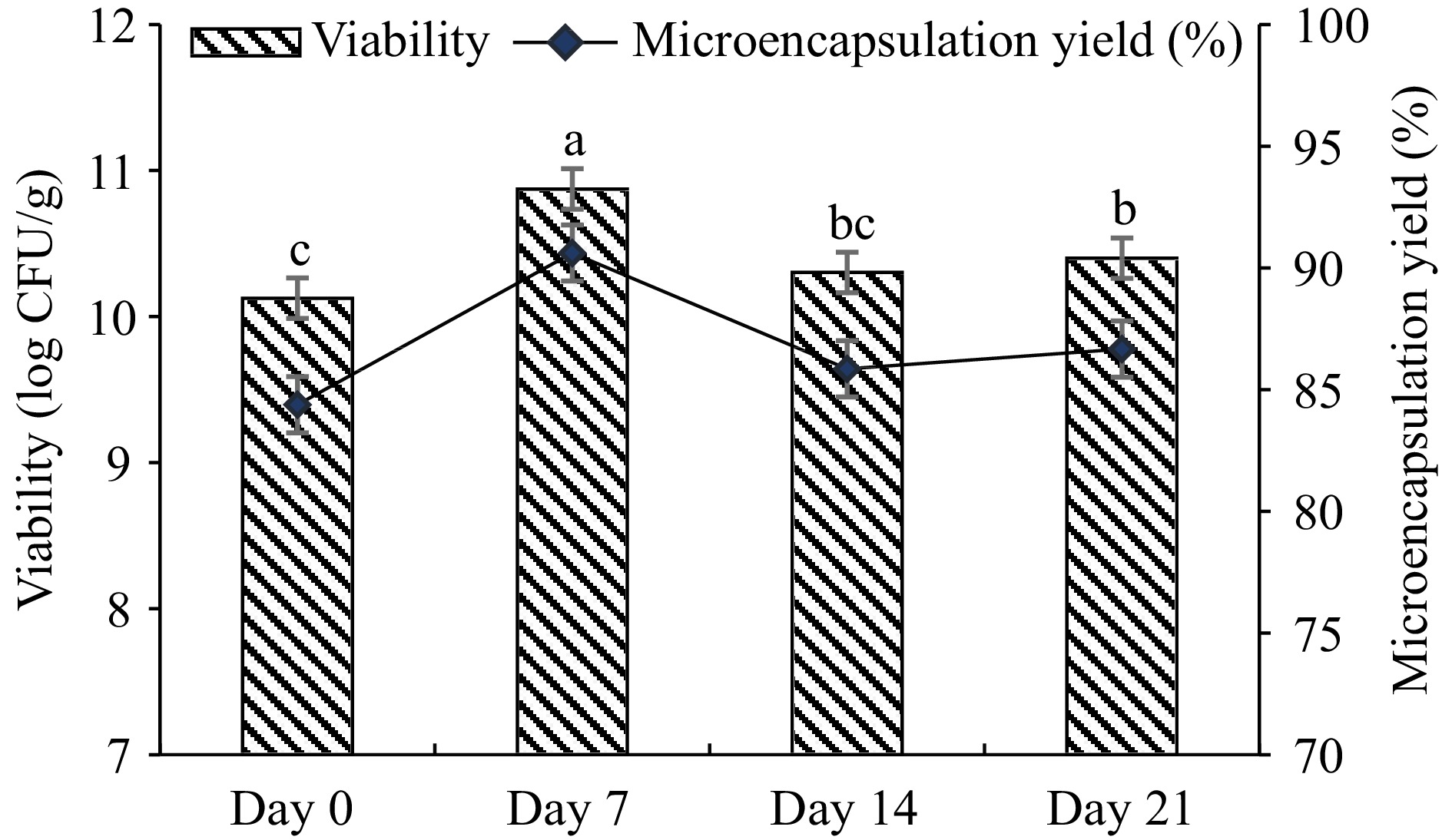
Figure 1.
Viability of microencapsulated L. plantarum (log CFU/g) and microencapsulation efficiency (%). Microcapsules were produced by a water-oil-emulsion technique using gum arabic-WPC complex, stored at 4 °C for 21 d. Results are shown as the means and standard deviations. Data are the average of duplicate samples, obtained in two independent assays.
%, indicating that still there are free cells present in the medium, e.g. 14%. It can be thought that low amount of free bacteria may utilize the sugar in the pudding and grow and increase in cell number slightly during storage.
Physicochemical properties of microencapsulated L. plantarum
-
The physicochemical properties of microencapsulated cells were evaluated. To determine the color of microencapsulated cells, the CIELab parameters; L* (brightness), a* (redness), and b* (yellowness) have been analyzed, and the values were found as 71.41 ± 0.21, 0.936 ± 0.08, and 32.84 ± 0.15, respectively. The pH value of diluted (1:10) samples was found as 5.09 ± 0.008. The water activity of the microencapsulated cells after the microencapsulation process was 0.069 at 26.25 °C. The moisture content of microencapsulated cells was 0.048%. The bulk density of microencapsulated cells was found to be 400 kg/m.
Morphology of microencapsulated cells
-
The morphology of microencapsulated cells was observed using an SEM. Figure 2 shows the surface views of the microcapsules. All microcapsules were shown to be spherical and regular in shape. Figure 2a shows the distribution of the microcapsules in the pudding, while Fig. 2b shows the spherical structure of the microcapsules and the homogeneity in their size. This observation suggests that the gum arabic – WPC complex has been successfully deposited onto the external surfaces of the L. plantarum cells. The diameter of the microencapsulated cells was recorded by SEM and the data showed that the microcapsules prepared with gum arabic-WPC complex were 1.044 ± 0.211 µm.
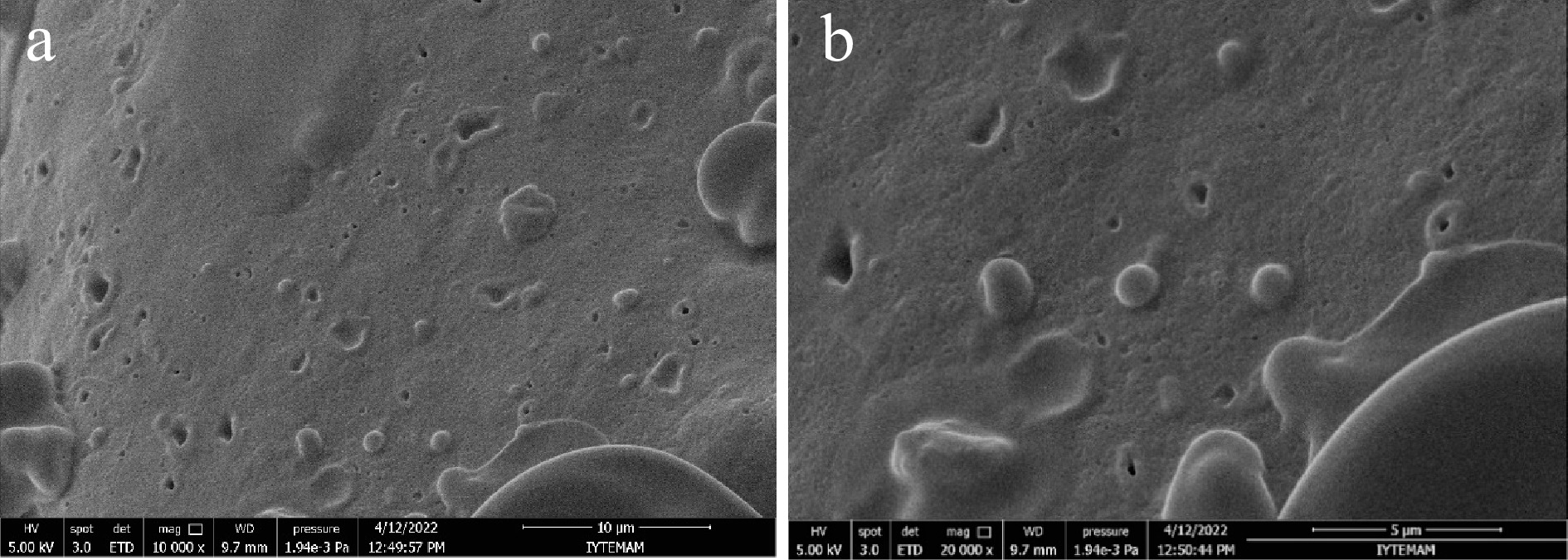

Figure 2.
Scanning electron micrographs of microencapsulated L. plantarum with gum arabic-WPC complex. (a) 10000× magnification, scale bar = 10 μm, (b) 20000× magnification, scale bar = 5 μm.
Viability of L. plantarum in the cocoa pudding
-
The L. plantarum was microencapsulated with gum arabic - WPC complex and included in the pudding. Microencapsulated and non-microencapsulated L. plantarum viability in pudding samples was evaluated during 21-d storage. Microencapsulated cell numbers increased on day 7 of storage at 4 °C; non-microencapsulated cells showed slow log reduction. The results obtained showed that the number of non-microencapsulated cells did not differ significantly during storage, but the viability of microencapsulated cells increased.
pH changes were determined to show the storage stability of pudding samples for 21 d. The pH values of all pudding samples are given in Fig. 3. At the beginning of storage, the pH values of all pudding samples were between 6.91 and 7.12, while at the end of the storage pH value of CP and MP decreased to 5.83 and 5.3, respectively. The pH value of the FP sample dropped significantly to 4.59 during storage.
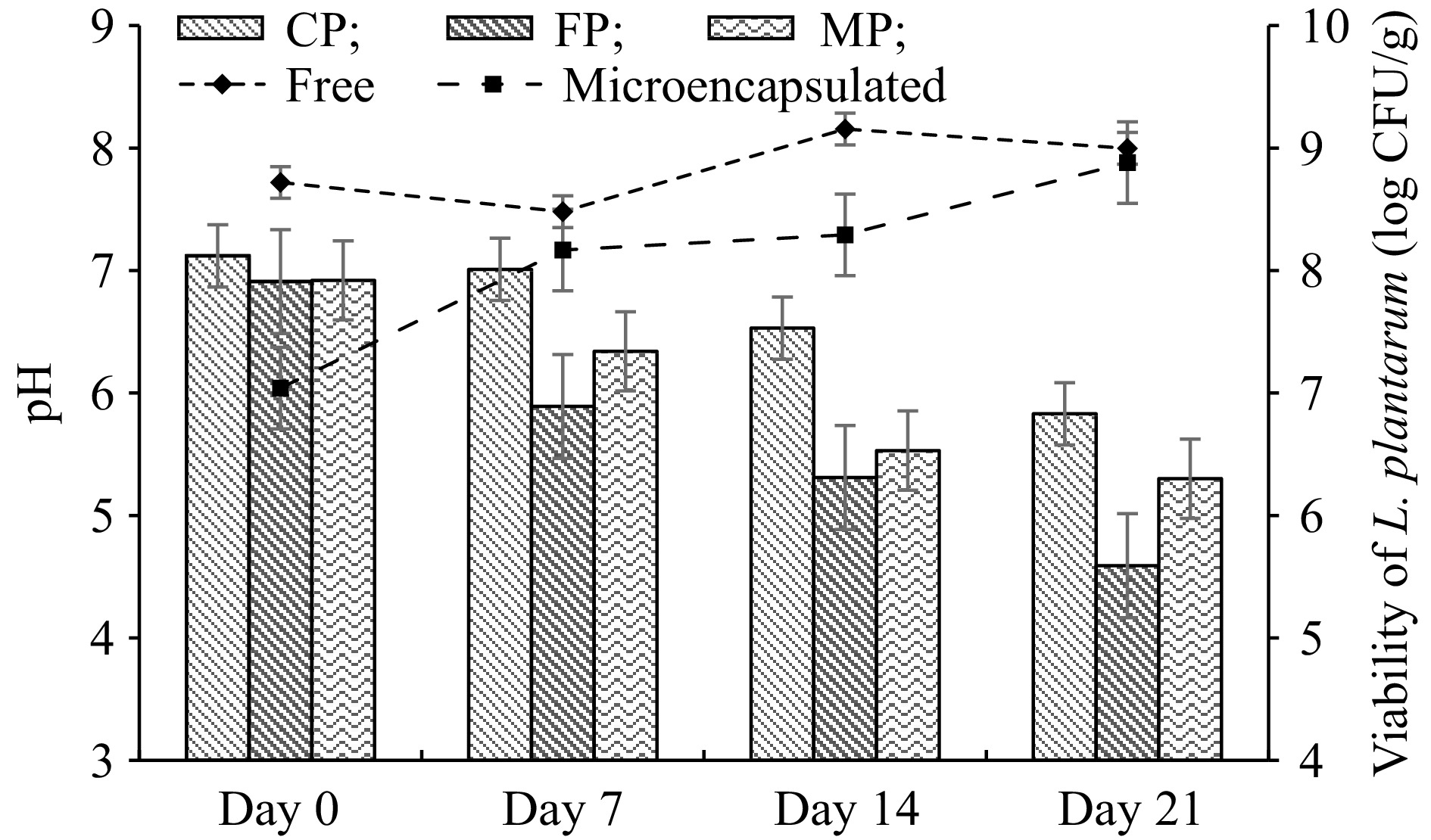
Figure 3.
Changes in pH and viable cell counts of free and microencapsulated L. plantarum in cocoa pudding during the storage at 4 °C. Results are shown as the means and standard deviations. Data are the average of duplicate samples, obtained in two independent assays. MP: Cocoa pudding samples including microencapsulated L. plantarum. FP: Cocoa pudding samples including free cells (non-microencapsulated) of L. plantarum. CP: Control cocoa pudding samples without L. plantarum.
Total coliform bacteria, yeast, and mold were evaluated during the storage time of cocoa puddings to prove microbiological safety. Based on the findings, no total coliform bacteria, yeasts, and molds were detected in all pudding samples.
Simulated gastrointestinal digestion
-
Microencapsulation processes are used to protect probiotics from harsh environmental conditions such as low pH, bile salts, and enzymes in the gastrointestinal tract. This study determined the influence of gastrointestinal conditions on the survivability of free (non-microencapsulated) and microencapsulated L. plantarum. Additionally, L. plantarum contained pudding samples that were evaluated to show the survival of bacterial cells (Fig. 4). After exposure to SGF for 2 h, there was a 2.34 and 1.52 log CFU/mL decrease in viability of the microencapsulated and free cells, respectively. However, a low viability reduction was observed in MP and FP samples, 0.05 and 0.14 log CFU/mL, respectively. Then digested SGF was passed to SIF for 2 h, in which there was a 1.0 and 1.34 log CFU/mL decrease in the viability of microencapsulated and free cells. A 2.0 and 1.18 log CFU/mL viability reduction was observed in MP and FP samples.
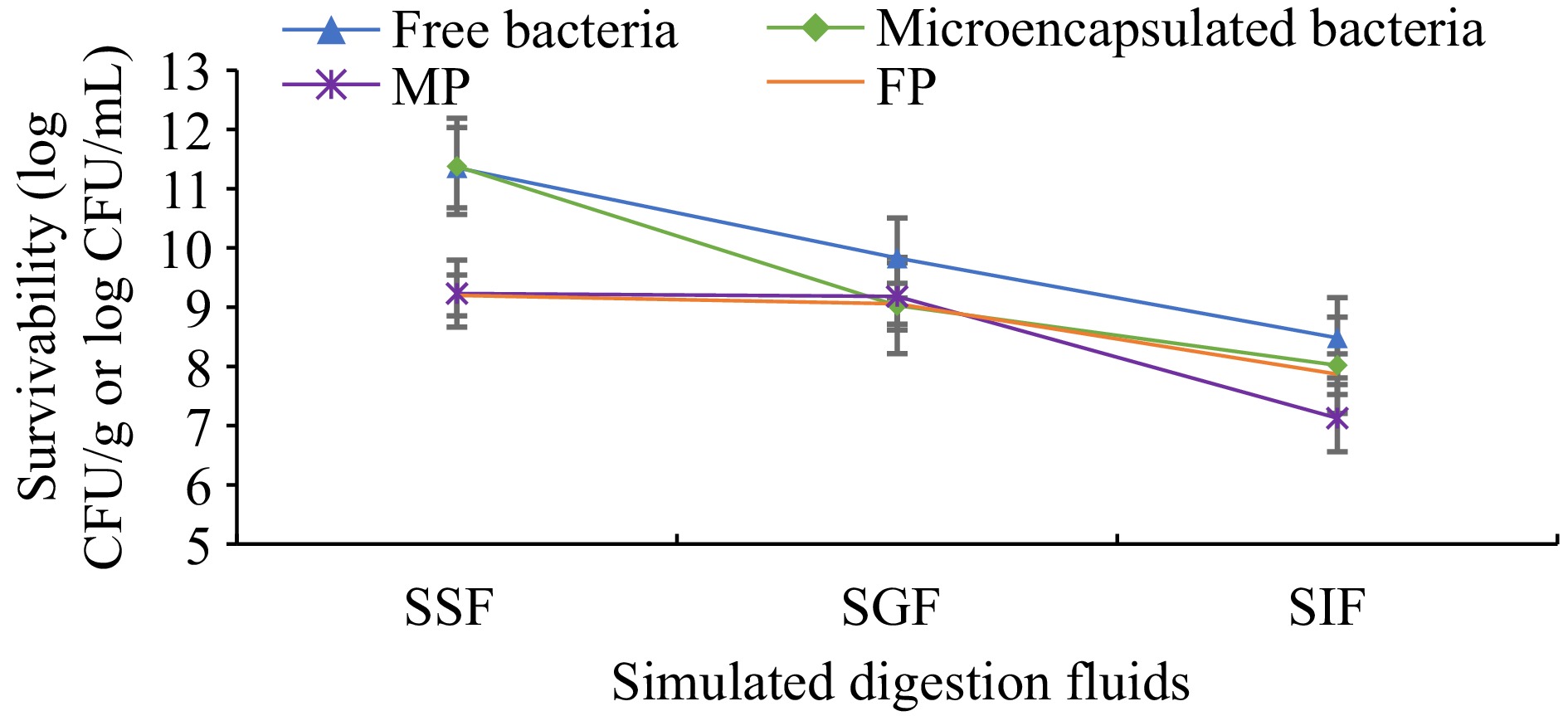
Figure 4.
Survivability of free and microencapsulated L. plantarum. Results are shown as the means and standard deviations. Data are the average of duplicate samples, obtained in two independent assays. SSF: Simulated saliva fluids. SGF: Simulated gastric fluids. SIF: Simulated intestinal fluids. MP: Cocoa pudding samples including microencapsulated L. plantarum. FP: Cocoa pudding samples including free cells (non-microencapsulated) of L. plantarum.
During the simulated digestion test, a 3-log reduction was observed in both free and microencapsulated L. plantarum cells. In pudding samples (MP and FP) only 2-log reduction was observed, this can be expected due to the pudding materials may show a protective barrier for microorganisms. After the 3-week storage period, the survivability of L. plantarum was between 77% and 85% in pudding samples, while the survivability of L. plantarum (not incorporated in the pudding) was found between 70% and 74%.
Sensory analysis
-
L. plantarum incorporated pudding samples and controls were evaluated shortly after the production, by 20 untrained panelists about the appearance, texture, taste, smell, consistency in mouth, and general acceptance. The sensory analysis results are reported in Table 1. The sensorial properties of pudding samples received scores between 3.84 and 4.40 on the 5-point hedonic scale. The interaction between cocoa pudding and microencapsulated and free L. plantarum did not demonstrate any significant effect (p > 0.05). This result demonstrated that a concentration of L. plantarum of approximately 7–8 log CFU/g did not influence the pudding characteristics.
Table 1. Sensory evaluation of pudding samples.
Pudding samples Appearance Texture Taste Smell Consistency in mouth General acceptance CP 4.400 ± 0.957A 4.080 ± 0.862A 4.160 ± 0.850A 4.120 ± 0.781A 4.040 ± 1.060A 4.200 ± 0.866A FP 4.280 ± 0.542A 4.200 ± 0.866A 4.200 ± 0.707A 4.040 ± 0.676A 4.160 ± 0.898A 4.160 ± 0.624A MP 4.120 ± 0.781A 4.320 ± 0.945A 4.280 ± 0.843A 3.840 ± 0.800A 4.240 ± 0.970A 4.160 ± 0.800A Results are shown as means ± standard deviation. Different capital letters on the same column show a significant difference by Tukey's test (p < 0.05), n = 25. CP: Control cocoa pudding samples without L. plantarum. MP: Cocoa pudding samples including microencapsulated L. plantarum. FP: Cocoa pudding samples including free cells (non-microencapsulated) of L. plantarum. -
L. plantarum was used in this study to produce functional pudding because of its probiotic properties. A recent article highlighted the probiotic properties of L. plantarum, such as resistance to gastrointestinal conditions, adherence to the bowel mucosa, the capacity to promote intestinal integrity, inhibit the growth of pathogens by altering the intestinal flora, modulate immune function, and the ability to treat or cure diseases[23].
The increase in viability of microencapsulated L. plantarum at the refrigerator temperature during storage was observed (Fig. 1), which can be explained by the absence of toxic waste accumulation due to decreased cell metabolism[24]. The subsequent decrease in viability can be attributed to the oxidation of lipids and protein denaturation during storage. During storage, the protective mechanism of WPC and gum arabic may form a semi-permeable wall around living cells, reducing the water content from the intermediate medium. In a previous study, microencapsulation with xylan-WPC complex of L. plantarum DSM 1954 showed no significant change in the viability, which was kept at 4 °C for 4 weeks[25]. A previous study showed that the viability of Streptococcus thermophilus CCM4757 decreased from 10.26 log CFU/g to 9.94 log during 6 months of storage when gum arabic and WPC complex was used as a coating material in the microencapsulation process[17].
Recommended water activity is below 0.6 to stabilize microbial growth[26]. In this study, obtained water activity value is lower than recommended to stabilize microencapsulated cells. In addition, a lower water activity value provides longer shelf life because less water is available to catalyze biochemical reactions[27]. Likewise, the lower water activity; low moisture content limited microbial growth and spoilage. In a study, water activity and moisture content of encapsulated L. plantarum were found as 0.34 and 4.5%, respectively[28]. Also, spray-dried L. plantarum microcapsules’ water activity was 0.196, and the moisture level was 3.73%[29]. Bulk density is important to process, packaging, and store the powders. In addition, morphology, particle size distribution, and moisture content can affect the bulk density of powders[19]. The bulk density of spray-dried L. plantarum (MTCC 5422) with fructooligosaccharide as coating material microcapsules ranged from 405.11 to 564.72 kg/m³[19]. The high bulk density of powders is more advantageous because large amounts of powders can be stored in smaller containers than powders with lower bulk densities. Higher bulk density can suggest a lower amount of stored air among particles[30]. Coating materials with high bulk density can exhibit a more protective impact on probiotic cells during storage because of a lower air volume[31].
There is no ideal particle size, the size depends on the specific application intended and can range from a few micrometers to several millimeters in foods[32]. The particle size of microcapsules of L. plantarum ATCC 8014 obtained by spray-drying with whey was found as 7.0 ± 1.0 μm[29]. The larger particle sizes of encapsulated L. plantarum were obtained around 66.07 ± 3.24 μm and 105.66 ± 3.24 μm[28]. The color parameters of microencapsulated cells showed consistency with a previous study, which carried out a spray-drying process with whey retentate to obtain L. plantarum microcapsules[29].
During the storage period of pudding samples, fluctuations in the viability of microencapsulated L. plantarum were observed (Fig. 3). The encapsulation efficiency of L. plantarum was 86.66%, indicating that still there are free cells present in the medium, e.g., 14%. It can be thought that a low amount of free bacteria may utilize the sugar in the pudding and grow and increase in cell number slightly while the pH value of samples is reduced during storage. This may be desirable up to some extent since the slight decrease in pH can prevent the product from spoilage microorganisms without influencing the sensory attributes of the product.
Likewise, the cell viability fluctuations were also seen in a study, in which kefir was produced using free and encapsulated L. plantarum[33]. According to a novel study, the viable cell number of L. plantarum in dairy dessert remained constant for 15 d of storage at 4 °C[7]. According to studies probiotic cell viability must be maintained during the processing and storage of food products, and foods containing probiotic bacteria must have at least 106 CFU/g or CFU/mL at the time of consumption to exhibit beneficial effects[31]. As can be seen in Fig. 3, viable L. plantarum count in FP and MP has been higher than the recommended daily intake. Besides, L. acidophilus viability was detected as higher than 6.5 log CFU/g in rice pudding samples during the shelf life[34].
The pH value of all pudding samples decreased during storage, but a higher decrease in the pH of FP was found (Fig. 3), the reason is that bacteria in free form break down carbohydrates and produce acid. According to the findings, microencapsulation technology provides the most desirable combination of quality and shelf-life of products. Furthermore, L. plantarum fortified dairy desserts showed a low pH value (5.29) at the end of the 15th-d of shelf life[7]. In a novel study, Konjac root powder added chocolate milk was enriched with 2% of free or microencapsulated lactic acid bacteria. The findings indicated a gradual decrease in all samples' pH values during refrigerated storage[35].
Probiotics have the ability to maintain viability throughout the gastrointestinal tract. Although there were decreases in viability due to the high acidity of gastric fluid, more than 8 log viability was observed in both microencapsulated and non-microencapsulated L. plantarum at the end of the simulated digestion (Fig. 4). In a novel study, microencapsulated L. plantarum 21,805 showed only a 0.51 log CFU/mL decrease in viability after exposure to SGF[33]. In a study, the encapsulation process combined with double emulsification and complex coacervation was carried out using gum arabic and gelation. After in vitro simulated gastrointestinal conditions, encapsulated L. plantarum viability was found as 80.4%[28].
-
This study demonstrated that L. plantarum was effectively microencapsulated with gum arabic-WPC complex. Microencapsulated L. plantarum, after storage at 4 °C for 21 d, showed higher cell counts than the minimum number of probiotics required for functional food products. Physicochemical properties, survival after exposure to the simulated gastrointestinal tract, and maintenance of viability in pudding indicated that this microencapsulation technique is suitable for preserving probiotics. The viability of microencapsulated and non-microencapsulated L. plantarum was higher than 7 log CFU/g in the cocoa pudding after 21 d of storage at 4 °C. Dairy desserts such as pudding appear to be effective food matrices for the transportation of probiotics without affecting sensory properties. Future studies should be conducted to evaluate the health-beneficial effects of probiotic pudding.
-
We are grateful to the Izmir Institute of Technology Integrated Research Centers for technical support.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2023 by the author(s). Published by Maximum Academic Press on behalf of Nanjing Agricultural University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Silkin B, Onen B, Elvan M, Harsa HS. 2023. Cocoa pudding fortified with microencapsulated Lactiplantibacillus plantarum DSM 1954. Food Materials Research 3:22 doi: 10.48130/FMR-2023-0022
Cocoa pudding fortified with microencapsulated Lactiplantibacillus plantarum DSM 1954
- Received: 31 March 2023
- Accepted: 09 July 2023
- Published online: 08 October 2023
Abstract: Probiotic pudding can be served as a functional food with high probiotic viability during production and storage. The aim of this study was to investigate the microencapsulation of Lactiplantibacillus plantarum DSM 1954 with a gum arabic-whey protein concentrate complex using a water-in-oil emulsion technique and to evaluate the stability of microencapsulated and non-microencapsulated L. plantarum in the cocoa pudding and to determine the main quality parameters and sensory characteristics of pudding during storage at 4 °C for 21 d. The efficacy of microencapsulation on the viability over 21 d was determined and an encapsulation efficiency of 86.66% was achieved. Whole milk, cocoa, corn starch, and gum arabic, as well as microencapsulated and non-microencapsulated L. plantarum were used to produce probiotic-fortified cocoa pudding. L. plantarum was added to pudding for 21 d to test their viability and stability. pH values and sensory analysis of pudding were conducted. Microencapsulated and non-microencapsulated L. plantarum cell counts were approximately 9 log CFU/g in pudding samples at the end of 21 d. With storage time, the pH of pudding containing non-microencapsulated bacteria decreased more than that of pudding containing microencapsulated bacteria. The addition of bacteria to the pudding did not have a significant effect on the taste, odor, and texture. Since both microencapsulated and non-microencapsulated bacteria maintain significant viability in pudding during storage, pudding can be considered a potential carrier of probiotics.
-
Key words:
- Functional foods /
- Gum arabic /
- Probiotic /
- Microencapsulation












