-
Soil salinity is a major restraint to agricultural production worldwide, and has been recognized as a major challenge to food security by the Food and Agricultural Organization (FAO, 2022). According to the reports, salt-affected soils are estimated to be around 1 billion ha, globally[1]. According to CSSRI, Karnal Haryana, salt-affected soils are estimated to increase by ~16 mha in India by the year 2050, due to anthropological changes. Soil salinization has made a significant part of land area unproductive or less productive[2]. This land degradation in salt-affected areas in India are threatening the development and economic growth of the country. In arid and semi-arid regions of India, desertification is reaching irreversible levels due to environmental degradation. Continued usage of improper farming techniques and activities has intensified soil salinization[3−5]. However, there is a dire need to adapt promising approaches in agriculture, which have the potential to meet global food demand while reducing the environmental footprint of agricultural systems[6,7]. The estimates show that 50% of arable land around the world will become salt-affected by the year 2050, therefore escalating food insecurity[8,9]. According to the Central Ground Water Board (CGWB), 84.87% of groundwater resources are classified as 'saline' in Haryana, largely due to overexploitation of groundwater resources and improper agricultural practices, which has led to the depletion of groundwater reserves and increased salinity. Soil salinity has a significant negative impact on crops, reducing their growth, yield and quality. This is due to the interference of salt with the nutrient and water uptake ability of plants, disrupting the morphological, physiological, biochemical, molecular, and metabolic processes resulting in a lowering of osmotic potential and an increase in ion toxicity[10,11]. An increase in the toxicity of ions causes the production of reactive oxygen species (ROS), contributing to a decrease in protein content and accumulation of proline and other osmolytes[12]. The global population is also expected to grow to 9.8 billion by 2050, which will require at least a 50% rise in agricultural production from 2012 levels.
Sorghum bicolor (L.) Moench, is a C4 crop of the family Poaceae and is ranked among the top five cereal crops globally. It is a multipurpose crop that is mainly grown for food, fodder, fuel, and bioethanol production[13]. It is well known for its adaptability to moderately salt-stressed conditions and is therefore widely grown in arid and semi-arid areas. It is known for its high nutritional content, gluten-free nature, and low environmental impact, making it an ideal choice for promoting sustainable food systems. According to the United States Department of Agriculture (USDA) report of 2022, the world's total sorghum production is 58.6 million tonnes, with India contributing 7.51% of the world's production. In India, Maharashtra is the largest producer of sorghum, contributing 47%, followed by Karnataka (22%) and Rajasthan (8%). In Haryana, sorghum is mainly grown as a fodder crop, with an area of 40.3 thousand hectares and a production of 21.3 thousand tonnes, with an average yield of 528 kg per hectare, according to the Department of Agriculture (DOA) of Haryana. The year 2023 has also been declared as the International Year of Millets to raise awareness about potential nutritional benefits, resilience, and the role of millets in sustainable agriculture. It is anticipated to emerge as a vital crop in the current climate change scenario due to its significant tolerance to high temperatures, water deficit, and saline conditions, which makes it an indispensable feed reserve[14]. Sorghum is a rich source of various phytochemicals including tannins, phenolic acids, anthocyanins, phytosterols, and policosanols and its fractions possess high antioxidant activity in vitro relative to other cereals. It is gluten-free and can be a good alternative for those with celiac disease or gluten sensitivity. It is also low on the glycemic index, making it a good choice for people with diabetes.
Silicon (Si) is a beneficial element which acts as a mitigator of abiotic and biotic stresses and increases growth and production in higher plants[15,16]. It is present in the earth's crust in the form of SiO2 in high concentration but it can't be utilized by plants in this form. Plants uptake Si in the form of ortho-silicic acid (OSA) or mono-silicic acid (H4SiO4). It is the only biologically active form of silicon and is easily soluble in water. Silixol is a stable formulation of OSA which contains 0.8% ortho-silicic acid, and has been generally used in foliar spray as a Si fertilizer in different concentrations which vary from crop to crop to enhance plant growth. Si helps in the alleviation of salt stress by reducing uptake of Na+[17] and Cl− ions[18] which contributes to the maintenance of optimum K+/Na+ ratio under saline conditions. The effect of Si on the transcriptional regulation of genes involved in water transport and stress-related pathways, including the jasmonic acid pathway, ABA-dependent or independent regulatory pathway, and phenyl propanoid pathway, has been proposed and the role of Si in stress mitigation in rice, wheat, maize, tomato, and soybean have been established by various researchers[19]. Silicon also alleviates salt-induced osmotic stress by upregulating the aquaporin's activity, thereby increasing the root hydraulic conductance[20]. Si is associated with higher tolerance due to involvement in the protection of various essential mechanisms such as energy metabolism, photosynthesis, transcription/translation, and hormonal signaling under salinity using proteomics studies[21]. Si supplementation via both foliar and root application effectively attenuated salinity in sorghum and sunflower plants by contributing to an antioxidative defense mechanism leading to Na+ detoxification[22]. Si application also enhanced nutrient uptake and nutrient use efficiency (NUE) contributing to higher dry weight accumulation[23]. The foliar application of manganese combined with silicon resulted in increased quantum efficiency of PSII, micronutrient uptake, relative chlorophyll index and subsequently enhanced dry mass accumulation in corn and sorghum[24]. The above findings indicate that silicon plays a crucial role in enhancing crop resilience and its nutritional quality under salinity stress. Therefore, the present investigation with the aim to mitigate the negative impact of salt stress on the growth, physiological, and biochemical parameters of sorghum by exogenous application of ortho-silicic acid was undertaken.
-
Seeds of two sorghum genotypes (CSV33MF and SSG 59-3) were collected from the Forage Section, Department of Genetics and Plant Breeding, Chaudhary Charan Singh Haryana Agricultural University, Hisar (Haryana) and were sown in pots under screen house conditions in screen house of the Department of Botany and Plant Physiology on July 17, 2021. The seeds were surface sterilized in 1% sodium hypochlorite (NaOCl) solution for 5 min and were sown in plastic pots containing 10 kg of dune sand. Before sowing pots were saturated with desired salt levels i.e., control (0), 4, 6, and 8 dS·m−1. The nutrient solution was given at regular intervals according to the method of Arnon[25]. Ortho-silicic acid (1.5 and 2.5 mg·L−1) was applied exogenously with the help of a manual sprayer 30 d after sowing (DAS). Three plants per pot were maintained to study several parameters.
Growth/morphological attributes
-
Fresh weight (FW) of leaf, root and stem, fresh weight (FW) of leaf, root and stem height (HT), dry weight (DW) and leaf area (cm2·plant−1) at 40 Days After Sowing (30 DAS and 10 Days after OSA application).
Physiological attributes
-
Photosynthesis rate, transpiration rate, and stomatal conductance were calculated using a third leaf from the top by using a plant Infrared Gas Analyzer (IRGA, LCi-SD, ADC Bioscience, USA)[26].
Relative water content (RWC) was calculated according to Barrs & Weatherley[27] using the formula:
$ \mathrm{R}\mathrm{W}\mathrm{C}\;\left({\text{%}}\right)=\dfrac{\mathrm{F}\mathrm{r}\mathrm{e}\mathrm{s}\mathrm{h}\;\mathrm{w}\mathrm{e}\mathrm{i}\mathrm{g}\mathrm{h}\mathrm{t}-\mathrm{D}\mathrm{r}\mathrm{y}\;\mathrm{w}\mathrm{e}\mathrm{i}\mathrm{g}\mathrm{h}\mathrm{t}}{\mathrm{T}\mathrm{u}\mathrm{r}\mathrm{g}\mathrm{i}\mathrm{d}\;\mathrm{w}\mathrm{e}\mathrm{i}\mathrm{g}\mathrm{h}\mathrm{t}-\mathrm{D}\mathrm{r}\mathrm{y}\;\mathrm{w}\mathrm{e}\mathrm{i}\mathrm{g}\mathrm{h}\mathrm{t}}\times 100 $ To assess the water potential of leaves, a pressure chamber (Model 3005, Soil Moisture Corporation, Santa Barbara, CA, USA) was used[28]. The third leaf from the top was separated from the plant with the help of a sharp edge knife and sealed in the pressure chamber one by one with the cut end protruding outside, and pressure was developed until the sap just appeared at the end.
$ 1,000\; \mathrm{ }\mathrm{m}\mathrm{m}\mathrm{o}\mathrm{l} \cdot \mathrm{k}\mathrm{g^{-1^{ }}}=\mathrm{ }2.5\; \mathrm{ }\mathrm{M}\mathrm{P}\mathrm{a}\mathrm{ }=\mathrm{ }25\; \mathrm{ }\mathrm{b}\mathrm{a}\mathrm{r}\mathrm{s} $ Chlorophyll fluorescence in plants was measured on a sunny day using a chlorophyll fluorometer)[29]. For 2 min, a fully expanded leaf was acclimated to darkness using a clip, and the leaf-adapted darkness was then continuously irradiated for 1 s (1,500 mol·m−2·s−1) by an array of three light-emitted diodes in the sensor.
Relative stress injury was quantified by determining the ratio of ion leakage into the external aqueous medium to the total ion concentration of the stressed tissue, as assessed through the electrical conductivity of the external medium[30]. The membrane injury was calculated as:
${ \rm RSI}\; ({\text{%}}) = 1-\dfrac{Ec1}{Ec2}\times 100 $ Biochemical attributes
-
Dried samples were ground to 1 mm particles and crude protein (CP), lignin, and fiber were analyzed by near-infrared reflectance spectroscopy (NIRS) using the method given by Fekadu et al.[31]. All results are reported as % dry mass (% DM).
Total soluble sugars were determined using the method given by Yemm & Willis[32].
Statistical analysis
-
The data was analyzed statistically for ANOVA using complete randomized design (CRD) by using SPSS 13.0 (Statistical Package for the Social Sciences) (SPSS Inc., Chicago, IL, USA) using Tukey's HSD test at 0.05 significance level and was expressed as mean ± standard errors. Treatments were compared with CD values at a 5% level of significance. Pearson's correlation analysis was conducted to explore potential associations among the traits, aiming to identify any potential relationships between variables. Correlation analysis was done using different packages of R software version 4.0.5.
-
The growth attributes were significantly influenced by both salt stress and OSA treatments. The data is presented in Tables 1 & 2. According to Table 1, the effect of ortho-silicic acid (OSA) on the fresh weight (FW), plant height (HT), dry weight (DW), and leaf area of sorghum genotype CSV 33 MF was evaluated under different levels of salt stress.
Table 1. Effect of ortho-silicic acid (OSA) on fresh weight stem (FWS), fresh weight leaf (FWL), fresh weight root (FWR), plant height (PH), dry weight stem (DWS), dry weight leaf (DWL), dry weight root (DWR) and leaf area (LA) of Sorghum genotype CSV 33MF grown under different levels of salt stress.
Salt level OSA
(mg·L−1)FWS
(g·plant−1)FWL
(g·plant−1)FWR
(g·plant−1)PH
(cm)DWS
(g·plant−1)DWL
(g·plant−1)DWR
(g·plant−1)LA
(cm2)0 dS·m−1 0 29.73cd ± 0.64 11.40bc ± 0.55 10.06abcd ± 0.29 114.0ab ± 8.39 9.01bc ± 0.19 2.84abc ± 0.14 3.05abc ± 0.09 689.0abc ± 33.4 1.5 33.90ab ± 0.55 12.50ab ± 0.47 11.66ab ± 0.23 129.7a ± 4.18 9.97ab ± 0.16 3.12ab ± 0.12 3.38ab ± 0.08 756.6ab ± 28.7 2.5 34.83a ± 0.46 13.76a ± 0.61 12.13a ± 0.27 136.0a ± 5.29 10.85a ± 0.14 3.43a ± 0.15 3.67a ± 0.07 832.0a ± 35.2 4 dS·m−1 0 28.20de ± 0.83 11.16bc ± 0.38 9.86bcde ± 0.73 112.0ab ± 3.79 8.26cd ± 0.25 2.70bcd ± 0.18 2.98abc ± 0.22 663.6bc ± 43.4 1.5 30.83bcd ± 0.44 12.06ab ± 0.17 11.43abc ± 0.23 120.3a ± 7.36 9.03bc ± 0.15 2.92abc ± 0.05 3.20ac ± 0.05 717.3ab ± 11.2 2.5 32.26abc ± 0.23 12.56ab ± 0.32 11.76ab ± 0.29 124.3a ± 5.90 9.49abc ± 0.06 3.14ab ± 0.08 3.41ab ± 0.23 769.6ab ± 18.4 6 dS·m−1 0 21.03gh ± 0.43 8.60de ± 0.06 7.80ef ± 0.42 82.6cd ± 4.42 5.81ef ± 0.55 2.18d ± 0.01 2.36cde ± 0.13 479.6de ± 03.0 1.5 24.10fg ± 0.40 9.16d ± 0.55 9.33de ± 0.69 93.0bc ± 1.16 6.50e ± 0.53 2.36cd ± 0.28 2.48cde ± 0.36 499.3d ± 29.6 2.5 25.20ef ± 0.55 9.76cd ± 0.27 9.40cde ± 0.12 95.3bc ± 1.33 6.73de ± 0.38 2.46cd ± 0.07 2.75bcd ± 0.07 548.3cd ± 16.0 8 dS·m−1 0 14.40i ± 1.46 5.56f ± 0.37 5.90f ± 0.32 47.7e ± 2.33 3.85g ± 0.45 1.40e ± 0.05 1.80e ± 0.21 302.0f ± 12.2 1.5 17.00i ± 0.89 6.03f ±0.13 6.76f ± 0.41 58.7de ± 4.33 4.64fg ± 0.27 1.47e ± 0.04 2.01de ± 0.10 318.0f ± 05.0 2.5 17.83hi ± 0.53 6.70ef ±0.15 7.16f ± 0.46 63.0de ± 2.08 4.88fg ± 0.16 1.54e ± 0.01 2.11de ± 0.10 340.0f ± 05.1 MSE 1.43 0.43 0.505 12.5 0.298 0.045 0.085 49.8 C.D 2.03 1.11 1.20 13.9 0.92 0.36 0.49 82.8 S.E (m) 0.69 0.38 0.41 4.74 0.31 0.12 0.17 28.2 S.E (d) 0.98 0.54 0.58 6.71 0.45 0.17 0.24 39.9 Data having the same letters in the column do not differ significantly while groups with different letters suggest a significant difference (Tukey's HSD test p < 0.05) with error degree of freedom = 24; MSE (Mean Square Error) at 5%. * Values are presented as mean ± standard error (n = 3). Table 2. Effect of ortho-silicic acid (OSA) on fresh weight stem (FWS), fresh weight leaf (FWL), fresh weight root (FWR), plant height (PH), dry weight stem (DWS), dry weight leaf (DWL), dry weight root (DWR) and leaf area (LA) of Sorghum genotype SSG 59-3 grown under different levels of salt stress.
Salt level OSA
(mg·L−1)FWS
(g·plant−1)FWL
(g·plant−1)FWR
(g·plant−1)PH
(cm)DWS
(g·plant−1)DWL
(g·plant−1)DWR
(g·plant−1)LA
(cm2)0 dS·m−1 0 29.10a ± 0.47 11.53abc ± 1.47 10.03abc ± 0.69 118.7b ± 4.8 8.82b ± 0.14 2.88ab ± 0.37 3.04ab ± 0.21 697.0c ± 31.5 1.5 31.57a ± 0.69 13.77a ± 0.76 10.56ab ± 0.32 132.3ab ± 1.3 9.55ab ± 0.20 3.42a ± 0.18 3.23ab ± 0.14 829.0ab ± 15.6 2.5 32.56a ± 0.54 13.97a ± 0.43 11.23a ± 1.02 140.0a ± 2.5 10.47a ± 0.42 3.63a ± 0.12 3.38a ± 0.40 879.0a ± 19.7 4 dS·m−1 0 27.73ab ± 1.65 11.27abc ± 1.33 9.67abc ± 0.57 116.7bc ± 4.1 8.06bc ± 0.51 2.82abc ± 0.33 2.92ab ± 0.33 681.7c ± 23.7 1.5 29.67a ± 1.30 12.50ab ± 0.35 11.10a ± 0.35 128.0ab ± 2.9 8.58b ± 0.40 3.12a ± 0.08 3.14ab ± 0.10 755.0bc ± 19.9 2.5 31.80a ± 1.67 13.30a ± 0.50 10.26ab ± 0.30 132.3ab ± 0.9 9.01ab ± 0.41 3.32a ± 0.12 3.21ab ± 0.19 804.0ab ± 30.1 6 dS·m−1 0 19.27cd ± 0.38 8.03cdef ± 0.70 7.67cdef ± 0.24 87.0d ± 1.5 5.77d ± 0.12 1.94cdef ± 0.17 2.29bc ± 0.16 360.7de ± 16.2 1.5 21.60c ± 0.66 8.50cde ± 0.45 8.40bcde ± 0.32 98.0d ± 1.5 6.29d ± 0.16 2.06bcde ± 0.11 2.42abc ± 0.12 389.3d ± 03.2 2.5 22.90bc ± 1.78 8.93bcd ± 0.43 8.83abcd ± 0.09 100.7cd ± 5.1 6.56cd ± 0.37 2.18bcd ± 0.13 2.57abc ± 0.23 434.7d ± 14.8 8 dS·m−1 0 12.60e ± 0.50 4.30f ± 0.25 5.70f ± 0.40 53.3e ± 1.2 3.75e ± 0.22 1.12f ± 0.07 1.66c ± 0.08 247.7f ± 17.0 1.5 14.37de ± 0.68 4.63ef ± 0.73 5.87ef ± 0.18 62.7e ± 1.6 4.04e ± 0.21 1.23ef ± 0.09 1.72c ± 0.15 269.0ef ± 25.5 2.5 15.90de ± 0.47 5.07def ± 0.72 6.77def ± 0.46 66.0e ± 2.8 4.21e ± 0.04 1.28def ± 0.10 1.85c ± 0.06 284.7ef ± 08.1 MSE 3.22 1.76 0.772 22.9 0.272 0.100 0.123 19.82 C.D 3.04 2.25 1.49 9.28 0.88 0.54 0.60 59.8 S.E (m) 1.04 0.77 0.51 3.16 0.30 0.18 0.20 20.4 S.E (d) 1.46 1.08 0.72 4.47 0.43 0.26 0.29 28.8 Data having the same letters in the column do not differ significantly while groups with different letters suggest a significant difference (Tukey's HSD test p < 0.05) with error degree of freedom = 24; MSE (Mean Square Error) at 5%. * Values are presented as mean ± standard error (n = 3). Fresh weight
-
The fresh weight of stem, leaf, and root showed a drastic decline under salt stress. OSA was able to mitigate the negative effects of salt stress to some extent. At control (0), the highest percent increase in FW was observed at 2.5 mg·L−1 of OSA, with stem FW increasing by 17.1%, leaf FW by 20.7%, and root FW by 20.6% in CSV 33MF. Exogenous OSA application also resulted in increased FW of stem, leaf, and root at 4 dS·m−1 with the highest increase observed at 2.5 mg·L−1, with stem FW increasing by 14.4%, leaf FW by 12.5 %, and root FW by 19.3%. Under 8 dS·m−1 salt level, the highest decline in fresh weight was observed which was mitigated to some extent by OSA application which increased FW of stem, leaf, and root. The highest percent increase in FW was observed at 2.5 mg·L−1 of OSA, with stem FW increasing by 23.8%, leaf FW by 20.5%, and root FW by 21.4%. Similar trends were observed in Table 2 demonstrating the impact of salt stress and OSA on sorghum genotype SSG 59-3. The imposition of elevated salt stress levels resulted in a significant decrease in the fresh weight of leaves, with the maximum reduction of 62.6% observed at 8 dS·m−1 in comparison to the control. However, the application of ortho-silicic acid (OSA) at concentrations of 1.5 and 2.5 mg·L−1 led to an increase in the fresh weight of leaves, with more pronounced enhancements observed at 2.5 mg·L−1.
Dry weight
-
The dry weight of the leaf, stem and root showed a declining trend with the increasing salt level from control to 8 dS·m−1. Fold change was found maximum in leaf dry weight at 8 dS·m−1 of salt stress i.e. 0.59 and 0.55 in CSV33MF and SSG 59-3 respectively, with respect to the control. Dry weight enhancement was noticed after exogenous OSA application. Maximum increase was observed at 2.5 mg·L−1 OSA application. After the application of 2.5 mg·L−1 of OSA, the percent increase was maximum at the control of salt stress in both genotypes i.e. CSV33MF (20.6%) and SSG 59-3 (11.2%) with respect to the control. Likewise, the dry weight of leaves exhibited a declining trend with escalating salt stress levels in the SSG 59-3 genotype. The highest percentage decline of 45.39% was observed at 8 dS·m−1 of salt stress compared to the control. However, the foliar application of OSA resulted in an increment in the dry weight of leaves, with the maximum enhancement observed at 2.5 mg·L−1. Similar results were also obtained for dry weight of stem and root under different levels of salt stress and foliar application of ortho-silicic acid. The contribution towards dry weight was more from the stem as compared to the root.
Plant height
-
The plant height was reduced in both genotypes due to the effect of salt stress. At 0 dS·m−1 salt level, plant height was recorded highest (136.0 ± 5.29 cm) at 2.5 mg·L−1 of OSA demonstrating a fold change of 1.19 as compared to control conditions of treatment. The plant height also showed a percent increase of 12.6% and 15.7% at 1.5 mg·L−1 and 2.5 mg·L−1 of OSA, respectively as compared to the control at 6 dS·m−1. The plant height exhibited a significant percent increase of 32.2% at 8 dS·m−1 after the exogenous application of 2.5 mg·L−1 OSA as compared to the control of the treatment. Plant height also experienced a significant reduction with increasing levels of salt stress in the SSG 59-3 genotype. At 8 dS·m−1, there was a higher percentage decline of 55.4% compared to the control level. The foliar application of OSA at concentrations of 1.5 and 2.5 mg·L−1 resulted in significant enhancements in plant height, with the maximum increase of 23.8% observed at 8 dS·m−1 at 2.5 mg·L−1 OSA.
Leaf area
-
Leaf area per plant decreased with increasing salt levels ranging from control to 8 dS·m−1. The highest decline in leaf area was observed at 8 dS·m−1 of salt level, but on comparison of both genotypes, SSG 59-3 (64.4%) had a maximum decrease in leaf area per plant as compared to CSV 33MF (56.2%) with respect to control. Leaf area per plant was increased after the foliar spray of ortho-silicic acid at 1.5 and 2.5 mg·L−1, but the maximum rise in leaf area was observed at 2.5 mg·L−1 of OSA. The leaf area showed a significant percent increase of 20.7% at 2.5 mg·L−1 of OSA compared to the control at 0 dS·m−1. After exogenous application of 2.5 mg·L−1 OSA, the leaf area also exhibited a significant percent increase of 15.9% at 4 dS·m−1 and 14.3 % at 6 dS·m−1 in CSV33MF genotype. The most drastic reduction in leaf area was observed at 8 dS·m−1 which was 56.16% in CSV33MF and 64.56% in SSG 59-3. Exogenous application of 2.5 mg·L−1 OSA mitigated the effects of salt stress on leaf area and increased leaf area by fold change of 1.13 in CSV33MF and 1.15 in SSG 59-3.
Overall, both genotypes exhibited reductions in fresh weight, dry weight, plant height, and leaf area per plant as salt stress levels increased. Nevertheless, the application of OSA mitigated these adverse effects and led to improvements in these growth parameters, with greater enhancements observed at 2.5 mg·L−1 of OSA.
Physiological attributes
-
The leaf water potential values became increasingly negative as salt levels were incremented from the control to 8 dS·m−1 in both genotypes at 40 Days after sowing (DAS) (Tables 3 & 4). A comparison of the two genotypes revealed that SSG 59-3 exhibited the most negative value (−1.15 MPa) compared to CSV33MF (−1.13 MPa) at the 8 dS·m−1 salt level. The water potential of the leaves became less negative when different concentrations of ortho-silicic acid were applied to both genotypes. SSG 59-3 displayed a less mean negative value (−0.50 MPa) compared to CSV33MF (−0.57 MPa) after the application of 2.5 mg·L−1 OSA at control. The relative water content (RWC) progressively decreased with increasing salt stress levels at 40 DAS in both genotypes. The maximum decrease in RWC was observed at the 8 dS·m−1 salt level in both genotypes, with a decrease of 18.3% in CSV33MF and 18.5% in SSG 59-3 compared to their respective controls. The foliar application of 2.5 mg·L−1 of ortho-silicic acid increased the RWC at each level of salt stress in both genotypes. In CSV33MF, the RWC increased from 81.2% to 85.5% at 4 dS·m−1, from 71.6% to 76.1% at 6 dS·m−1, and from 66.2% to 72.0% at 8 dS·m−1. A similar enhancement in RWC was observed in SSG 59-3. Relative Stress Injury exhibited an increasing trend with the imposition of salt stress, ranging from the control to 8 dS·m−1. The maximum leakage was estimated at the 8 dS·m−1 salt level in both genotypes, with SSG 59-3 showing more damage (42.3%) compared to CSV33MF (36.8%) with respect to the control. The foliar application of 2.5 mg·L−1 of ortho-silicic acid prevented electrolyte leakage to some extent.
Table 3. Effect of ortho-silicic acid (OSA) on water potential (Ψw), Relative Water Content (RWC), Relative Stress Injury (RSI), Transpiration Rate (E), Stomatal Conductance (gs), Assimilation Rate (A), Chlorophyll Fluoroscence (CHLF) and Chlorophyll content (CHL) of Sorghum genotype CSV33MF grown under different levels of salt stress.
Salt level OSA (mg·L−1) Ψw (MPa) RWC (%) RSI (%) E (mmol H2O·m−1·s−1) gs (mmol H2O·m−1·s−1) A (μmol CO2·m−1·s−1) CHLF (Fv/Fm) CHL (SPAD) 0 dS·m−1 0 −0.673bc ± 0.023 84.45abc ± 5.49 11.13f ± 0.39 3.58ab ± 0.07 0.103abc ± 0.012 11.12bcd ± 0.80 0.729abc ± 0.011 36.6abc± 4.3 1.5 −0.583ab ± 0.015 85.89ab ± 2.14 10.54f ± 0.72 3.88ab ± 0.25 0.120ab ± 0.010 12.09abc ± 1.10 0.739ab ± 0.001 39.2ab ± 2.6 2.5 −0.496a ± 0.012 87.74a ± 2.10 10.45f ± 0.33 4.14a ± 0.06 0.147a ± 0.009 14.68a ± 0.31 0.772a ± 0.008 42.5a ± 1.4 4 dS·m−1 0 −0.783de ± 0.013 81.19abcd ± 1.98 21.30de ± 2.12 3.35bc ± 0.17 0.093abc ± 0.012 10.78bcd ± 0.53 0.703bcde ± 0.008 33.5bcde± 1.3 1.5 −0.700cd ± 0.012 84.20abc ± 3.03 20.63de ± 1.91 3.69ab ± 0.07 0.107abc ± 0.009 11.87abc ± 0.25 0.710bcd ± 0.006 35.0bcd ± 1.9 2.5 −0.667bc ± 0.018 85.50ab ± 1.38 18.30e ± 0.55 4.04ab ± 0.13 0.127ab ± 0.015 12.55ab ± 0.31 0.719abcd ± 0.004 38.7abc ± 2.3 6 dS·m−1 0 −0.996h ± 0.009 71.59de ± 4.01 28.97bc ± 0.55 1.80ef ± 0.10 0.050bc ± 0.010 7.84de ± 1.29 0.679cde ± 0.010 28.7def ± 1.3 1.5 −0.893fg ± 0.003 74.32cde ± 0.39 27.30bc ± 0.72 2.09de ± 0.12 0.063bc ± 0.009 8.94cde ± 1.03 0.723abcd ± 0.003 32.5cde ± 2.3 2.5 −0.833ef ± 0.018 76.07bcde ± 2.15 24.03cd ± 0.07 2.71cd ± 0.23 0.083abc ± 0.012 9.27bcde ± 0.45 0.736abc ± 0.003 34.3bcd ± 1.9 8 dS·m−1 0 −1.127i ± 0.023 66.17e ± 4.00 36.80a ± 0.61 1.09f ± 0.11 0.030c ± 0.010 6.21e ± 0.24 0.650e ± 0.005 21.8g ± 1.2 1.5 −1.010h ± 0.012 70.30e± 1.48 34.23a ± 0.61 1.57ef ± 0.16 0.047bc ± 0.009 7.31e ± 0.33 0.665de ± 0.009 25.6fg ± 1.0 2.5 −0.957gh ± 0.035 71.97de ± 4.42 31.87ab ± 1.12 2.16de ± 0.06 0.070abc ± 0.010 8.20de ± 0.10 0.687bcde ± 0.015 27.5efg ± 2.7 MSE 0.001 5.89 3.02 0.06 0.0008 1.372 0.0004 4.84 C.D 0.052 5.13 2.96 0.42 0.031 1.98 0.023 3.73 S.E (m) 0.018 1.76 1.00 0.14 0.011 0.68 0.008 1.27 S.E (d) 0.025 2.49 1.42 0.20 0.015 0.96 0.011 1.80 Data having the same letters in the column do not differ significantly while groups with different letters suggest a significant difference (Tukey's HSD test p < 0.05) with error degree of freedom = 24; MSE (Mean Square Error) at 5%. * Values are presented as mean ± standard error (n = 3). Table 4. Effect of ortho-silicic acid (OSA) on water potential (Ψw), Relative Water Content (RWC), Relative Stress Injury (RSI), Transpiration Rate (E), Stomatal Conductance (gs), Assimilation Rate (A), Chlorophyll Fluoroscence (CHLF) and Chlorophyll content (CHL) of Sorghum genotype SSG 59-3 grown under different levels of salt stress.
Salt level OSA (mg·L−1) Ψw (MPa) RWC (%) RSI (%) E (mmol H2O·m−1·s−1) gs (mmol H2O·m−1·s−1) A (μmol CO2·m−1·s−1) CHLF (Fv/Fm) CHL (SPAD) 0 dS·m−1 0 −.713bc ± 0.026 82.47ab ± 1.98 12.93f ± 0.43 3.30bc ± 0.13 0.097abc ± 0.009 10.85abc ± 0.61 0.713abc ± 0.006 33.5cd ± 0.7 1.5 −0.630ab ± 0.015 84.84a ± 2.44 11.77f ± 0.73 3.74ab ± 0.08 0.117a ± 0.009 11.54ab ± 0.30 0.738a ± 0.014 37.4ab ± 1.1 2.5 −0.570a ± 0.025 86.48b ± 1.69 11.33f ± 0.33 4.02a ± 0.15 0.136a ± 0.015 13.34a ± 0.76 0.751a ± 0.001 40.1a ± 0.6 4 dS·m−1 0 −0.837cd ± 0.017 79.29ab ± 1.88 22.83e ± 0.58 3.15c ± 0.06 0.093abcd ± 0.008 10.14bcd ± 1.17 0.710abc ± 0.001 31.1de ± 0.5 1.5 −0.743bc ± 0.015 82.32ab ± 1.72 22.06e ± 0.80 3.42bc ± 0.07 0.103ab ± 0.012 11.90ab ± 0.12 0.714abc ± 0.001 34.2bcd ± 0.6 2.5 −0.713bc ± 0.020 84.59a ± 2.10 19.57e ± 0.58 3.58abc ± 0.09 0.128a ± 0.015 12.71ab ± 0.16 0.724ab ± 0.006 37.3abc ± 0.4 6 dS·m−1 0 −1.023ef ± 0.052 69.96cd ± 0.71 32.47c ± 0.60 1.54ef ± 0.16 0.030e ± 0.010 6.84e ± 0.21 0.657cd ± 0.009 26.4fg ± 1.0 1.5 −0.943de ± 0.029 74.45bc ± 0.78 30.60c ± 0.78 2.04de ± 0.08 0.040cde ± 0.010 7.40de ± 0.07 0.700abc ± 0.001 29.5ef ± 0.9 2.5 −0.877d ± 0.009 75.76bc ± 1.78 26.90d ± 0.10 2.55d ± 0.08 0.057bcde ± 0.009 8.27cde ± 0.08 0.715ab ± 0.008 32.2de ± 0.5 8 dS·m−1 0 −1.150f ± 0.042 63.96d ± 0.68 42.33a ± 0.69 0.92g ± 0.14 0.017e ± 0.008 5.81e ± 0.23 0.637d ± 0.001 20.0h ± 0.8 1.5 −1.063ef ± 0.044 68.76cd ± 1.09 39.40ab ± 0.71 1.32fg ± 0.06 0.020e ± 0.009 6.12e ± 0.33 0.665cd ± 0.014 23.5gh ± 0.9 2.5 −1.030ef ± 0.010 69.45cd ± 1.82 36.63a ± 1.30 1.61ef ± 0.07 0.037de ± 0.010 7.11e ± 0.70 0.678bcd ± 0.001 25.7fg ± 0.8 MSE 0.002 8.28 1.45 0.31 0.0008 0.96 0.0004 1.65 C.D 0.084 4.88 2.04 0.16 0.031 1.30 0.021 2.18 S.E (m) 0.029 1.66 0.69 0.05 0.011 0.45 0.007 0.74 S.E (d) 0.040 2.35 0.98 0.07 0.015 0.63 0.010 1.05 Data having the same letters in the column do not differ significantly while groups with different letters suggest a significant difference (Tukey’s HSD test p < 0.05) with error degree of freedom = 24; MSE (Mean Square Error) at 5%. * Values are presented as mean ± standard error (n = 3). The chlorophyll content also declined with increasing salt stress levels from the control (0) to 8 dS·m−1 in both genotypes. The maximum fold change was observed at the 8 dS·m−1 salt level, contributing to a fold change of approximately 0.6 in both CSV33MF and SSG 59-3 compared to the control. A progressive increase in chlorophyll content was observed in both genotypes with the application of different concentrations of ortho-silicic acid. The fold change was maximum at 2.5 mg·L−1 of ortho-silicic acid in CSV33MF (1.285) and SSG 59-3 (1.207) at the 8 dS·m−1 salt level compared to the control. Chlorophyll fluorescence or photochemical quantum yield also declined with the imposition of salt stress in both genotypes at 40 DAS. The quantum yield values decreased from the control to 8 dS·m−1, ranging from 0.73 to 0.65 in CSV33MF and 0.71 to 0.64 in SSG 59-3, respectively. The application of 2.5 mg·L−1 of ortho-silicic acid resulted in a progressive increase in quantum yield at each salt level, as well as in the control. The maximum increase was noticed at the 6 dS·m−1 salt level in CSV33MF, where it increased from 0.68 to 0.74.
The rate of photosynthesis decreased with increasing salt stress levels from the control (0) to 8 dS·m−1 in both genotypes at 40 DAS. The percent decrease in photosynthetic rate was 44.2% in CSV33MF and 46.5% in SSG 59-3 at the 8 dS·m−1 salt level compared to the control. The foliar application of 2.5 mg·L−1 of ortho-silicic acid enhanced the rate of photosynthesis in both genotypes. The percent increase was noticed at each level of salt stress in CSV33MF (14.1%, 15.4%, and 32.2% at 4, 6, and 8 dS·m−1, respectively) compared to the control. Similar increments were also observed in genotype SSG 59-3. A progressive decline was noticed in the rate of transpiration with an increase in salt levels from the control to 8 dS·m−1. The fold change in transpiration rate was highest at the 8 dS·m−1 salt level, with 0.305 in CSV33MF and 0.282 in SSG 59-3 compared to the control. At the highest salt level (8 dS·m−1), the application of 2.5 mg·L−1 of ortho-silicic acid showed a significant increase in transpiration rate (88.7% in CSV33MF and 73.1% in SSG 59-3). Stomatal conductance also decreased with an increase in salt levels in both genotypes. The percent decline in stomatal conductance observed at 4, 6, and 8 dS·m−1 salt levels was 5.0%, 50.2%, and 70.1%, respectively, in CSV33MF, and 5.2%, 68.4%, and 84.4% in SSG 59-3 compared to their respective controls. The foliar application of ortho-silicic acid (1.5 and 2.5 mg·L−1) led to an improvement in stomatal conductance in both genotypes, while the maximum increase was observed at 2.5 mg·L−1 OSA at 8 dS·m−1 leading to a fold change of 2.33 in the CSV33MF genotype.
Biochemical parameters
-
A decrease in protein content was observed with the increase in salt levels from control to 8 dS·m−1 at 40 DAS as described in Fig. 1. In the sorghum genotypes CSV33MF and SSG 59-3, protein content decreased from 8.89 to 6.61 and 8.59 to 6.72% DM, respectively, with the onset of salt stress (0 to 8 dS·m−1). The foliar application of ortho-silicic acid (2.5 mg·L−1) increased protein content under both stressed and control conditions. The maximum increase in protein content (23.9% in CSV33MF and 15.2% in SSG 59-3) was observed with 2.5 mg·L−1 OSA at 8 dS·m−1 salt level compared to their respective controls.
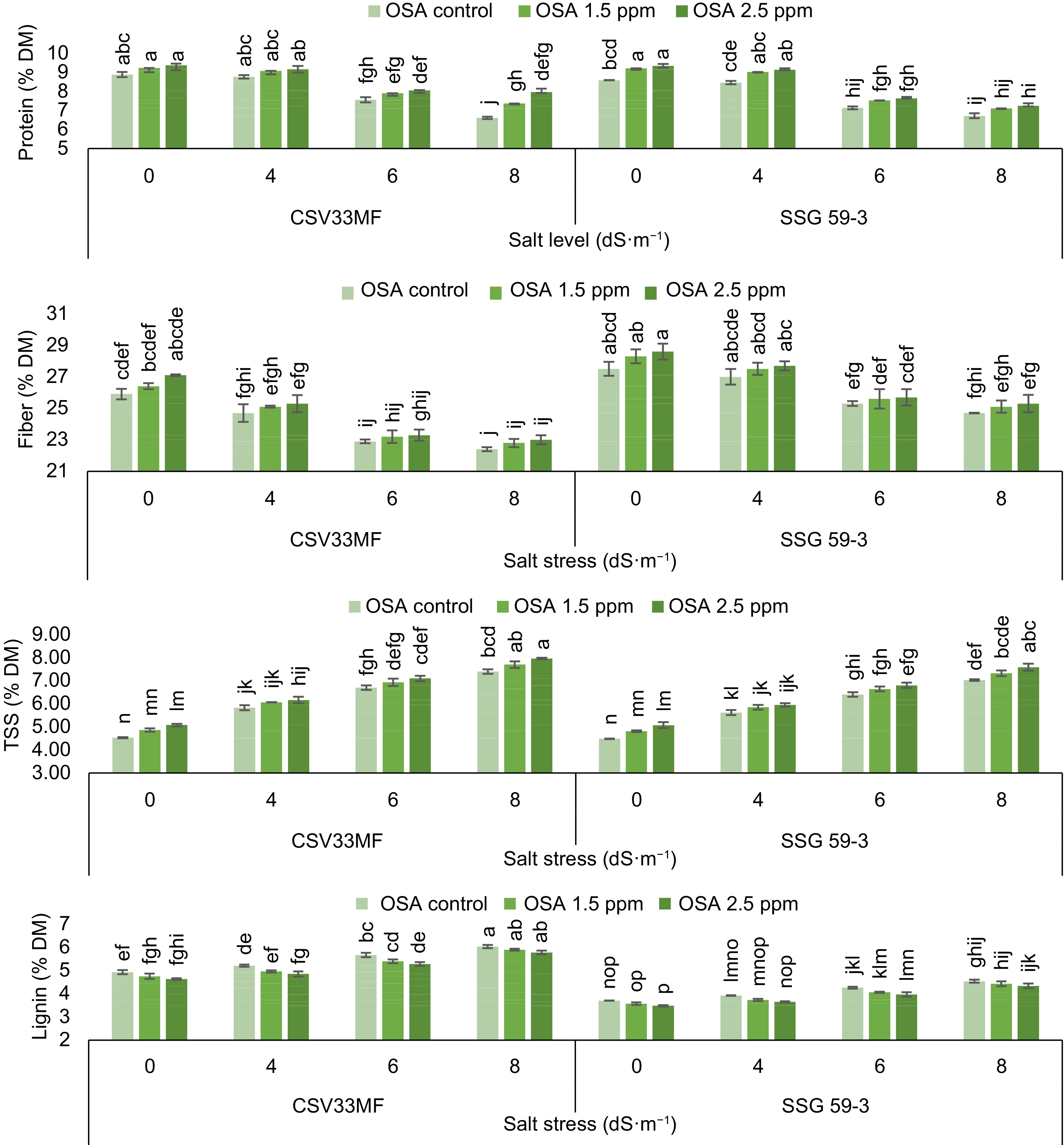
Figure 1.
Effect of ortho-silicic acid (OSA) on protein (% DM), fiber (% DM), Total Soluble Sugar (TSS) (% DM) and lignin (% DM) of Sorghum genotypes CSV33MF and SSG 59-3 grown under different levels of salt stress at 40 DAS. Data having the same letters in the column do not differ significantly while groups with different letters suggest a significant difference (Tukey's HSD test p < 0.05) with error degree of freedom = 48.
The total soluble sugar (TSS) content of both sorghum genotypes was significantly influenced by both salt stress (4, 6, and 8 dS·m−1) and OSA (1.5 and 2.5 mg·L−1). The results revealed that plants grown under stress conditions exhibited higher TSS values compared to the control. The maximum TSS content was estimated at 8 dS·m−1 salt level in both genotypes (CSV33MF and SSG 59-3). Significant increases in TSS content were observed with the application of ortho-silicic acid at 1.5 and 2.5 mg·L−1 concentrations. The highest increase was observed in CSV33MF at 8 dS·m−1 salt level, where TSS content increased from 7.40 to 7.97 at 2.5 mg·L−1 OSA.
The fiber content significantly decreased with increasing levels of salt stress from control to 8 dS·m−1 in both genotypes at 40 DAS. The decrease was 14.1% in CSV33MF and 10.8% in SSG 59-3 at 8 dS·m−1 compared to the control. Application of all concentrations of ortho-silicic acid caused an increase in fiber content in both genotypes under stressed and control conditions, but the maximum increment was observed with 2.5 mg·L−1 OSA in CSV33MF compared to SSG 59-3 under control conditions. Lignin content increased with each increment in salt levels in both genotypes at 40 DAS. The percentage increase in lignin content was 5.6%, 14.9%, and 18.3% at 4, 6, and 8 dS·m−1 salt levels, respectively, in CSV33MF compared to their respective controls. Foliar application of OSA (1.5 and 2.5 mg·L−1) led to an increase in lignin content in both genotypes, with the maximum increase observed at 2.5 mg·L−1. The percentage increase was higher at 8 dS·m−1 salt level in CSV33MF (4.5%) and at 6 dS·m−1 in SSG 59-3 (6.8%) compared to their respective controls.
-
Salt stress has a detrimental influence on plant development (germination, vegetative growth, and reproductive development), physiology and biochemical aspects along with its water and nutrient uptake either directly or indirectly. A comprehensive review of the literature revealed a scarcity of well-maintained work on the effects of ortho-silicic acid on the morpho-physiological and biochemical responses of sorghum to salt stress. The nutritional quality of sorghum also relies on these components, therefore evaluation of these parameters under salt stress might reveal the relative importance of these traits in maintaining nutritional quality which can be further improved by crop improvement program of forage sorghum for salt stress tolerance.
Morpho-physiological changes in response to OSA treatment under salt stress
-
Growth variables are an indispensable tool for assessing crop productivity across various species. It is influenced by an array of factors, including genetic makeup, physiological and biochemical factors along with environmental factors. Under salt stress, both the genotypes of sorghum (CSV33MF and SSG 59-3) exhibited a significant decline in plant height, fresh and dry weight, leaf number, leaf area per plant, and specific leaf weight. This reduction in growth parameters intensified with increasing salt concentration, validating findings also reported by Bimurzayev et al.[33]. The presence of salts in the soil lowers the solute potential, leading to a reduction in water imbibition by roots, which subsequently impairs plant growth and development. This decrease in growth could be linked to negative effects on physiological processes such as osmotic imbalance, ion stresses, transpiration, and photosynthesis[34]. Increased salt tolerance in plants is often associated with higher antioxidant activity and cell membrane protection, contributing to decreased lipid peroxidation as well as improved leaf photosynthetic rate and stomatal conductance. For this, in-depth investigations are warranted. For instance, a recent opportunity has suggested that mapping the movement of cell apoplastic ion and metabolite patterns could offer novel insights into the function of guard cells[35], particularly under salinity. This understanding could shed light on the maintenance of photosynthesis and biomass accumulation with silicon application under salinity. OSA has shown potential in alleviating salt stress by regulating antioxidant activity. Foliar application of OSA mitigates salt-induced damage and enhances photosynthetic characteristics of plants, evidenced by improved photosynthetic pigment content, photosynthetic rate, stomatal conductance, and intercellular CO2 concentration[36]. Under abiotic stress, OSA can promote meristem division and plant development through improved nutrient uptake and reduced oxidative damage caused by ROS generation[37]. Additionally, OSA contributes to plant growth by enhancing multiple adaptive responses, including organic acids and phenol exudation, accumulation of compatible solutes and hormonal regulation[38].
Biochemical changes in response to OSA treatment under salt stress
-
In leaves, intense soluble sugar accumulation was observed following salt treatment in all the genotypes of wheat and barley. This accumulation is attributed to increased invertase activity, which converts sucrose to fructose and glucose, thus raising the total soluble sugar content[39]. Elevated levels of sugar metabolites in the leaves of salt-tolerant cultivars, functioning as osmo-protectants, helping the plants cope with osmotic stress[40].
Silicon (Si) plays a crucial role in altering the gene expression of Si transporters (Lsi1 and Lsi2) and stress-related proteins, leading to increased silica accumulation and higher levels of compatible solutes in Si-supplemented plants[41]. Similarly, maize plants grown from seeds treated with Si for 12 h showed increased growth, leaf-relative water content, and levels of photosynthetic pigments, soluble sugars, soluble proteins, total free amino acids, potassium, and activities of SOD, CAT, and POD enzymes, compared to untreated plants[42]. Increased lignin deposition has been observed in Si-treated (Si+) plants compared to untreated (Si−) plants. Deposition of lignin at the inner tangential cell wall starts together with the formation of the silica aggregates in Si+ roots[43]. Upregulation of phenylpropanoid biosynthesis-related genes under -Si conditions. Due to this -Si plants exhibit increased mechanical strength and calorific value in cell walls due to elevated lignin deposition. Limiting the Si supply also significantly increased the thioglycolic acid lignin content and thioacidolysis-derived syringyl/guaiacyl monomer ratio[44]. Flores et al. reported that silicon application improved the physiological quality, dry mass, and fibre production of Sorghum bicolor[45]. The accumulation of large amounts of silicon in the endoderm tissues of sorghum increased the crop's tolerance to stress including water deficit as the endoderm cells play an important role in water transportation by roots. Also, the application of silicon increased the protein content in sorghum seeds. The study focused on the nutritional and functional properties of sorghum-based products and the potential valorization of sorghum bran. The addition of silicon, along with micro minerals and soybeans, enhanced the protein content in polished sorghum-based products[46]. In the present study, OSA alleviated the negative impact of stress on growth and yield, consistent with previous findings. Si application enhanced levels of photosynthetic pigments, relative water content, protein content, and carbohydrate content in both wheat varieties (WH-1105 and KRL-210). It also reduced proline content, malondialdehyde (MDA) content (an indicator of lipid peroxidation), and electrolyte leakage, thereby improving salt stress tolerance. Furthermore, Si reduces Na+ absorption, improves ion balance, and alleviates the adverse effects of salt stress in two licorice species, as demonstrated by previous studies[47,48].
All morphological parameters were observed to have a highly significant negative correlation with RSI, while conferring a highly significant positive correlation with A, gs, RWC, E and CHL. Figure 2 provides information regarding the correlation values and patterns among the different traits varying across all the treatments. The data is highly significant for approximately all parameters having a p-value of < 0.001. Correlations between different traits were determined using Pearson correlation coefficients (PCCs). Strong correlation was observed among all the physiological traits except RSI, and with the growth traits. Among the biochemical traits, protein and fibre showed a positive correlation with physiological and growth traits whereas lignin and TSS showed a negative correlation.
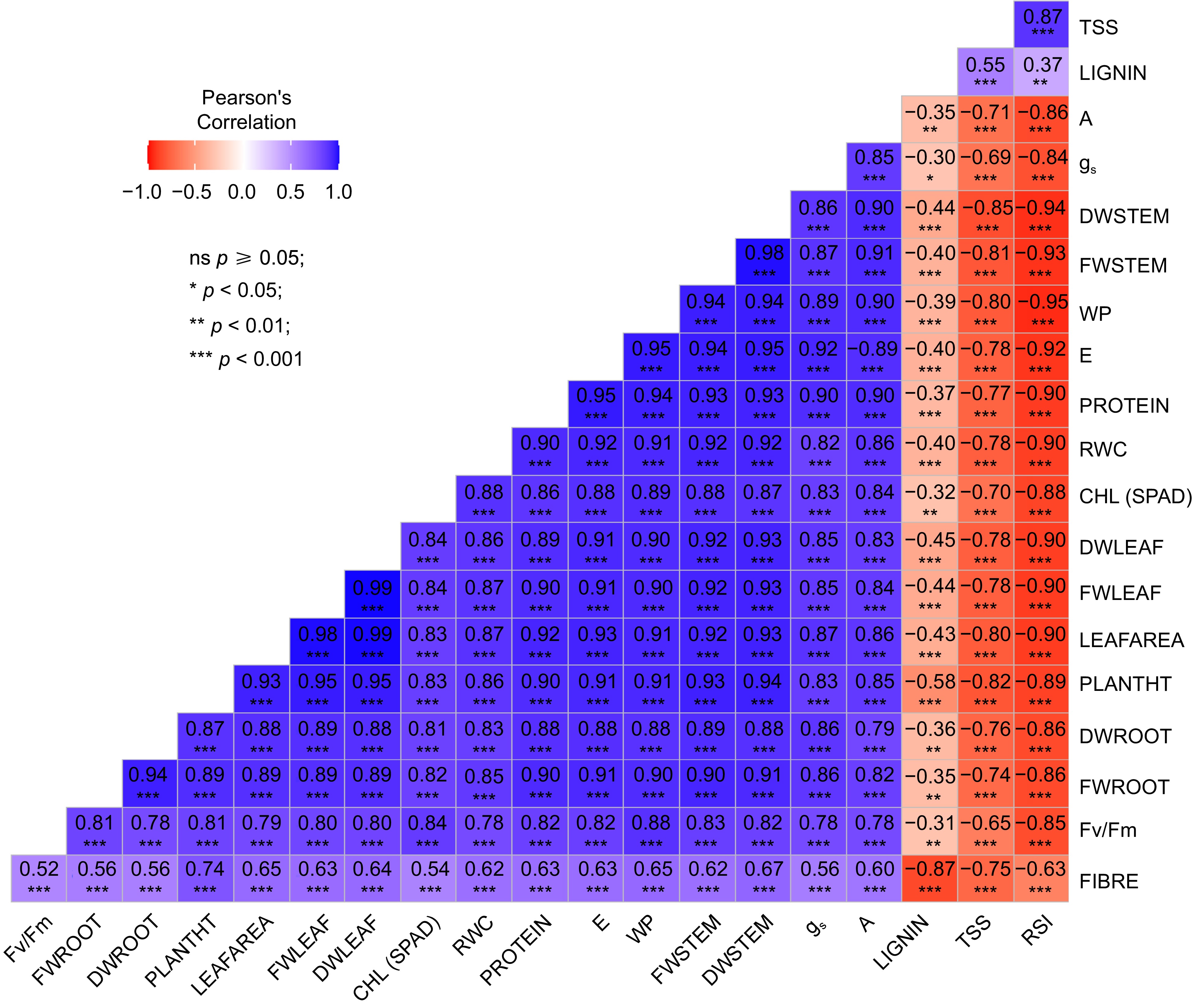
Figure 2.
Pattern of correlation and level of significance observed among different traits across all the treatments in both sorghum genotypes. TSS (Total Soluble Sugar), PROTEIN, FIBRE, LIGNIN, A (Assimilation Rate), gs (Stomatal Conductance), DW (Dry Weight of Stem, Leaf and Root), FW (Fresh Weight of Stem, Leaf and Root), WP (Water Potential), E (Transpiration Rate), RWC (Relative Water Content); PLANTHT (Plant Height), Fv/Fm (Chlorophyll Fluoroscence), CHL (Chlorophyll Content).
-
Salt stress is a major environmental factor limiting global plant growth and productivity. The sorghum varieties CSV33MF and SSG 59-3, grown in northern India, are significantly affected by salinity and sodicity. Silicon (Si) has shown potential as an abiotic stress mitigator. Salt stress negatively impacted morpho-physiological and biochemical traits in both sorghum genotypes, with SSG 59-3 experiencing greater reduction. Foliar application of ortho-silicic acid (OSA) improved various growth parameters, with 2.5 mg·L−1 OSA being more effective than 1.5 mg·L−1. OSA significantly mitigated the effect of salt stress up to 6 dS·m−1 and demonstrated improved productivity at all salt levels. OSA enhances osmotic balance and water use efficiency (WUE), preventing salt-induced damage in plants. Sorghum is a vital crop in the context of climate change due to its abiotic stress resilience ensuring food security in increasingly adverse conditions. These findings suggest the potential inference of using exogenous ortho-silicic acid to mitigate salt stress.
-
The authors confirm contribution to the paper as follows: study conception and design: Kumari G, Satpal, Lakra N, Arya SS; data collection, data analysis and interpretation of results, draft manuscript preparation, writing, review and editing: Pankaj, Devi S, Ahlawat YK, Dhaka P. All authors reviewed the results and approved the final version of the manuscript.
-
All data generated or analyzed during this study are included in this published article, and are available from the corresponding author upon reasonable request.
This work was supported by the Chaudhary Charan Singh Haryana Agricultural University, Haryana, India.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Pankaj, Devi S, Dhaka P, Kumari G, Satpal, et al. 2024. Enhancing salt stress tolerance of forage sorghum by foliar application of ortho-silicic acid. Grass Research 4: e016 doi: 10.48130/grares-0024-0014
Enhancing salt stress tolerance of forage sorghum by foliar application of ortho-silicic acid
- Received: 27 April 2024
- Revised: 08 June 2024
- Accepted: 03 July 2024
- Published online: 16 July 2024
Abstract: Soil salinity poses a significant threat to global food security as salt-affected soils are expected to increase more under the influence of climate change. Sorghum is the world's 5th most important cereal crop and is moderately salt tolerant. Salt stress causes osmotic stress in sorghum and induces several physiological changes, such as membrane disruption, reactive oxygen species (ROS) generation, nutrient imbalance, decreased photosynthetic activity, and decreased stomatal aperture. This research focused on minimizing the detrimental effects of soil salinity on crop productivity by exploring the potential of ortho-silicic acid (OSA) as a mitigating agent for salt stress and analyzing its impact on growth, physiological and biochemical attributes. A pot experiment was performed under control and 4, 6, and 8 dS·m−1 NaCl with OSA concentrations of 1.5 and 2.5 mg·L−1. Results indicated that OSA application improved growth attributes, including fresh weight, plant height, dry weight, and leaf area, under various salt stress levels. Physiological attributes such as photosynthesis rate, transpiration rate, stomatal conductance, and relative water content were 23.7%, 32.4%, 51.3%, and 6.4% higher under 2.5 mg·L−1 OSA treatment, respectively compared to control. Nutritional attributes such as crude protein, fiber, total soluble sugars, and lignin were also improved under OSA treatment. The concentration of 2.5 mg·L−1 OSA treatment was found to be more effective under saline and non-saline conditions for increasing sorghum productivity. This research offers a promising strategy to increase crop productivity and resilience in the face of escalating soil salinity due to climate change.
-
Key words:
- Climate change /
- OSA /
- Photosynthesis /
- Silicon /
- Sorghum bicolor (L.)












