-
The intestinal mucosal barrier is composed of epithelial cells and intercellular connections, which can effectively regulate the transportation of large molecules in the intestinal lumen, and prevent microorganisms, harmful solutes, toxins, and intraluminal antigens from entering bodies[1]. Mucosal barrier factors such as trefoil factor (TFF) family, diamine oxidase (DAO), and transform growth factor-ɑ (TGF-α) have protective and restorative effects on intestinal mucosal integrity, which are synthesized and secreted by intestinal mucosa[2−4]. The damage to intestinal barrier function can cause the invasion of antigens and bacteria in the lumen, and eventually lead to intestinal diseases, including diarrhea, inflammatory bowel disease (IBD), and Crohn's disease[5−7].
The Notch signaling pathway plays a crucial role in a series of cellular processes, including proliferation, differentiation, development, migration, and apoptosis. Research suggests that the Notch signaling pathway is involved in intestinal development, and it can be connected to intestinal cell lineage specification[8]. Furthermore, the Notch signaling pathway can regulate intestinal stem cells, CD4+T cells, innate lymphoid cells, macrophages, and intestinal microbiota, and intervene in the intestinal mucosal barriers in cases of ulcerative colitis[9]. The activated Notch signaling pathway can suppress the goblet cells differentiation and mucus secretion, which would result in the disruption of the intestinal mucosal barrier[10]. In addition, the absence of the Notch signaling pathway would cause the dysfunction of tight junctions and adherens junctions of intestinal mucosa, which could lead to increased permeability of epithelial cells and exposure of luminal contents to the immune system and inflammation[11]. Phytochemicals, such as cucurbitacin, honokiol and quercetin have been reported to have therapeutic effects on intestinal diseases by targeting the Notch signaling pathway[12−14].
Plant-based diets rich in phytochemicals, such as phenolics, anthocyanins, and vitamins, have been related to the prevention of human diseases[15]. Anthocyanins belong to a subclass of flavonoids, and are mainly in the form of different anthocyanin combined with glucose, galactose, and arabinose[16]. The physiological action of anthocyanins, such as antioxidant activity, anti-inflammation, and anti-obesity effects have been widely reported[17,18]. M3G is found naturally in plants and is reported to be the most common anthocyanin in different blueberry varieties[19,20]. Previous researchers reported that M3G from blueberry suppressed the growth and metastasis potential of hepatocellular carcinoma cells, modulated gut microbial dysbiosis, and protected TNF-α induced inflammatory response injury in vascular endothelial cells[21−23]. Anthocyanins with diverse molecular structures and from different dietary sources are bioavailable at diet-relevant dosage rates, and anthocyanins from berry fruit are absorbed and excreted by both humans and rats[24]. Studies have shown that anthocyanins have a protective effect on intestinal barrier damage by regulating gut microbiota, tight junction (TJ) protein expression, and secretion of MUC2[25−27]. In addition to physiological indicators related to the intestinal barrier, the regulation of the Notch signaling pathway was involved in this study. Therefore, it is speculated that M3G could improve the colonic mucosal barrier function via the Notch signaling pathway.
To prove this hypothesis, the effects of M3G on the colonic mucosal barrier function were investigated in DSS-induced colitis mice. The pathological morphology of the colon tissue, markers of intestinal physical barrier function and immune barrier function and the Notch signaling pathway were evaluated to reveal the mechanism of M3G in improving colonic mucosal barrier function. The results are expected to lay the foundation for the utilization of anthocyanins as a promising natural product for improving intestinal diseases.
-
M3G (CAS: 30113-37-2) used in this study was purchased from Xinyi Science and Technology Instrument Business Department, Baoji, Shanxi, China (HPLC ≥ 98%).
Animal experiments
-
Five-week-old male C57BL/6J mice (Wanlei Bio Co., Ltd., Shenyang, Liaoning, China) weighing 16−18 g were housed and given AIN-93M diet (Wanlei Bio Co., Ltd., Shenyang, Liaoning, China) feeding. The animal experiment was carried out according to the guidelines of the Standards for Laboratory Animals of China (GB 14922-94, GB 14923-94, and GB/T 14925-94) and the Ethics Committee of Shenyang Agricultural University (IACUC Issue No.: 2023022401). All animal housing and experiments were conducted in strict accordance with the institutional guidelines for the care and use of laboratory animals.
After acclimating to the breeding environment for one week, the mice were randomly divided into two groups: control group (CG group, n = 6) and model group (n = 12). On days 1−7, the mice of the model group were given 2.5% DSS (CAS: 9011-18-1) dissolved in drinking water, while the mice of the CG group were given the same volume of drinking water as the model group. On days 8−14, mice of the model group were randomly divided into two equal groups (n = 6): DSS group and M3G group. The mice of the CG and DSS groups were administered intragastrically via drinking water, and the mice of the M3G group were administered intragastrically by M3G (5 mg/kg body weight (BW)/d) dissolved in drinking water, and the liquid volume was controlled to be the same. For all groups of mice, the body weight, food consumption, stool consistency, and bloody stool were measured daily during the experiment. On day 15, mice were sacrificed after a 12 h fast, and the colon tissue samples of the mice were collected. The length of the colon tissue was measured and recorded.
Histopathological analysis
Hematoxylin-eosin (HE) staining
-
Colon tissue was embedded with paraffin and then cut into sections. After being dewaxed from paraffin, the tissue was placed in water, and stained with hematoxylin and eosin solution. The stained tissue was dehydrated and sealed for observation. A microscope (BX53, Olympus Co., Ltd., Tokyo, Japan) was used to observe the stained tissue and photographed using 100× magnification. The damage in epithelial cells of colon tissue was evaluated.
Periodic acid-schiff (PAS) staining
-
The colon tissue was dewaxed and placed in water after being embedded with paraffin, and then stained with schiff and hematoxylin solution. The stained tissue was dehydrated and sealed for observation. A microscope was used to observe the stained tissue and photographed using 100× magnification. The mucosal thickness and the population of goblet cells of the colon tissue were measured and recorded.
Real-time polymerase chain reaction (RT-PCR)
-
The expression of MUC2 in colon tissue was detected by RT-PCR. Total RNA was extracted from colon tissue, and the concentration was determined using an ultraviolet spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The single-stranded cDNA of the extracted RNA (0.1 μL) was synthesized using a Transcriptor First Strand cDNA Synthesis Kit (Roche Co., Ltd., Basel, Kanton Basel, Switzerland). ExicylerTM 96 (Bioneer Corporation, Daejeon, Korea) was used to analyze the fluorescence quantitative cDNA. The reaction conditions were as follows: 94 °C for 5 min, 94 °C for 10 s, 60 °C for 20 s, 72 °C for 30 s, then followed by 40 cycles of 72 °C for 2 min 30 s, 40 °C for 1 min 30 s, and then melting from 60 to 94 °C, and incubating at 25 °C for 1−2 min. The primer sequences are shown in Table 1.
Table 1. The primer sequences used in the RT-PCR analysis.
Gene Prime Sequence (5'-3') Size (bp) MUC2 Forward TGTGCCTGGCTCTAATA 17 Reverse AGGTGGGTTCTTCTTCA 17 β-actin Forward CTGTGCCCATCTACGAGGGCTAT 23 Reverse TTTGATGTCACGCACGATTTCC 22 Western blot analysis
-
The whole proteins from the colon tissue (200 mg) were extracted using a Whole Cell Lysis Assay kit (BioTeke Co., Ltd., Beijing, China) according to the manufacturer’s protocol. Briefly, colon tissue was cut into pieces, and then mixed with phenylmethylsulfonyl fluorid (PMSF), followed by adding the protein extraction reagents A and B to prepare the tissue homogenate. Protein concentration was then determined using a Bradford Kit (BioTeke Co., Ltd., Beijing, China) according to the manufacturer’s protocol. Proteins were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). The membranes were blocked with 5% non-fat dry milk in TBST buffer for 1 h, followed by incubating overnight at 4 °C with the appropriate monoclonal primary antibody (detailed information in Table 2). Then membranes were washed to remove non-bound antibodies, and then incubated with the secondary antibody (goat anti-rabbit immunoglobin G-horseradish peroxidase (IgG-HRP), 1:5000; Wanlei Bio Co., Ltd., Shenyang, Liaoning, China) at 37 °C for 45 min. The enhanced chemiluminescence (ECL) western blotting detection reagent (Wanlei Bio Co., Ltd., Shenyang, Liaoning, China) was used to detect the protein bands, and then the protein bands were visualized by a Gel Imaging System (Beijing Liuyi Biotechnology Co., Ltd., Beijing, China).
Table 2. Details of the primary antibodies used in the experiment.
Primary antibody Dilution ratio Manufacturer Claudin-1 1:500 Wanlei Bio Co., Ltd. Occludin 1:500 Wanlei Bio Co., Ltd. ZO-1 1:500 Wanlei Bio Co., Ltd. iFABP 1:1000 ABclonal Technology Co., Ltd. DLL1 1:500 Wanlei Bio Co., Ltd. DLL4 1:1000 ABclonal Technology Co., Ltd. Notch1 1:500 Wanlei Bio Co., Ltd. NICD 1:1000 Affinity Biosciences Co., Ltd. Hes1 1:1000 ABclonal Technology Co., Ltd. TFF3 1:1000 Affinity Biosciences Co., Ltd. β-actin 1:1000 Wanlei Bio Co., Ltd. Enzyme-linked immunosorbent assay (ELISA)
-
The level of SIgA in colon tissue was detected by SIgA ELISA kit (Model number: EM1362, Wuhan Fine Biotech Co., Ltd., Wuhan, Hubei, China). The experiment was carried out according to the ELISA kit instructions.
Flow cytometry (FCM)
-
CD4+T (CD3+CD4+) cells and CD8+T (CD3+CD8+) cells in colonic lamina propria monocytes (LPMC) were detected by FCM. The colonic epithelial cells were isolated by adding digestive solution to the colon tissue. After digestion, screening, re-suspension precipitation, and centrifugation, the precipitate was collected as colonic LPMC. Labeled antibodies were added to the flow tube and incubated according to the instructions. After washing with phosphate buffered saline (PBS), the cells were re-suspended in PBS solution. The CD4+T (CD3+CD4+) cells and CD8+T (CD3+CD8+) cells were detected by flow cytometric (Agilent Technologies, Inc., Santa Clara, CA, USA).
Statistical analysis
-
Data are expressed as mean ± standard deviation (SD) based on six replicates. Differences between two groups were assessed using Student's t tests. The results were considered statistically significant at p < 0.05. Data were analyzed using Graph Pad Prism 8.0 (Graph Pad Software, San Diego, CA, USA) and SPSS 17.0 (IBM Corporation, Armonk, NY, USA).
-
The body weight of mice was monitored daily during the experimental period. As shown in Fig. 1a, after being fed with DSS for 7 d, the body weight of the mice in the DSS group was reduced significantly (p < 0.01) compared with that of the CG group. While the body weight of the mice in the M3G group was increased, and exhibited significantly higher (p < 0.01) body weight gain compared with the DSS group. The DAI score of the mice in the DSS group was significantly higher (p < 0.01) than that of the CG group, but M3G supplementation significantly lowered (p < 0.01) the DAI score (Fig. 1b), which demonstrates that the DSS-induced mice colitis model was established successfully and M3G inhibited colon tissue damage in DSS-fed mice. Compared with the CG group, the food intake of the mice fed with DSS was significantly reduced (p < 0.01), and M3G supplementation significantly increased (p < 0.05) the food intake (Fig. 1c).
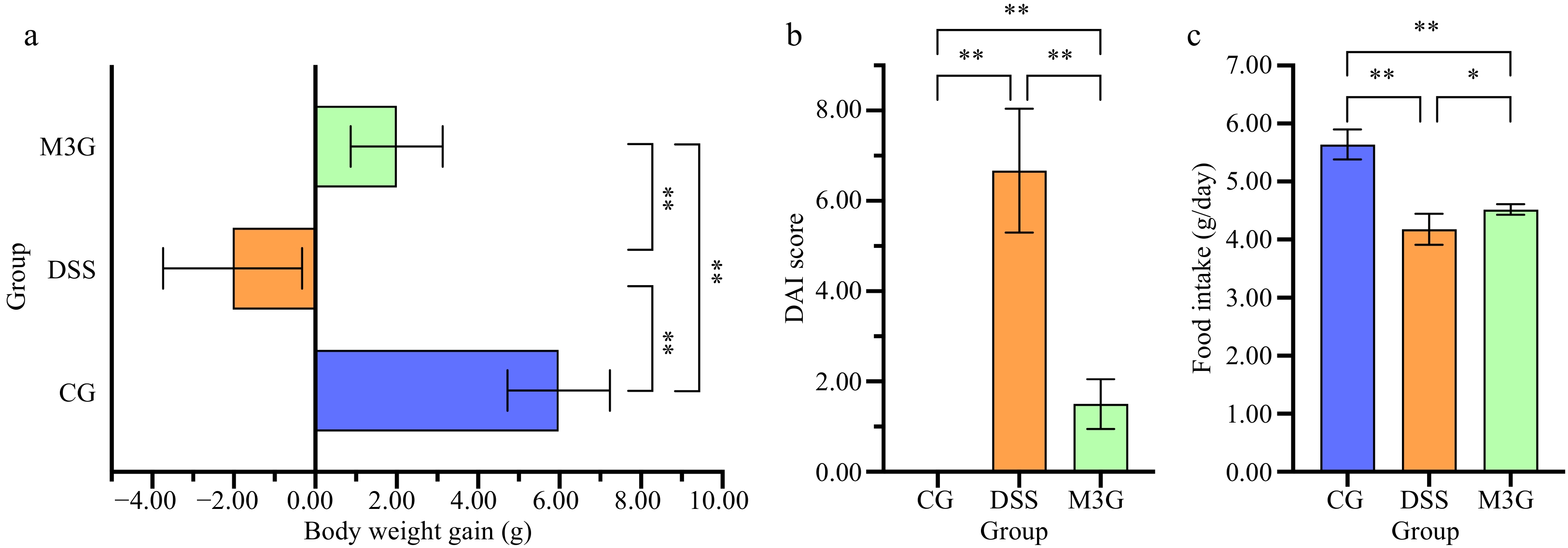
Figure 1.
Effect of M3G on body weight gain, DAI score, and food intake in mice. (a) Body weight gain. (b) DAI score (Fecal consistency: 0 = normal, 1 = semi-formed, 2 = soft stool, 3 = diarrhea or watery stool; bloody stool: 0 = occult blood negative, 1 = occult blood positive, 2 = visible bloody stool, 3 = massive hemorrhage; weight loss: 0 = 0%, 1 = 1%−5 %, 2 = 6%−10% ; 3 = 11%−15% reduction). (c) Food intake. Results are expressed as the mean ± SD (n = 6). * p < 0.05 and ** p < 0.01 indicate significant differences between two groups.
Effects of M3G on the colonic mucosal barrier function of mice with DSS-induced colitis
Pathological morphology changes of the colon tissue
-
Representative HE and PAS-stained sections of the colon are shown in Fig. 2a & b. The morphology of colon tissue in the CG group did not exhibit any damage, while damage in epithelial cells and goblet cells, inflammatory cells infiltration, and separation of the muscle layer and mucosal muscle layer were observed in the DSS group. The above damage to the tissue was changed for the better by M3G supplementation, with only small levels of damage in epithelial cells and goblet cells, inflammatory cell infiltration, and separation of the muscle layer and mucosal muscle layer compared to that in the CG group.
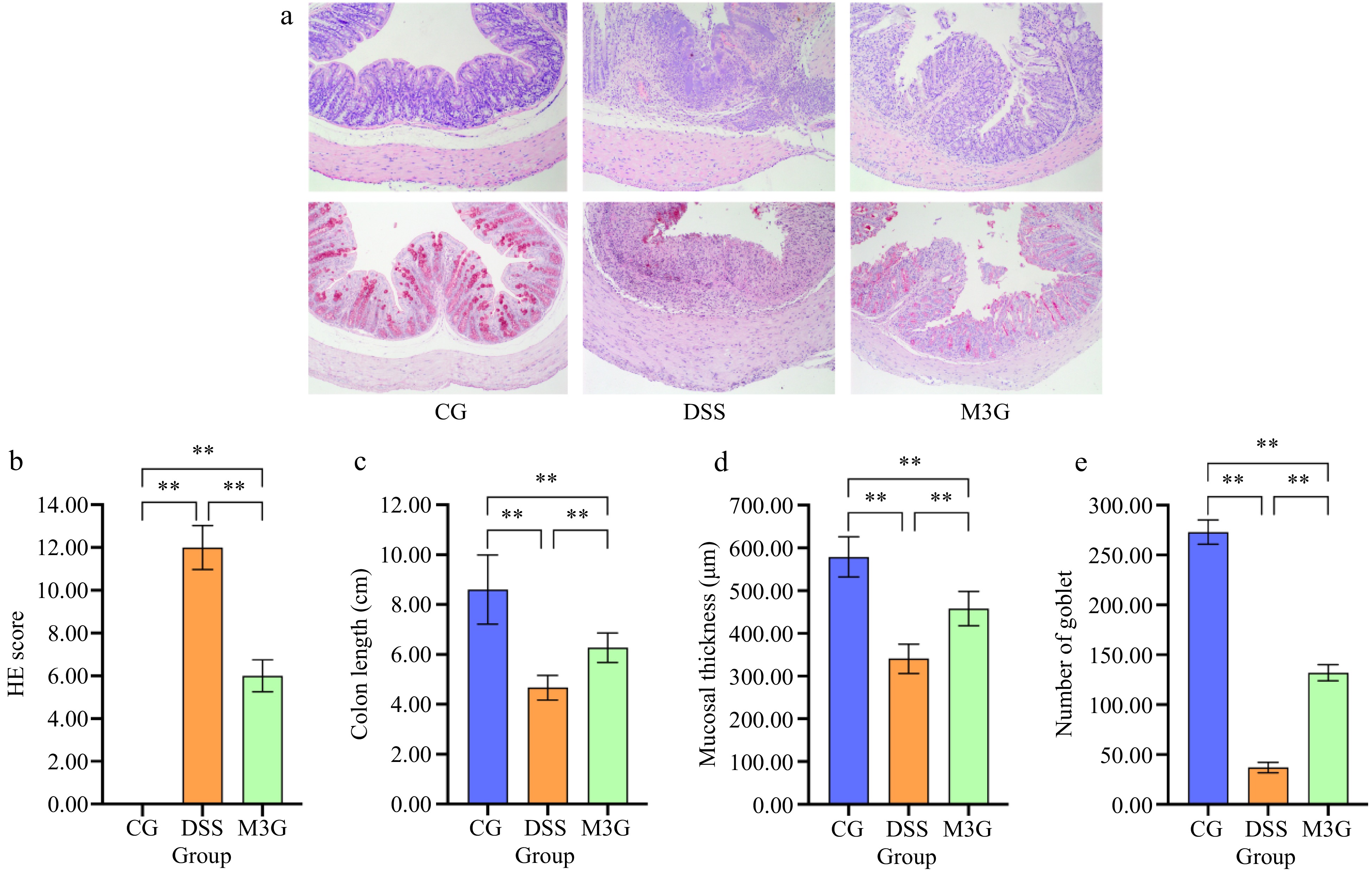
Figure 2.
Effect of M3G on the pathological morphology of colon tissue. (a) HE staining of colon tissue. Original magnifications: 100×. (b) PAS staining of colon tissue. Original magnifications: 100×. (c) The damage score of colon tissue. (1) Epithelial cell damage: 0 = normal morphology; 1 = regional destruction of the epithelial surface; 2 = diffuse epithelial destruction and/or mucosal ulcers involving submucosa; 3 = severe epithelial damage. (2) Inflammatory cell infiltration: 0 = no infiltration or less than 5 cells; 1 = mild infiltration of the inherent layer; 2 = moderate infiltration of the muscular mucosa; 3 = high infiltration of the muscular mucosa; 4 = severe infiltration involving submucosa. (3) Separation of muscle layer and mucosal muscle layer: 0 = normal; 1 = moderate; 2 = severe. (4) Goblet cell depletion: 0 = no depletion; 1 = presence of disordered goblet cells; 2 = 1 to 3 regions without goblet cells; 3 = more than 3 regions without goblet cells; 4 = complete depletion of goblet cells. (d) The colon length. (e) The mucosal thickness of colon tissue. (f) Number of goblet cells in colon tissue. Results are expressed as the mean ± SD (n = 6). * p < 0.05 and ** p < 0.01 indicate significant differences between two groups.
HE score was calculated to evaluate the degree of colon tissue damage. The HE score in the DSS group was significantly higher (p < 0.01) than that in the CG group. Supplementation of M3G significantly reduced (p < 0.01) the score, which decreased by almost 50% of that in the DSS group (Fig. 2c). The colon length in the DSS group was significantly lower (p < 0.01) than that in the CG group, which was significantly increased (p < 0.01) by M3G supplementation (Fig. 2d). The mucosal thickness and the number of goblet cells of the colon tissue were determined by PAS staining. Compared with the CG group, the mucosal thickness and the number of goblet cells of the colon tissue were significantly decreased (p < 0.01) in the DSS group, but those were significantly increased (p < 0.01) in the M3G group (Fig. 2e & f). These results suggest that M3G supplementation can decrease the pathological damage in the colon tissue induced by DSS.
Effects of M3G on the colonic mucosal barrier function
-
The mRNA expression level of MUC2 in the colonic mucosa tissue was determined. Compared with the CG group, DSS significantly decreased (p < 0.01) the mRNA level of MUC2, but this effect was significantly suppressed (p < 0.01) by M3G supplementation (Fig. 3a). The protein levels of claudin-1, occludin, ZO-1, iFABP, and TFF3 in the colon tissue were subsequently assessed. The protein expression levels of claudin-1, occludin, and ZO-1 in the DSS group were significantly lower than those in the CG group (p < 0.01). M3G supplementation significantly inhibited (p < 0.01) the reduction in the expression of these proteins (Fig. 3b−d). Regarding the protein expression level of iFABP, it was significantly higher (p < 0.01) in the DSS group than that in the CG group. M3G supplementation significantly decreased (p < 0.01) iFABP expression level compared with the DSS group (Fig. 3e). The protein expression level of TFF3 was significantly higher (p < 0.01) in the DSS group than that in the CG group. However, M3G supplementation significantly intensified (p < 0.01) the increase of TFF3 expression compared with the DSS group (Fig. 3f). These results indicate that M3G can regulate colonic epithelial barrier function.
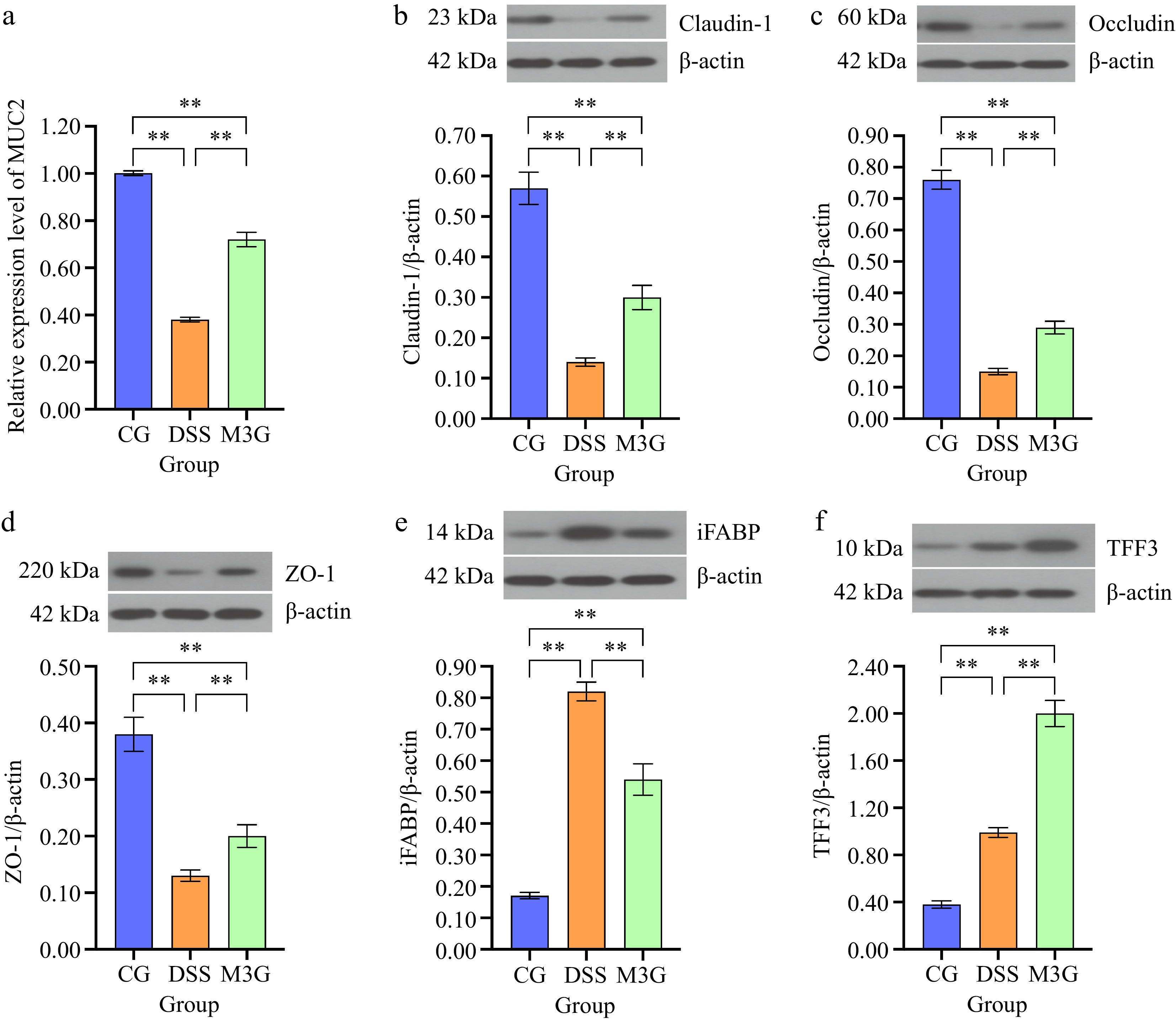
Figure 3.
Effects of M3G on the colonic epithelial barrier disruption. (a) The mRNA expression level of MUC2. (b) The protein expression level of claudin-1. (c) The protein expression level of occludin. (d) The protein expression level of ZO-1. (e) The protein expression level of iFABP. (f) The protein expression level of TFF3. Results are expressed as the mean ± SD (n = 6). * p < 0.05 and ** p < 0.01 indicate significant differences between two groups.
Effects of M3G on the colonic immune barrier function
-
The content of CD4+T (CD3+CD4+) and CD8+T (CD3+CD8+) cells in colonic LPMC of mice in each group were detected by FCM as shown in Fig. 4 and Supplemental Fig. S1. Compared with the CG group, the percentages of CD4+T (CD3+CD4+) and CD8+T (CD3+CD8+) cells in the DSS group were significantly increased (p < 0.01), and those were significantly decreased (p < 0.05) after M3G supplementation (Fig. 4a & b). SIgA was measured as an indicator of immune barrier function in the colon tissue. The level of SIgA in the DSS group was significantly lower (p < 0.01) than that in the CG group. Supplementation of M3G significantly raised (p < 0.01) the level of SIgA (Fig. 4c). These results indicate that M3G can improve the colonic immune dysfunction induced by DSS.
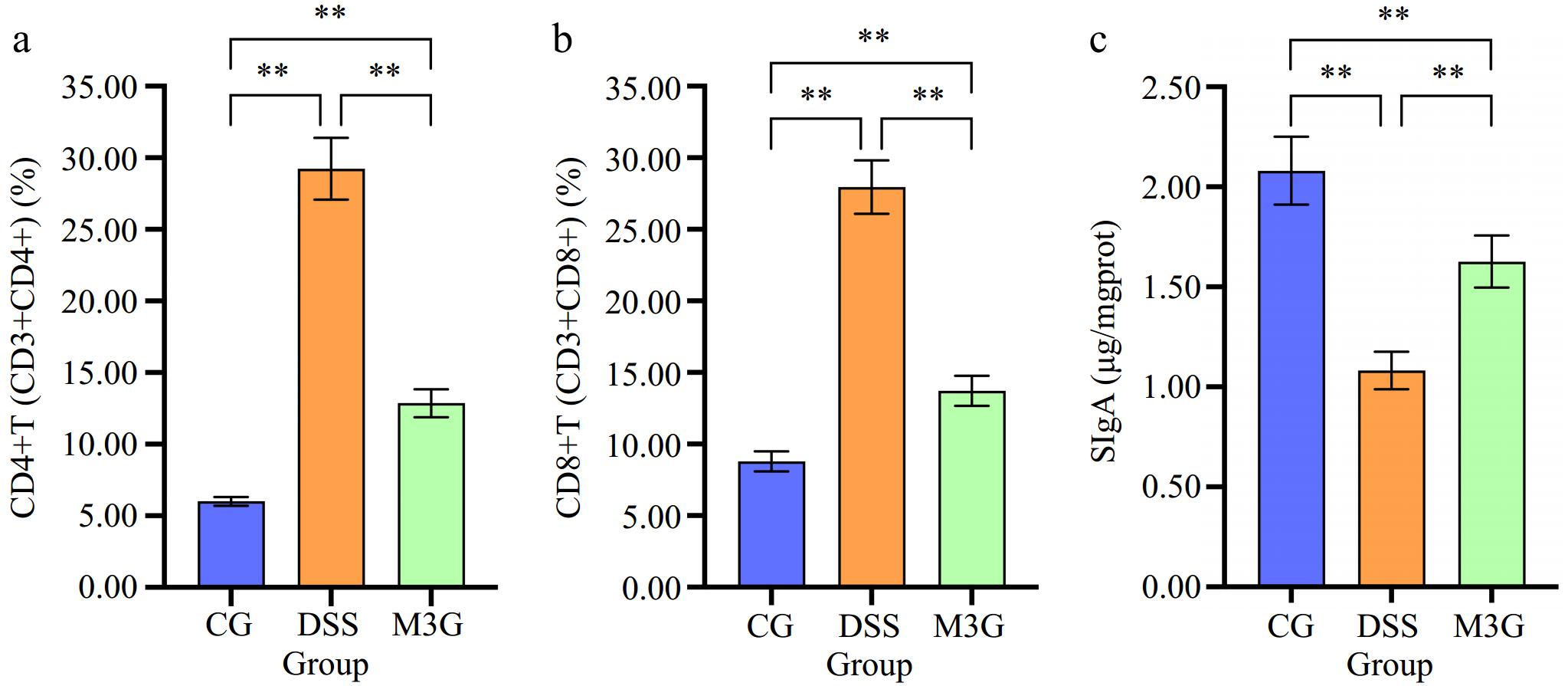
Figure 4.
Effect of M3G on the colonic immune barrier function. (a) The percentage of CD4+T (CD3+CD4+) cells in the colon tissue. (b) The percentage of CD8+T (CD3+CD8+) cells in the colon tissue. (c) The level of SIgA in the colonic mucosal tissue. Results are expressed as the mean ± SD (n = 6). * p < 0.05 and ** p < 0.01 indicate significant differences between two groups.
Effects of M3G on the Notch signaling pathway in colon tissue
-
Subsequently, the protein expression levels of Notch1, NICD, DLL4, DLL1, and Hes1 in the colon tissue were measured (Fig. 5). DSS significantly up-regulated (p < 0.01) these protein expression levels compared with the CG group. M3G supplementation significantly (p < 0.01) down-regulated the protein expression levels of Notch1, NICD, DLL4, DLL1, and Hes1 in comparison with the DSS group. The results indicate that M3G can inhibit the over-activation of the Notch signaling pathway.
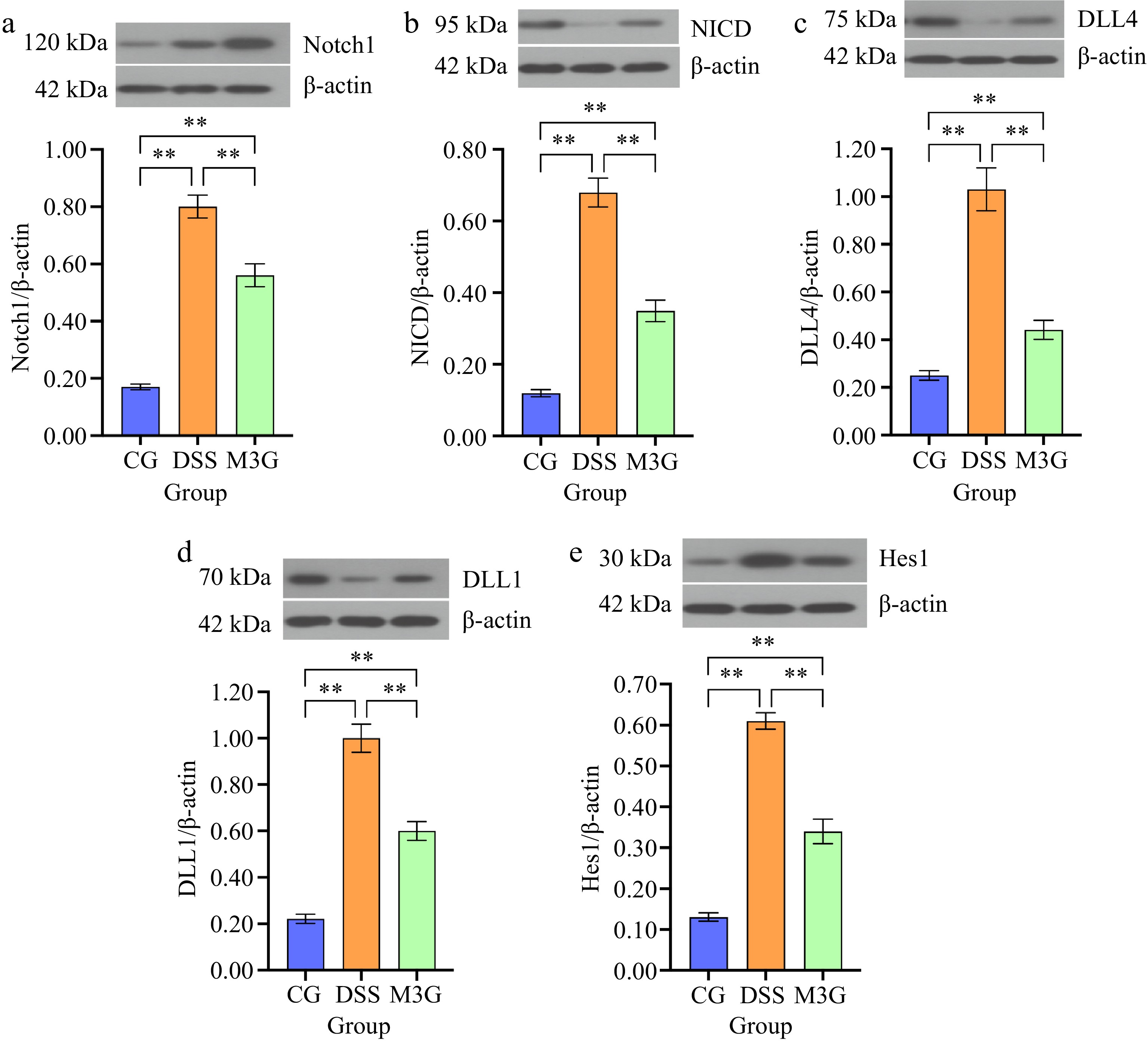
Figure 5.
Effects of M3G on the protein expression levels of Notch1, NICD, DLL4, DLL1, and Hes1 in the colon tissue. (a) The protein expression level of Notch1. (b) The protein expression level of NICD. (c) The protein expression level of DLL4. (d) The protein expression level of DLL1. (e) The protein expression level of Hes1. Results are expressed as the mean ± SD (n = 6). * p < 0.05 and ** p < 0.01 indicate significant differences between two groups.
Associations between physiological index and the Notch signaling pathway
-
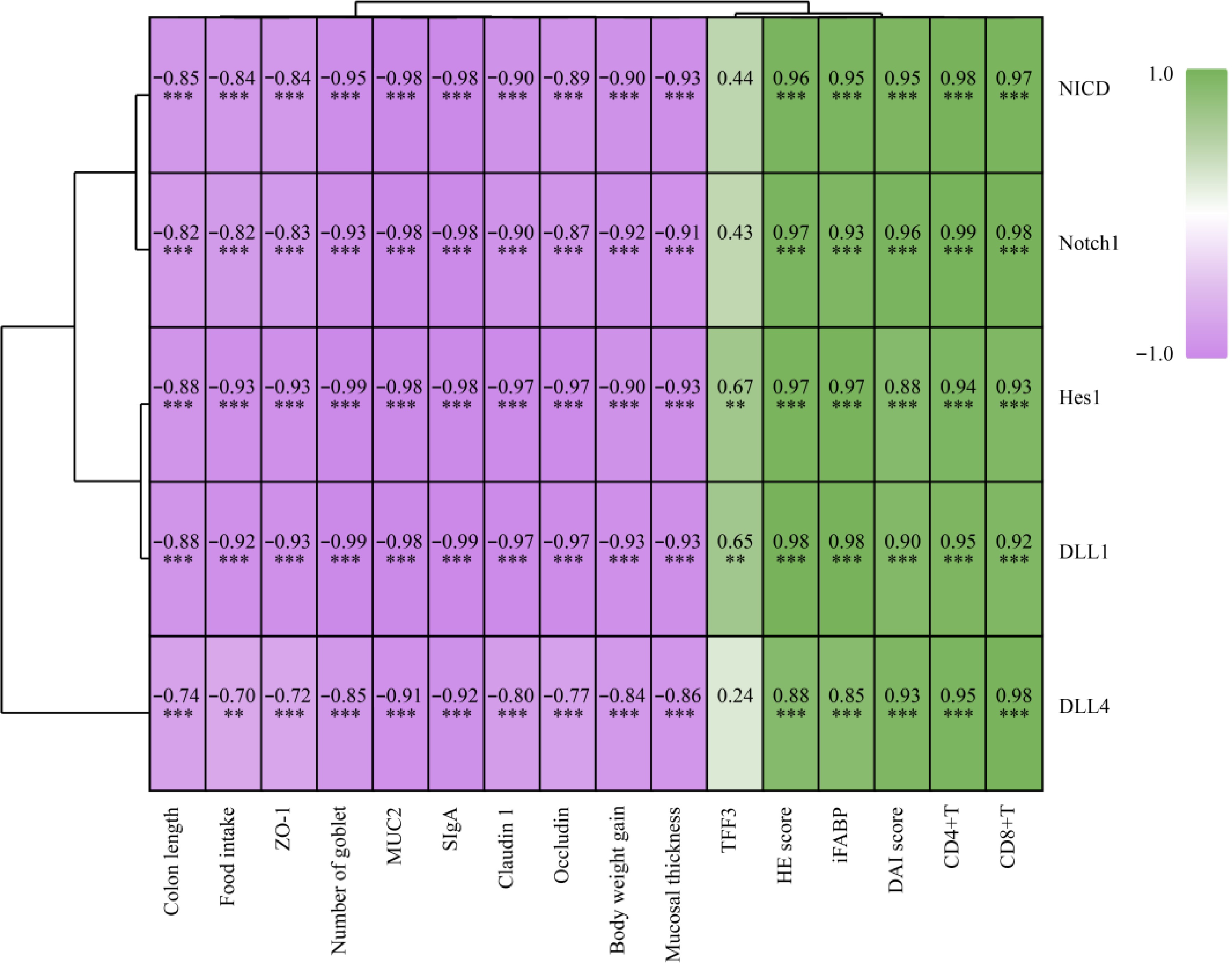
To explain the relationships between them, the Spearman r correlations between biomarkers and the Notch signaling pathway were analyzed, and represented using a heatmap. As shown in Fig. 6, colon length, food intake, ZO-1, number of goblet, MUC2, SIgA, claudin-1, occludin, body weight gain, and mucosal thickness were significantly (p < 0.01 or p < 0.05) positively correlated with Notch1, NICD, DLL4, DLL1, and Hes1. Conversely, HE score, iFABP, DAI score, CD4+T, and CD8+T were significantly (p < 0.01) negatively correlated with Notch1, NICD, DLL4, DLL1 and Hes1. In particular, TFF3 was only significantly (p < 0.05) positively correlated with DLL1, and Hes1. The results indicate that M3G may ameliorate the colonic mucosal barrier dysfunction via modulation of the Notch signaling pathway.
-
Within the last five years, special attention is being paid to the therapeutic effects of anthocyanins. The recent studies could give evidence to prove the therapeutic potential of anthocyanins from different sources against various diseases via in vitro, in vivo, and epidemiological experiments[28]. The anthocyanin supplementation has been demonstrated to have positive effects on intestinal health[29]. The intestinal barrier is one of the crucial factors which can affect intestinal health and normal intestinal barrier function not only maintains intestinal health but also protects overall health by protecting the body from intestine injury, pathogen infection, and disease occurrence[30]. The disruption of intestinal barrier integrity is regarded as an important factor leading to IBD, obesity, and metabolic disorders[31−33]. In this study, the effects of M3G on regulating the colonic physical barrier function and colonic immune barrier function in DSS-induced colitis mice were explored.
M3G supplementation reduced the DAI score and the HE score of colon tissue, and restored colon length, mucosal thickness, and the number of goblet cells in the colon tissue, indicating that M3G can alleviate DSS-induced colon tissue damage and colitis symptoms in mice. The DAI score was developed as a simplified clinical colitis activity index to assess the severity of colitis[34]. Zhao et al. have found that black rice anthocyanin-rich extract can significantly decrease the DAI and HE scores of colon tissue in DSS-induced colitis mice[35]. Intestinal goblet cells are mainly differentiated from multipotential stem cells. Intestinal stem cells were located at the base of the crypt and distributed in intestinal mucosal epithelial cells, composing around 50% of colon epithelial cells[36]. The mucus layer is formed by goblet cell secretion, it separates the intestinal epithelium from the intestinal lumen, thereby preventing the invasion of pathogenic microorganisms and the translocation of intestinal microbiota[37]. The results suggested that M3G might repair the colonic mucosa by increasing the number of goblet cells.
MUC2 is the most abundant mucin secreted by goblet cells of the colon, goblet cells, and MUC2 play important roles in maintaining and protecting the intestinal mucosal barrier[38]. M3G supplementation restored the level of MUC2 by nearly double that of the DSS group. Claudin-1, occludin, and ZO-1 are TJ proteins, which is an important component of the intestinal physical barrier[39]. M3G supplementation mitigated the down-regulation of claudin-1, occludin, and ZO-1 in DSS-induced colitis mice. Wang et al. have found that Lonicera caerulea polyphenols can increase the expression levels of occludin in HFD rats[40]. Chen et al. have found that purple-red rice anthocyanins alleviated intestinal barrier dysfunction in cyclophosphamide-induced mice by up-regulating the expression of tight junction proteins[41]. iFABP can serve as a biomarker of small bowel damage in coeliac disease and Crohn's disease, and supplementation of M3G down-regulated the expression level of iFABP in colon tissue[42]. It has been reported that TFF3 alleviated the intestinal barrier function by reducing the expression of TLR4 in rats with nonalcoholic steatohepatitis, and supplementation of M3G increased the expression level of TFF3 in this research[43]. Although the effect of M3G on epithelial TJ proteins and other related proteins were limited, the beneficial effect of M3G on the colonic mucosal barrier function was supported by histological evaluation.
The intestinal immune barrier is composed of intestinal mucosal lymphoid tissue and intestinal plasma cell secreted antibodies. Inflammatory damage in acute colitis influences the gut microbiota, epithelial barrier, and immune function in subsequent colitis[44]. Fructose can influence colon barrier function by regulating some main physical, immune, and biological factors in rats[45]. SIgA, CD4+T cells, and CD8+T cells play important roles in the functioning of the human immune system. The results indicated that M3G supplementation can maintain the colonic immune barrier function by modulating the level of SIgA, and the percentages of CD4+T cells and CD8+T cells of colon tissue.
Notch signaling pathways are important for the maintenance of intestinal epithelial barrier integrity, and its abnormal activation is related to IBD and colon cancer[46]. Among the four Notch receptors in mammals, the most scattered in the intestine is Notch1[47]. NICD is the active form of the Notch receptor, NICD enters the nucleus, binds to the recruitment co-activator transcription complex, and then combines with the Hes gene to regulate the fate of cells[48,49]. DLL1 and DLL4 can serve as ligands for Notch signaling receptors[50]. Lin et al. found that qingbai decoction had beneficial effects on the mucus layer and mechanical barrier of DSS-induced colitis by inhibiting Notch signaling[51]. Supplementation with M3G significantly down-regulated Notch1, NICD, DLL4, DLL1, and Hes1 expression levels in the colon tissue, and there is a significant correlation between biomarkers and Notch signaling pathway-related proteins, suggesting that M3G might sustain the colonic barrier function via inhibiting the Notch signaling pathway.
-
The present results showed that M3G exerted an improvement effect on the colonic mucosal barrier (physical barrier and immune barrier) function in DSS-induced colitis mice. The positive impact of M3G on the colonic mucosal barrier function may be ascribed to the modulation of permeability, stability, and integrity of the colonic mucosa by down-regulating the Notch signaling pathway. The results provide theoretical support for anthocyanins as the raw materials of functional products related to intestinal health. However, this study also has limitations, such as lacking intensive research on the effect mechanisms through cell experiments. Therefore, intensive research and clinical tests are the future direction.
-
The authors confirm contribution to the paper as follows: study conception and design: Zhang C, Jiao X; data collection: Zhang L; analysis and interpretation of results: Zhang C, Zhang L, Zhang B, Zhang Y; draft manuscript preparation: Zhang C, Wang Y, Tan H, Li L, Jiao X. All authors reviewed the results and approved the final version of the manuscript.
-
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
We would like to thank the Liaoning Provincial Department of Education Project - General Project (LJKMZ20221060).
-
The authors declare that they have no conflict of interest.
- Supplemental Fig. S1 (a) Flow cytometric analysis of CD4+T (CD3+ CD4+) cells. (b) Flow cytometry analysis of CD8+T (CD3+ CD8+) cells.
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press on behalf of China Agricultural University, Zhejiang University and Shenyang Agricultural University. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zhang C, Zhang B, Zhang L, Adel Ashour A, Wang Y, et al. 2024. Malvidin-3-O-galactoside ameliorates colonic mucosal barrier function via the Notch signaling pathway. Food Innovation and Advances 3(3): 279−287 doi: 10.48130/fia-0024-0026
Malvidin-3-O-galactoside ameliorates colonic mucosal barrier function via the Notch signaling pathway
- Received: 24 May 2024
- Revised: 19 July 2024
- Accepted: 05 August 2024
- Published online: 23 August 2024
Abstract: The colonic mucosal barrier is an important component of the intestinal barrier, and its integrity is crucial for maintaining digestive tract homeostasis and normal metabolism in the body. This study aimed to elucidate the mechanisms by which malvidin-3-O-galactoside (M3G) might ameliorate colonic mucosal barrier function, from the perspective of physical barrier function and immune barrier function. Male C57BL/6J mice were given dextran sulfate sodium (DSS) to establish a mice model for colitis and then administrated with or without M3G for one week. The results showed that M3G supplementation significantly improved the disease activity index (DAI) score and colon tissue injury in mice with DSS-induced colitis. M3G improved the colonic physical barrier function by modulating the expression of mucin2 (MUC2), claudin-1, occludin, zona occludens 1 (ZO-1), and intestinal fatty acid binding protein (iFABP) in the colonic mucosa. Additionally, M3G also relieved the colonic immune barrier of mice by increasing the level of secretory immunoglobulin A (SIgA) in colon tissue and the percentages of CD4+T (CD3+CD4+) and CD8+T (CD3+CD8+) cells in colon lamina propria monocytes in mice. Furthermore, M3G down-regulated Notch signaling pathway-related proteins such as Notch1, notch intracellular domain (NICD), delta-like ligand 4 (DLL4), delta-like ligand 1 (DLL1), and hairy/enhancer of split 1 (Hes1) of colon tissue. The present results demonstrated that M3G can improve colonic mucosal barrier function by inhibiting the Notch signaling pathway.
-
Key words:
- Malvidin-3-O-galactoside /
- Colonic mucosal barrier /
- Notch signaling pathway /
- Colitis