-
Cenchrus is a highly diversified genus[1], with around 145 extant species[2]. At least for the Elephant, Taiwan, and Mott varieties, its production potential has been proven under adequate irrigation and fertilization conditions[3]. It is a species native to Africa, and it has been introduced to tropical and subtropical countries, where it has been used as a fodder crop due to its high biomass, nutritional content, and adaptability to diverse climates and soil types[4]. However, weather conditions in the tropics entail well-defined rain and drought periods, which affect crop productivity[5].
Cenchrus purpureus (Schumach.) Morrone, exhibits extensive genetic diversity, which contributes to its adaptability to various environmental conditions. This diversity, stemming from natural variation and selective breeding, has led to cultivars suited to different climates and soils[4]. The species is highly adaptable, thriving in areas with varying rainfall levels[5], and poor soil conditions[4,6]. Its deep root system enhances drought resilience, making it a critical species for fodder production, bioenergy production, and soil conservation in sustainable agriculture, where it helps reduce erosion and improve soil structure[4].
C. purpureus provides a range of economic advantages for local farmers and livestock, such as enhanced feed quality, reduced costs, and the possibility of sustainable agricultural practices. For example, previous studies[7−9] showed the current yield, crude protein, metabolizable energy, neutral and acid detergent fiber of C. purpureus grass to be 26.0−30.0 t/ha/year, 6.5%−9.6%, 1.89−2.06 Mcal/kg, 64.1%−67.3%, and 38.8%−43.1%, respectively, suggesting an opportunity for significant improvement of both the yield and nutritive value of this grass. Likewise, other authors[10,11] highlight the potential of various techniques to enhance the quality of C. purpureus grass and these improvements could lead to increased economic profitability and sustainability in livestock farming within tropical regions.
The use of C. purpureus accessions, such as King grass, Maralfalfa, Elephant grass, Merkeron, Purple grass, CT-115, and Taiwan, is widespread in tropical regions of Mexico. King grass is favored for its height and regrowth under intensive management[12,13]. Maralfalfa is valued for its fast growth and high digestibility, making it ideal for meat and milk production, Elephant grass is famous for its resilience and versatility[13]. Merkeron is preferred in drought-prone areas, and Purple grass is used for high yields and pest resistance[13], while CT-115[12,13], and Taiwan[12−14] are favored for their nutritional value and productivity in intensive systems.
This fodder crop is a multifunctional species which not only supports agricultural productivity but also plays a crucial role in environmental sustainability through its contributions to soil conservation, carbon sequestration, and the provision of ecosystem services[11]. Soil pH, base exchange capacity, and cation exchange capacity, along with phosphorus, potassium and magnesium levels, decrease after ten years of continuous grazing despite a slight increase in organic matter, suggesting increased acidity and a reduction in essential nutrients[11]. On the other hand, the use of robust methodologies, such as variance analysis and regressions in growth analysis, allows for the identification of factors that maximize biomass yield and improve the sustainability of C. purpureus cultivation under different environmental conditions[15].
Fodder production and quality must be carefully evaluated, and proper management decisions should be based on this evaluation to improve productivity in cattle farming units, and to adapt to the conditions created by climate change[16]. The modified environmental conditions (climate variability) have not been thoroughly investigated, nor has their global impact been evaluated[17]; in the case of HAV, the months are classified as abnormally dry have increased to 10 from 2016 to 2021, and the region suffered from extreme drought in some years within this period[18]. It was hypothesized that some of these genotypes might be capable of producing higher yields despite adverse environmental conditions. Thus, fresh weight, dry matter, and agronomic traits of seven accessions of Cenchrus purpureus (Taiwan grass, King grass, Elephant grass, Merkeron, CT-115, Purple grass, and Maralfalfa) were evaluated under the current climate conditions of the HAV region.
-
The research was carried out at the experimental field of the Instituto Tecnológico Superior de Tantoyuca, in the municipality of Tantoyuca, state of Veracruz, Mexico, located in the HAV region, between parallels 97°59' W and 98°24' W, and 21°06' N and 21°40' N, at an altitude of up to 300 masl. This region has a warm subhumid climate, with rainy summers, a mean annual temperature between 22 and 26 °C, an average annual rainfall between 1,100 and 1,300 mm, and a relative humidity of 44%[19]. According to FAO classification, the soil where the experiment was performed is classified as a pellic vertisol with clayey texture, a pH of 7.8, light salinity, 18.3 ppm N-NO3, 50 ppm phosphorus, 49 ppm potassium, and organic matter (> 5.1%), according to the analysis performed at the Soil Laboratory of the Veracruz University.
Evaluated germplasm and experiment design
-
The vegetal material used for this study belongs to the Cenchrus purpureus (Schumach.) Morrone species, and the evaluated accessions, taken as treatments, were King grass, Maralfalfa, Elephant grass, Merkeron, Purple grass, CT-115, and Taiwan, obtained from the 'La Posta' experimental field, belonging to the National Institute of Forestry, Agricultural and Livestock Research (INIFAP), in the town of Paso del Toro, Veracruz state, Mexico. A randomized block experimental design was used, with seven treatments and four replications. The assessment period ran from July 2019 to April 2020 under rainfed conditions.
Experiment site and planting
-
Twenty eight plots of 25 m2 (5 m × 5 m) were created in a 1,222 m2 area, distributed in four blocks (seven plots p/block), with a distance of 2 m between plots and blocks. Planting of the assessed accessions was done with two stems (phytomers with three internodes) per planting point, introducing two internodes in the seedbed, and placing the stems at a 45° angle. The distance between crop rows and sowing points was 80 cm, resulting in a plant density of 36 tillers per 25 m2 in every experimental unit. The seeded ground was only prepared by primary tillage (plowing and harrowing). Planting was done in September 2018, and the uniform cutting was made in July 2019, at 15 cm above the ground.
Sampling and Harvest Frequency (HF)
-
A sample area of 2 m2 (1 m × 2 m) was marked, leaving a 2 m border in each experimental unit to avoid the edge effect. Harvest frequency was determined based on a mean intercepted radiation rate of ≥ 95 in the four replications of each treatment. Four consecutive cuts were performed[20].
Evaluated variables
Fresh weight (FW) and dry matter (DM)
-
Cuts were done at 15 cm above the ground for FW assessment purposes, and all the biomass of the representative sample (2 m2) was weighed immediately after cutting, using a digital scale with a minimum precision of 1 g. Five whole plants (leaf and stem) were randomly selected to determine the DM and obtain a 300 g subsample, which was dried in a forced air lab oven at 65 °C to a constant weight[21]. The results were extrapolated to 10,000 m2 (ha).
Leaf: stem ratio (LSR) and accumulated dry leaf matter rate (ADLMR)
-
Five plants were selected from the biomass obtained from the sample, and the leaves were separated from the stem. Once separated, the stems were all placed in a forced air lab oven at 65 °C to a constant weight. The LSR was estimated by dividing the dry weight of the leaf samples by the dry weight of the stem samples. The ADLMR was calculated by dividing the dry leaf matter yield by the number of days between cutting periods. To estimate the FW, DM, LSR, and ADLMR variables per season (dry and rainy), the values of each variable per accession and season were added up. The annual values of all variables were obtained by adding up the data from the four cuts[22].
Height (H)
-
The H was determined using a measuring tape, calibrated in meters, centimeters, and millimeters, to measure five plants of each sample at every cutting period, from the soil level to the flag leaf.
Climate variables
-
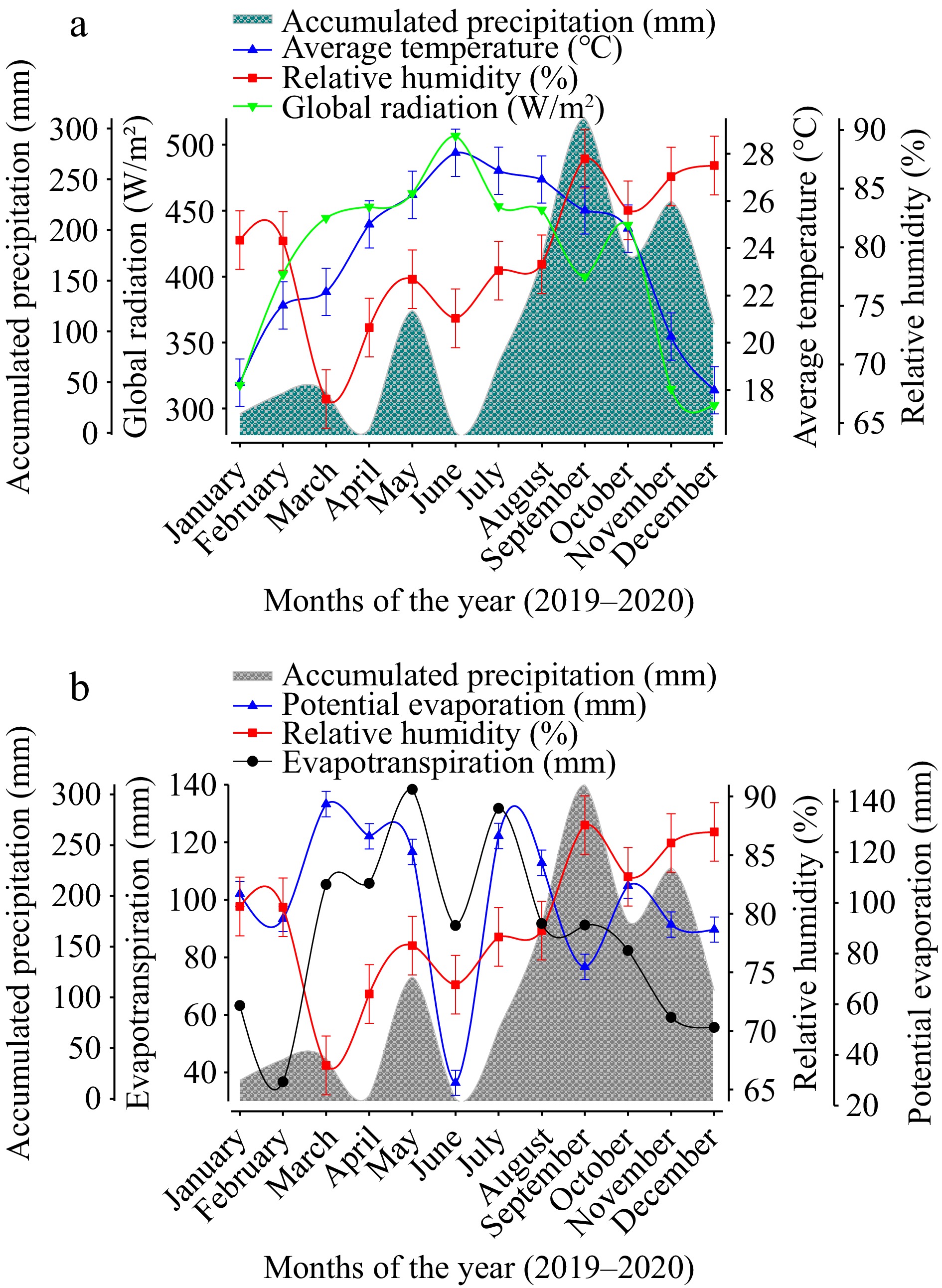
Some climate variables that are important for biomass production, such as average rainfall (mm), relative humidity (%), global radiation (W/m2), mean temperature (°C), and reference evapotranspiration (mm), were taken during the development of the research activities (Fig. 1). These data were obtained from the weather station located at the experimental field of the Instituto Tecnológico Superior de Tantoyuca. For the purposes of this research, 'dry season' is defined as the months of the year in which reference evapotranspiration is greater than the average rainfall because, in these conditions, crops are in a state of stress due to a lack of moisture, and this period is considered key in determining the need for irrigation[23].
Statistical analysis
Analysis of variance (ANOVA)
-
The Statistic software (STATISTICA V10, 2013) was used for statistical analysis; the general linear model (GLM) was as follows:
$\rm Y_{ijk}={\text μ} + {\text α}_i+{\text β}_j+{\text ε}_{ijk} $ Where: Yijk = the quantitative response variable of the i-th accession and the j-th replication. μ = Overall mean. αi = fixed effect of the i-th Cenchrus purpureus accession.
$\, {\text β}_{\rm j} $ Multivariate analysis
-
Using the studied response variables from each cutting performed, and the monthly average climatological variables of the research site, a multiple linear analysis was executed to ascertain the weight climatological variables have on the assessed accessions. The MLR model is as follows:
$ {\text γ}_{\iota }={\text β}_{0}+{\text β}_{1}{\rm X}_{1}+{\text β}_{2}{\rm X}_{2}+{\text β}_{3}{\rm X}_{3}+{\text β}_{4}{\rm X}_{4}+{\rm e}\mathfrak{i} $ Where:
$ {\text γ}_{{\iota }} $ ${\rm e}\mathfrak{i} $ Lastly, Principal Component Analyses (PCA) were used to observe the possible grouping of the agronomic and yield variables, as well as the accession grouping (STATISTICA V10, 2013).
-
Merkeron grass had the highest value for ADLMR, statistically different (p < 0.05) from Taiwan and King grass accessions, thus demonstrating its capability to accumulate dry matter in its leaves (Table 1). Consequently, this accession had an LSR of 0.88, the greatest of all the accessions, showing a statistical difference (p < 0.05) with the CT-115 accession, which had the lowest value (0.58). CT-115 showed the highest capacity for dry matter accumulation on its stems, which is made more evident by its being one of the tallest accessions, only shorter than King grass and Maralfalfa. An important characteristic, from the perspective of productivity, is harvest frequency (HF), that is, the time (d) it takes for the plant to reach maturity and harvest time. Merkeron, Elephant, and Purple grasses were harvested at intervals ~60 d, while the rest of the accessions at ~80 d (p < 0.05) (Table 1).
Table 1. Agronomic variables of seven Cenchrus purpureus accessions in the HAV in the rainy season.
Accession Variable ADLMR (kg/ha/d) LSR H (m) HF (d) Elephant 202.44ab 0.83a 1.85b 63.00b Merkeron 220.02a 0.88a 2.06ab 62.50b Purple 200.72ab 0.85a 1.90b 62.50b Taiwan 161.95b 0.82a 2.28a 83.00a King grass 154.07b 0.78a 2.16ab 83.00a CT-115 189.82ab 0.58b 2.05ab 83.00a Maralfalfa 170.04ab 0.70ab 2.15ab 83.00a s.e. 20.35 0.06 0.13 4.13 ADLMR: Accumulated dry leaf matter rate. LSR: Leaf:stem ratio. H: Height. HF: Harvest frequency. Means within columns followed by different letters differ by Fisher test at 5% probability. Elephant grass had the highest ADLMR, showing a statistical difference (p < 0.05) with Merker grass, Taiwan A-144, King grass and CT-115. In the present study, the highest LSR was observed in Purple grass (p < 0.05). However, despite this, Purple grass had the lowest H, which was notably different from that of Elephant grass and Taiwan. There was a slight variation in HF, with Elephant grass having the lowest value, similar to King grass, and CT-115 (p > 0.05). Merkeron grass had the highest HF of all accessions (p < 0.05) (Table 2).
Table 2. Agronomic variables of seven Cenchrus purpureus accessions in the HAV on the dry season.
Accession Variables ADLMR(kg/ha/d) LSR H (m) HF (d) Elephant grass 228.51a 0.93b 1.94a 79.00e Merkeron grass 108.84bc 0.91b 1.79ab 104.50a Purple grass 198.68a 1.22a 1.70b 98.00ab Taiwan 79.52c 0.92b 1.92a 94.00cb King grass 110.97bc 0.89b 1.84ab 85.50cde CT-115 91.02c 0.86b 1.72b 84.00de Maralfalfa 187.45ab 0.93b 1.86ab 92.50bcd s.e. 30.56 0.06 0.06 3.34 ADLMR: Accumulated dry leaf matter rate. LSR: Leaf:stem ratio. H: Height. HF: Harvest frequency. Means within columns followed by different letters differ by Fisher test at 5% probability. The ADLMR variable in Elephant grass, which had a value of 228.51 kg/ha/d, was the highest in all treatments, showing statistical differences (p < 0.05) with Merkeron grass, Taiwan, King grass, and CT-115, and a greater capability to deposit dry matter on the leaves (Table 3). Even though Elephant grass showed an adequate LRS, it was surpassed by Purple grass, which was statistically different (p < 0.05) from King grass, CT-115, and Maralfalfa. Measure H was not statistically different (p > 0.05) among the assessed accessions. The shortest HF was 71 d, and it corresponded to Elephant grass, while the longest corresponded to the Taiwan accession, with statistical differences between the two (p < 0.05).
Table 3. Annual analysis of agronomic variables in seven Cenchrus purpureus accessions in the HAV region.
Accession Variables ADLMR (kg/ha/d) LSR H (m) HF (d) Elephant grass 215.47a 0.88ab 1.89a 71.0b Merkeron grass 164.42ab 0.89ab 1.92a 83.5ab Purple grass 199.70ab 1.03a 1.80a 80.2ab Taiwan 120.73b 0.87ab 2.09a 88.5a King grass 132.51b 0.83b 2.00a 84.2ab CT-115 140.42ab 0.72b 1.88a 83.5ab Maralfalfa 178.74ab 0.81b 2.00a 83.5ab Standard error 19.08 0.04 0.07 3.2 Season Rainy 185.58a 0.78b 2.06a 74.2b Dry 143.56b 0.95a 1.82b 91.0a s.e. 10.20 0.02 0.37 1.7 ADLMR: Accumulated dry leaf matter rate. LSR: Leaf:stem ratio. H: Height. HF: Harvest frequency. Means within columns followed by different letters differ by Fisher test at 5% probability. The differences in climatic factors between the dry and rainy seasons (Fig. 2) influenced the agronomic behavior of the assessed accessions. Noteworthy values of ADLMR, H, and HF were reported during the rainy season. On the other hand, LSR was greatest in the dry season (p < 0.05).
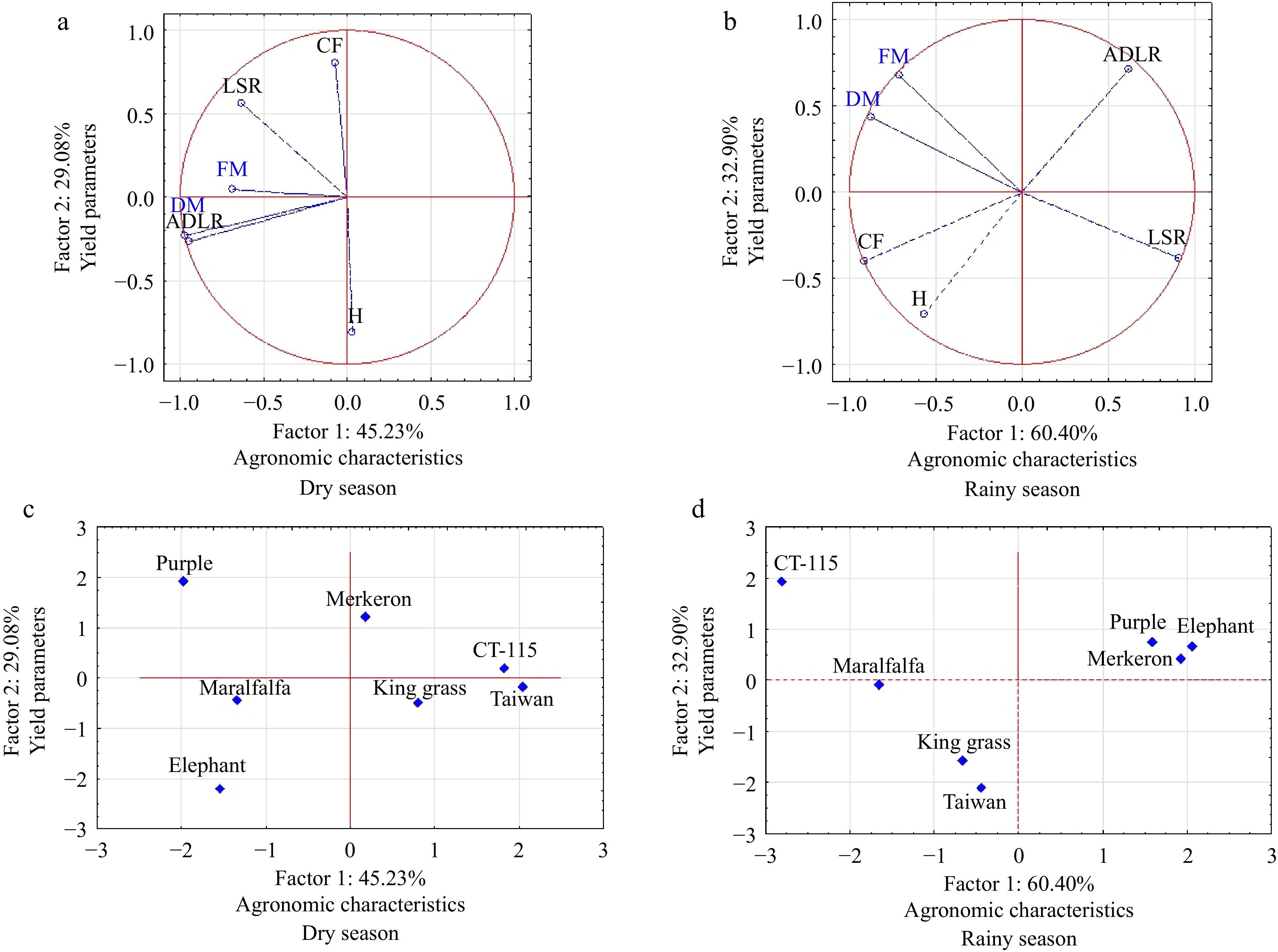
Figure 2.
Grouping of Cenchrus purpureus accessions according to two main components: Factor 1. Agronomic characteristics and Factor 2. Yield parameters. (a), (c) dry season, (b), (d) rainy season.
The FW of the assessed accessions was not statistically different during the rainy season; nevertheless, the DM results indicate that CT-115 was quantitatively superior to all the other accessions, and statistically different (p < 0.05) from Elephant and Merkeron grass. Conversely, the FW of Maralfalfa was greatest in the dry season, and it was statistically different from CT-115 and Taiwan (p < 0.05). In turn, Elephant grass had the highest accumulation of dry matter (t/ha), followed by Purple grass and Maralfalfa (p < 0.05) in the same period. The annual cumulative fresh weight was greatest in Maralfalfa, and statistically different (p < 0.05) from Taiwan, while Elephant grass had the greatest amount of accumulated dry matter of all the accessions after a year of assessment (Table 4).
Table 4. Cumulative yields of fresh weight and dry matter per season and cumulative annual yield of seven accessions of Cenchrus purpureus in the HAV region.
Accession Rainy season Dry season Annual FW (t/ha) DM (t/ha) FW (t/ha) DM (t/ha) FW (t/ha) DM (t/ha) DM (%) Elephant grass 261.31a 54.39a 155.18abc 80.09b 416.50ab 134.49a 34.1a Merkeron grass 269.78a 55.60a 179.11a 46.71a 448.89ab 102.31ab 24.2b Purple grass 264.66a 61.65ab 153.46ab 69.30b 418.13ab 130.95a 32.7a Taiwan 252.66a 60.05ab 104.96c 29.21a 357.62a 89.27b 27.5ab King grass 259.79a 61.32ab 156.66ab 41.11a 416.46ab 102.44ab 26.3ab CT-115 321.14a 79.81b 120.01bc 32.20a 441.16ab 112.01ab 27.2ab Maralfalfa 295.95a 67.03ab 190.57a 67.37b 486.52b 134.41a 30.0ab s.e. 25.08 7.22 13.77 6.05 32.62 11.23 2.84 FW: Fresh weight. DM: Dry matter. Means within columns followed by different letters differ by Fisher test at 5% probability. Table 5 shows the average FW and DM per cutting period. The superior capability of Elephant grass to produce dry matter (DM) under the particular climatic conditions of the HAV is evident. Statistical differences exist in FW and DM variables between seasons (p < 0.05). The rainy season yielded the highest values for these variables.
Table 5. Average annual yields per cut of fresh weight and dry matter of seven Cenchrus purpureus accessions in the HAV region.
Accession Variable FW (t/ha) DM (t/ha) Elephant grass 208.25ab 67.24a Merkeron grass 224.44a 51.15ab Purple grass 209.06ab 65.47a Taiwan 178.81b 44.63b King grass 208.23ab 51.22ab CT-115 220.58ab 56.00ab Maralfalfa 243.26a 67.20a s.e. 15.01 6.19 Season Rainy 275.04a 62.84a Dry 151.42b 52.28b s.e. 8.02 3.31 FW: Fresh weight. DM: Dry matter. Means within columns followed by different letters differ by Fisher test at 5% probability. The correlation analysis used to determine the degree of association between the DM and other agronomic variables indicated positive correlations with H, ADLMR, and a negative correlation with LSR. Nevertheless, it was found that there is little association with the FW variable (r = 0.147), showing a value of p= 0.083. For the case of the H and LSR variables, the associations were low (r = 0.192 and r = 0.262, respectively). There was a high degree of association with the ADLMR variable (r = 0.889), with a value of p = 0.001 (Table 6).
Table 6. Correlation analysis between the DM and the agronomic variables of seven Cenchrus purpureus accessions grown in the HAV.
Variable r r2 α s.e. (a) p (a) β s.e. (b) p (b) DM H 0.192 0.037 12.826 6.743 0.059 8.218 3.580 0.023 LSR 0.262 0.069 44.658 5.333 0.001 -19.545 6.116 0.002 DLMAR 0.884 0.782 3.967 1.227 0.002 0.153 0.007 0.001 FW 0.147 0.022 26.466 1.525 0.001 0.000 0.001 0.083 ADLMR: Accumulated dry leaf matter rate. LSR: Leaf:stem ratio. H: Height. FW: Fresh weight. r2: Determination coefficient. a: intercept. b: slope. p: p-value. MLR was implemented for the creation of statistical models to describe the magnitude of the association between climatological and agronomic variables of Cenchrus purpureus in the HAV is presented in Table 7. From all the performed analysis, the one that obtained a medium to high value in the goodness-of-fit indicators (r = 0.73 and r2adjusted = 0.52) was the FW variable, where the Forward method included all the climatic variables. The goodness-of-fit indicators for the DM mathematical model (r = 0.22 and r2adjusted = 0.03) do not indicate a good fit of the data, and likewise with the LSR and ADLMR. The MLR model (Forward method) for the H variable showed a medium-high degree of association (r = 0.70 and r2adjusted = 0.48), only including temperature and global radiation.
Table 7. Multiple linear regression models of the agronomic variables of seven Cenchrus purpureus accessions, associated with climatological variables of the HAV.
Variable Y = α + β1 + β2 + β3 + β4 + c r r2adjusted p MSE Method FW −27.2 0.10 −33.01 2.12 3.15 34.68 0.73 0.52 0.01 1202.92 F p value 0.84 0.10 0.01 0.01 0.01 DM 11.08 − −1.50 0.15 − 14.33 0.22 0.03 0.02 205.45 F p value 0.50 − 0.29 0.06 − H 3.64 0.00 −0.28 0.01 − 0.24 0.70 0.48 0.01 0.06 F p value 0.01 0.17 0.01 0.01 − LSR 2.90 − − −0.01 −0.01 0.16 0.53 0.28 0.01 0.02 B p value 0.01 − − 0.01 0.01 ADLMR 80.19 − −16.99 1.45 − 80.36 0.32 0.08 0.01 6458.51 F p value 0.39 − 0.036 0.01 − Y model = α + β1X1 + β2X2 + β3X3 + β4X4 + c; α: Intercept; β1−4: Slope factor; X1: Accumulated precipitation (mm). X2: Average temperature (°C). X3: Global radiation (W/m2). X4: Relative humidity (%). c: Estimate error. FW: Fresh weight. DM: Dry matter. r2adjusted: Adjusted correlation coefficient. p: Probability value. MSE: Mean Square Error. F: forward method. B: backward method. (−): Variable not included in the model. The principal component analysis (PCA) was conducted using all estimated variables (rainy and dry seasons) of the seven Cenchrus purpureus accessions, resulting in the creation of two groups of orthogonally independent variables. The first group (Factor 1) corresponds to those known as agronomic characteristics, and the second group (Factor 2) is made up of estimated yield variables (Fig. 2).
The PCA implemented in the dry and rainy seasons (Fig. 2) formed three groups of Cenchrus purpureus accessions: a) outstanding agronomic characteristics and fresh weight and dry matter, b) outstanding yield characteristics, and c) unremarkable agronomic and yield characteristics. The Cenchrus purpureus accessions integrating each group were different in both seasons. Their genetic characteristics and interactions with the environment define their performance.
-
The ADLMR of the Taiwan accession was greater than reported by Rueda et al.[24] in the south region of Veracruz state during the rainy season. Despite the application of fertilizers, the values that they obtained were lower than those reported in this study. The planting density was not the same, thus the greater accumulation of DM/ha/d may be attributed to the longer HF, and to a greater acidity of the soil where the experiment was carried out, which negatively affected the yield. The ADLMR statistics were led by Elephant grass, Purple grass and Maralfalfa (Table 3) these results were higher than those reported by Uvidia-Cabadina et al.[25], who reported that Maralfalfa had an average growth rate of 113 ka/ha/d at 75 days of age.
The LSR results of the evaluated accessions showed greater values during the dry season, in comparison with the rainy season. Purple grass stood out from the rest in this variable, showing a value of 1.03 (with a harvest frequency of 80 d). This does not coincide with the results reported by Rueda et al.[24], who evaluated this parameter at 185 days of regrowth; however, they found that LSR values decreased in the cultivars exposed to irrigation with nitrogen fertilization, showing that the stem proportions in this grass increased with age. The LSR decreased due to the increase in stem proportions, mainly in the rainy season, which favors apical growth. This was also described by Habte et al.[22] upon evaluating 45 genotypes of Cenchrus purpureus in the dry and rainy seasons of Bishoftu, Ethiopia, at 1,890 masl. However, it has been shown[26] that planting density plays an important role on the LSR of Pennisetum purpureus cultivars, in which values of 1.4 were obtained with 40,000 plants/ha. For this research, plant density per hectare in the research units was established at 15,625, which could partially explain the lower values found in the assessed accessions. Increased stem proportions in this genus reduces the quality of the fodder in terms of their neutral detergent fiber content, as indicated by Maldonado-Mendez et al.[27].
In the case of King grass harvested at 80 d, the LSR was 0.78 and 0.89 in the rainy and dry season, respectively (Tables 1 & 2). The results seem to be related to an increased availability of water, which differs from the reports of Rueda et al.[24], who observed an LSR value of 0.26 in the King grass accession after 185 d of regrowth, with the use of fertilizer at various doses and concentrations. However, when harvested after 60 d in that study, the value of this parameter oscillated between 0.66 and 0.70, which led to the conclusion that there is a negative correlation between the LSR and yield, contrasting with the results of this study.
In the present study, King grass was one of the accessions with the highest H value per harvest frequency (2.00 m). This result differs from the reports by Rueda et al.[24], who, upon application of fertilizers at several concentrations (a formula 200, 100, 100 kg/ha/year, N-P-K), managed to increase this agronomic characteristic. The heights obtained from accessions that they assessed at 125 d were generally higher than those reported in this study, which suggests a favorable response to fertilizer. Upon evaluating eight varieties of Cenchrus purpureus, including CT-115, Ruiz et al.[28], reported that the height values did not surpass those in this research. The results are similar to the results of our study, where the height of the assessed accessions differs among them (p > 0.05). Plant height in different C. purpureus genotypes partially depend on regrowth days; thus, the older the plant is, the taller it is[29], which could explain the observed difference.
Accessions with the greatest average FWs per cutting period were Maralfalfa and CT-115 (243 and 220 t/ha, respectively), at HF of 83 d. These FW results are higher than those presented by Maldonado-Mendez et al.[27], who assessed Maralfafa accession in a humid tropical region of Chiapas, Mexico. They established six cutting periods, ranging from 28 to 97 d, and coincided in that Maralfalfa is capable of accumulating fresh weight as days pass, but its nutritional value decreases over time. Arias et al.[29], reported that CT-115 assessed under irrigation conditions with organic fertilizer showed a fresh weight lower than those presented here when harvested at 150 d of regrowth. The region's low precipitation level could explain this variable's lower values. Measured yields were lower in the dry season, showing significance in them (p < 0.05). This same effect was reported by Habte et al.[22], who associated the high FW and DM with the months of highest rainfall. Moreover, they found no significant differences between the genotypes they assessed, unlike the results reported here (Table 5).
DM is a desirable attribute in fodder crops because it contains nutrients that could be beneficial to ruminants. Results presented here showed that some accessions were more capable of depositing dry matter, and the variation in climatological variables between the rainy and dry seasons did not affect the behavior of this indicator to any considerable degree, such as for Elephant grass, Purple grass, and Maralfalfa (Table 4). However, the climatic conditions of the rainy and dry season affected DM values of the rest of the assessed accessions (p < 0.05) This indicates that these genotypes are susceptible to climate fluctuations, as demonstrated in an experiment by Uvidia-Cabadina et al.[25] in the Ecuadorian Amazon region. They evaluated the Maralfalfa accession at different ages and climatic conditions, and found a correlation between dry matter, rainfall, outdoor temperature, and relative humidity. Taiwan showed greater DM in the rainy season, which duplicated the amount shown in the dry season. Maldonado-Mendez et al.[27] evaluated the effect of age on the DM of the Maralfalfa cultivar during the rainy season in Chiapas, Mexico, and concluded that age helps to duplicate this variable; however, their yields do not surpass those reported in this study, possibly due to the shortened HF and the characteristics of the soil. The photosynthetic and DM accumulation capabilities show variation within the Cenchrus genus, the higher yields of some accessions during the seasons of most abundant rainfall is due to their intrinsic potential and the higher amount of nutrients dissolved in the ground because of the greater humidity, as illustrated by Habte et al.[22], upon evaluation of the DM and FW of several genotypes of this genus in similar conditions.
Among the assessed accessions, DM per cut, per year, was greatest in Purple grass and Maralfalfa (Table 5). In this respect, it has been mentioned that these cultivars showed the highest DM when compared with King grass, CT-115, and Elephant grass, and highlighted that fertilization and longer HF were factors that increased the accumulation of DM[25,29,30] in all the assessed genotypes. This explains the greater DM found in this study.
Most of the evaluated agronomic variables showed a low association degree, with DM and FW showing a positive degree, although still low and not significant (Table 6). Therefore, it is possible to infer that the amount of water accumulated in the stems differs among the assessed accessions, which occupies a greater proportion of the plant (Table 3). In contrast, Habte et al.[22] found high positive correlations (r = 0.86) between the total amounts of DM and FW accumulated in several Cenchrus purpureus cultivars. Seemingly, the higher proportion of leaves (LSR 2.41 and 5.87 in the rainy and dry seasons, respectively) boosted this correlation. Moreover, these authors coincide in finding a correlation between DM and LSR (negative correlation), indicating that the higher the LSR, the higher the total proportion of leaves in the plant and, consequently, the amount of DM represented by the stem is lower.
The data that best fits the MLR model corresponds to FW variables, which is evidenced by their goodness indicators (Table 7); Araujo et al.[30] report similar values in these indicators for several species of C4 grass crops. Pannicum maximum (Tanzania and Mombasa), a grass species with an erect growth habit, like that of the Cenchrus genus, reached r2 values between 0.78 and 0.87 for the ADLMR and ambient temperature variables in the univariate linear regression analysis. The step-by-step execution with the Forward method used in this investigation for the multivariate regression model included all the used climatological variables. From these results, it can be observed that temperature has a negative angular value on biomass production. Andrade et al.[31] found that temperature was related to enzymatic physiological processes, such as photosynthesis, and other processes related to transport through cellular membranes. The production behavior of the assessed accessions may have been affected by the extreme climatological conditions under which this research was done (Fig. 1).
-
The assessed accessions showed variations in their agronomic behavior and yield indicators, attributed to changes in the climatic variables between the rainy and dry seasons, and to genetic variability. Elephant grass and Maralfalfa achieved the highest DM and FW in both seasons, even though there were minimal differences between them, proving their genetic capability to maintain a stable production in different environmental conditions. It is necessary to simultaneously evaluate the agronomic characteristics, yield, and quality indicators of the assessed accessions to identify those who show the best levels in these attributes.
The first author of the manuscript (Armando Arrieta-González) received a scholarship for Doctoral studies from the National Council of Humanities, Science and Technology (CONAHCyT). We thank the 'La Posta' experimental field belonging to the National Institute of Forestry, Agricultural and Livestock Research (INIFAP) for the accessions of Cenchrus purpureus.
-
The authors confirm contribution to the paper as follows: study conception and design: Arrieta-González A, Silva-Martínez KL, Domínguez-Mancera B; data collection: Arrieta-González A, Silva-Martínez KL, Vite-Cristóbal C; analysis and interpretation of results: Arrieta-González A, Silva-Martínez KL, Hernández-Beltrán A, Domínguez-Mancera B; draft manuscript preparation: Arrieta-González A, Silva-Martínez KL, Rodríguez-Andrade A, Domínguez-Mancera B. All authors reviewed the results and approved the final version of the manuscript.
-
The datasets generated during and/or analyzed during the current study are available from the corresponding author Belisario Domínguez-Mancera on reasonable request.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Arrieta-González A, Silva-Martínez KL, Vite-Cristóbal C, Rodríguez-Andrade A, Hernández-Beltrán A, et al. 2024. Agronomic traits of seven accessions of Cenchrus purpureus under rainfed conditions in the tropical region of Veracruz, México. Grass Research 4: e023 doi: 10.48130/grares-0024-0022
Agronomic traits of seven accessions of Cenchrus purpureus under rainfed conditions in the tropical region of Veracruz, México
- Received: 02 July 2024
- Revised: 30 October 2024
- Accepted: 01 November 2024
- Published online: 28 November 2024
Abstract: The purpose of this study was to determine the agronomic characteristics and yield of seven Cenchrus purpureus accessions (Taiwan grass, King grass, Elephant grass, Merkeron, CT-115, Purple grass, and Maralfalfa) to identify the best options to use as fodder resources in the Huasteca Alta Veracruzana (HAV) ruminant production units. The research was conducted at the Instituto Tecnologico Superior de Tantoyuca in 2019 and 2020, under rainfed conditions. Sowing density was established at 36 tillers per 25 m2, with a spacing between rows of 0.8 m and 0.8 m between plants. Fresh weight, dry matter, daily rates of dry matter accumulation in leaves, leaf/stem ratio, height, and harvest frequency were assessed. Rainfall amount, temperature, relative humidity, and global radiation levels were recorded to assess their effect on the evaluated variables. The one-way analysis of variance and multivariate analysis techniques were used, such as multiple linear regression and clustering analysis with Statistica V10 software. There were no statistical differences between the fresh weight of different accessions in the rainy season. On the other hand, CT-115 presented a higher dry matter content than Merkeron or Elephant grass (p < 0.05). The correlation analysis showed a high association between dry matter and daily rates of dry matter accumulation in leaves. The results varied due to weather phenomena and C. purpureus genetic variability. Quality parameters must be evaluated to identify the accessions with the best nutritional values and their effect on animal performance.
-
Key words:
- Mathematical models /
- Fodder /
- Yield /
- Drought /
- Tropic