-
Hypertension refers to a long-term elevated blood pressure that mainly results from either nonspecific lifestyle change, genetic factors, or an identifiable cause[1,2]. Hypertension is a common chronic medical problem worldwide and has been confirmed as a leading factor in the cause of cardiovascular mortality[3−5]. Globally, 17 million people die annually due to cardiovascular disease, which is nearly a third of all deaths. Annually, hypertension has been linked to an estimated 9.4 million fatalities worldwide. It is believed to have caused at least 45% of coronary heart disease deaths and the majority of stroke deaths (51%)[6]. Hypertension has also been reported as a major risk factor for many other chronic diseases, such as vision loss, chronic kidney disease, atrial fibrillation, and dementia[7]. During the initial phase of hypertension, certain biomarkers in the bloodstream can be beneficial in understanding the mechanism of hypertension and may be used as potential treatments, including Angiotensin A, Vasoconstruction Inhibiting Factor (VIF), EndoThelin 1,2, Leptin, IL-1, IL-6, Nitric Oxide (NO), CRP, Renin, BNP, Uric Acid, and VCAM-1[8]. It is well known that hypertension plays an essential role in vascular endothelial dysfunction. Moreover, prolonged high blood pressure can increase the shear stress on blood vessels, leading to structural and functional damage to the vascular endothelium[9,10]. Vascular endothelial cells take charge in secreting vasoconstrictive factors, including NO, Prostaglandin-I-2 (PGI2), endothelin (ET) and Angiotensin II (Ang II)[11−13]. These vasoconstrictive factors regulate the vasoconstriction of vascular endothelium and they exist in an equilibrium state under a normal physiological condition[14−16]. For example, NO, ET and Ang II have been confirmed to possess vasoconstrictive functions, whereas PGl2 is a key factor that could regulate vascular tone[17]. It has been reported that hypertension could alter the synthesis of these vasoconstrictive factors, which could result in the collapse of their equilibrium in vascular endothelium[16]. Additionally, hypertension could take place with systemic inflammation, and thus some inflammatory mediators could be used to indicate the vascular damage caused by high blood pressure[18]. For example, interleukin 6 (IL-6) and tumor necrosis factor-α (TNF-α) are two inflammatory mediators that can be secreted into vascular endothelium by vasoactive substances (adrenaline and Ang II). IL-6 could activate platelet activating factors to enhance platelet aggregation and produce blood clots, whereas TNF-α could damage the integrity of vascular endothelial cells and trigger the inflammation to the vascular wall[19−22].
Medicinal plants have gained more attention in the field of medical science due to their potential health beneficial features. Gaocha is one of the most traditional tea beverages in China and other Asian countries. Gaocha is made of the tender shoots of A.ginnala Maxim, and it has been reported to contain various health promoting nutrients, such as alkaloids, tannins, flavonoids, and organic acids[23−26]. Previous studies have reported that Gaocha possesses antioxidant activity and the consumption of Gaocha could inhibit tumor activity[27,28], hypoglycemic effect[29,30] and bacteriostatic action[31]. Gaocha has been used as a folk medicine to alleviate blood pressure. However, to the best of our knowledge, its antihypertensive mechanism has not been well studied. To this end, we treated spontaneously hypertensive rats (SHRs) with Gaocha extract at different doses and assessed the alteration of their blood pressure. Meanwhile, the vascular endothelial function factors (NO, ET, PGl2 and Ang II) and inflammatory mediators (IL-6 and TNF-α) of these SHRs under different Gaocha dose treatments were analyzed and compared. The findings from this study could provide useful insight on elucidating the mechanism of Gaocha on the hypertension alleviation.
-
Catechin, epicatechin, gallocatechin, epigallocatechin, galloyl acid and β-glucogallin were purchased from Sigma-Aldrich (St. Louis, MO, USA) with a purity of 99%. Methanol, acetonitrile, and acetic acid were of HPLC grade and purchased from Tedia Co., Ltd. (Fairfield, OH, USA). Captopril tablets are a product of Changzhou Pharmaceutical Co., Ltd (Jiangsu, China). Normal saline and chloral hydrate were purchased from Kaiyuan Pharmaceutical Co., Ltd (Tianjin, China) and Shanghai Qiangshun Chemical Reagent Co., Ltd (Shanghai, China), respectively. Rat IL-6, ET-1, PGl2, Ang II, NO and TNF-α were all purchased from Sigma-Aldrich (St. Louis, MO, USA).
Gaocha extract
-
The raw materials used for the Gaocha samples are sourced from Shucheng County, Lu'an City, Anhui Province (China) in 2017, and are processed by Anhui Lvyuan Tea Company. The production process of Gaocha involves picking, light withering, fixation, and baking to yield the final product[32]. The Gaocha sample was extracted using hot water at 100 oC with a 1:10 w/w ratio for 30 min under sonication. Afterwards, the resultant mixture was centrifuged at 12,000 rpm for 10 min to collect Gaocha extract. The Gaocha extract was then freeze-dried to yield the dryness. The dried extract was diluted using distilled water to a concentration of 1 g/mL. The resultant extract was filtered through a 0.22 μm membrane and then directly injected to liquid chromatography.
Ultra-Performance Liquid Chromatography (UPLC)
-
An Agilent 1260 series UPLC (Palo Alto, Santa Clara, USA) was used to analyze the phenolic compounds composition in the Gaocha extract. The injection volume was set at 5 μL. An Agilent Eclipse PlusC18 column (Palo Alto, Santa Clara, CA, USA) was used to separate phenolic compounds under a flow rate of 1.0 mL/min. The mobile phase consisted of (A) water and (B) acetonitrile. A gradient elution program was as follows: 0−7 min, 10%B to 30%B; 7−10 min, 30%B isocratic; 10−11 min, 30%B to 13%B; 11−16 min, 13%B isocratic; 16−17 min, 13%B to 10%B; and 17−20 min, 10%B isocratic. The column was maintained at 40 °C during the elution program. The detection wavelength on the diode array detection was set at 280 nm. The detected individual phenolic compounds were quantified using their corresponding standard.
Animal model
-
A total of 40 pathogen-free male spontaneously hypertensive rats (SHRs) were provided from Beijing Vital River Laboratory Animal Technology Co., Ltd with its laboratory animal production license of SCXK(Jing) 2012-0001 (Beijing, China). These SHRs were 12-weeks-old with a body weight of 250 ± 20 g. Meanwhile, 8-12-week-old pathogen-free normal male rats were also purchased from Beijing Vital River Laboratory Animal Technology Co. Ltd with the same body weight as the SHRs. Both SHRs and normal rats were acclimated on a AIN93 G diet at 24−26 °C in a 12 h light/dark cycle for 7 d before administration. The SHRs were randomly divided into five groups and eight rats in each group was placed into two cages with four rats per cage. The normal rats were also put into two cages with four rats per cage. The SHRs in the first group (disease control group) were fed with an equal volume of distilled water as the treated SHRs. The second SHRs group was fed with captopril (6.25 mg/kg of rat weight). The rats in the third, fourth, and fifth SHRs group were gavaged with the Gaocha extract with a dose of 12.0 mg/kg (high dose), 6.0 mg/kg (mid dose) and 3.0 mg/kg (low dose) per rat weight, respectively. The Gaocha dried extract was dissolved using distilled water to a concentration of 1.2, 0.6, and 0.3 mg/mL for oral gavage. The normal rats (healthy control group) were fed with an equal volume of distilled water as the treated SHRs. The drug/extract administration took place at 9:00 am once per day for 16 d. During the experiment these rats had access to the same diet and water, and hair color, growth and general behavior of the rats were monitored. All rats were penned individually and randomly assigned to the pens.
Tail artery blood pressure measurement
-
Tail artery blood pressure of all the rats were measured using the tail-cuff method at 0, 3, 7, 10, and 15 d under ALC-NIBP system (Shanghai Alcott Biotech. Co. Ltd., Shanghai, China). Each pressure measurement was conducted three times at 60 min intervals.
Serum Ang II, NO, ET, PGl2, IL-6 and TNF-α measurement
-
After the last oral gavage, all the rats were fasted for 12 h without any food or water. The rats were then intravenously administered with 0.01 ml/g 10% chloral hydrate to induce anesthesia and then 5 mL of blood was collected and sampled from their aorta abdominals. Subsequently, the blood sample was transferred to EDTA tubes and then to vacuum blood collection tubes. The blood samples were centrifuged at 3,000 r/min for 30 min to separate plasma and serum. The targets (Ang II, NO, ET, PGl2, IL-6 and TNF-α) were analyzed using ELISA on an RT-6000 ELISA analyzer (Rayto Life and Analytical Sciences Co. Ltd, Shenzhen, China) according to the manufacturer’s instructions.
Histomorphological examination of rat heart and kidney
-
After the blood was sampled, the corresponding organs were resected and soaked in 4% formaldehyde for 24 h. Afterwards, the organs (heart and kidneys) of the rats were placed in a Petri dish. The collected organ samples were washed with normal saline, dried using filter papers, and then fixed by 10% methanol. The fixed organs were dehydrated using a gradient series of ethanol and then passed through xylene solution to remove the ethanol and facilitate molten paraffin wax infiltration at 55 °C. Subsequently, the organs were embedded into a wax block and then cut into 4 mm thickness paraffin section. The paraffin sections were then stained with hematoxylin and eosin and visualized using a high-power microscope (OlympusBX51, Olympus Medical System Corp., Tokyo, Japan).
Statistical analysis
-
Data were expressed as the mean ± standard error. One-way analyses of variance (ANOVA) were used to compare the means using Duncan's range test on SPSS17.0 (Chicago, IL, USA). Homogeneity analysis was carried out using the least significant different test. A p ≤ 0.05 difference was considered significant.
-
Upon analysis of the functional component composition of Gaocha extract, it was found that the dried extract was affluent in phenolic compounds, with a total polyphenol content of 75.4 mg/g. Moreover, ultra-performance liquid chromatography revealed the presence of β-glucogallin, galloyl acid, gallocatechin, epigallocatechin, catechin, and epicatechin in the extract (Table 1). The predominant phenolic compound in the dried extract of Gaocha was revealed to be epicatechin, with a content of 32.68 mg/g, followed by epigallocatechin (5.69 mg/g) and gallocatechin (3.11 mg/g). Additionally, catechin was present in the extract at 1.95 mg/g. These findings demonstrate that Gaocha is rich in multiple polyphenols, potentially contributing to its beneficial effects.
Table 1. Concentration of phenolic compounds in dried Gaocha extract.
Phenolic compound Content (mg/g) Percentage (%) β-Glucogallin 0.50 ± 0.00 0.67 ± 0.00 Galloyl acid 0.74 ± 0.01 0.99 ± 0.02 Gallocatechin 3.11 ± 0.07 4.13 ± 0.14 Epigallocatechin 5.69 ± 0.08 7.55 ± 0.15 Catechin 1.95 ± 0.06 2.59 ± 0.05 Epicatechin 32.68 ± 0.76 43.35 ± 1.09 Polyphenol 75.40 ± 0.71 Polyphenol content was analysed using Folin-Cioalteu and expressed as mg galloyl acid/g dried Gaocha extract. Individual phenolic compounds were quantified using their corresponding standard and expressed as mg/g dried Gaocha extract. Data are the mean ± standard deviation of triplicate tests. Gaocha extract effect on hair growth and behavior of SHRs
-
To gain insight into the general behavioral effects of Gaocha extract on SHRs, we conducted the experiments involving feeding them the extract and monitoring their behavior. The rats in the healthy control group exhibited glossy hair and gentle behavior (data not shown). In contrast, the SHRs in the disease control group showed an aggressive behavior and their hairs were thinner and fluffier. The SHRs treated with the Gaocha extract had more closely distributed and glossy hair, and those with the high Gaocha dose administration had similar hair growth and behavior as the SHRs treated with captopril (positive control). All the rats were kept in the same conditions with free access to water and food during the experiment.
Gaocha extract effect on tail artery blood pressure of SHRs
-
In order to evaluate the potential of Gaocha extract in reducing blood pressure, we monitored the tail artery blood pressure of SHRs. The results in Table 2 showed the tail artery systolic blood pressure of these rats during the experimental period. It was observed that the rats in the healthy control group had a blood pressure ranging from 109.92 to 116.49 BP/mm Hg. The SHRs in the disease control group exhibited an elevated blood pressure compared to the normal rats during the whole experiment period and their blood pressure was about 166.34 to 167.69 BP/mm Hg. The SHRs in the positive control group (captopril) had a lower blood pressure during the study and their blood pressure remained around 110 BP/mm Hg. The Gaocha extract resulted in a significant decrease in the blood pressure of the SHRs, and such pressure alleviation relied on the administration dose of the extract (Table 2). For example, the SHRs orally gavaged with the low Gaocha extract dose reduced their pressure from 169 BP/mm Hg to about 134 BP/mm Hg. The Gaocha extract with the mid dose lowered the blood pressure of the SHRs to about 125 BP/mm Hg. It is noteworthy that the SHRs treated with the high Gaocha extract dose had a similar blood pressure as the normal control rats and the captopril treated SHRs, suggesting that the high Gaocha extract dose may be effective in reducing high blood pressure.
Table 2. Tail artery systolic blood pressure of normal and spontaneously hypertensive rats treated with different Gaocha extract levels during the administration period.
Group Blood pressure during administration period (BP/mm Hg) 0 day 3 day 7 day 10 day 15 day Normal rats (healthy control) 114.48 ± 2.47e 109.92 ± 5.75e 112.23 ± 2.40e 116.49 ± 6.43d 115.87 ± 3.56d SHRs (disease control) 166.78 ± 2.71b 167.69 ± 2.46a 166.81 ± 2.71a 167.52 ± 2.56a 166.34 ± 2.99a SHRs with captopril (6.25 mg/kg) 165.36 ± 3.23b 123.92 ± 7.29cd 114.65 ± 2.71de 116.86 ± 4.62d 109.72 ± 3.51e SHRs with high Gaocha dose (12 mg/kg) 171.77 ± 5.13a 122.18 ± 4.67b 117.77 ± 3.53b 122.75 ± 1.72b 121.82 ± 3.41b SHRs with mid Gaocha dose (6 mg/kg) 161.72 ± 3.77c 127.99 ± 4.67c 125.37 ± 2.25c 125.48 ± 3.07c 123.91 ± 2.12c SHRs with low Gaocha dose (3 mg/kg) 169.17 ± 9.12ab 139.46 ± 1.69d 134.41 ± 3.85d 134.25 ± 3.49c 133.09 ± 3.76c Data are the mean ± standard error. Different letters in each column represent significant difference at a 0.05 significant level. Gaocha extract effect on PGI2, Ang II and ET of SHRs
-
To explore the potential mechanism of Gaocha extract in lowering blood pressure, we analyzed its effects on PGI2, AngII, and ET level of SHRs. The healthy rats (healthy control group) had a PGI2 level of 30 ng/L, whereas its level was only about 10 ng/L in the SHRs of disease control group (Fig. 1a). Administrating captopril (positive control) to the SHRs for 16 d resulted in a level increase of PGI2 in the SHRs. The SHRs treated with the low Gaocha extract dose exhibited a similar PGI2 level as the disease control group. However, the mid and high Gaocha extract dose significantly increased the PGl2 level in the SHRs, and the SHRs with the high extract dose administration had a similar PGl2 level as the positive control SHRs (captopril).
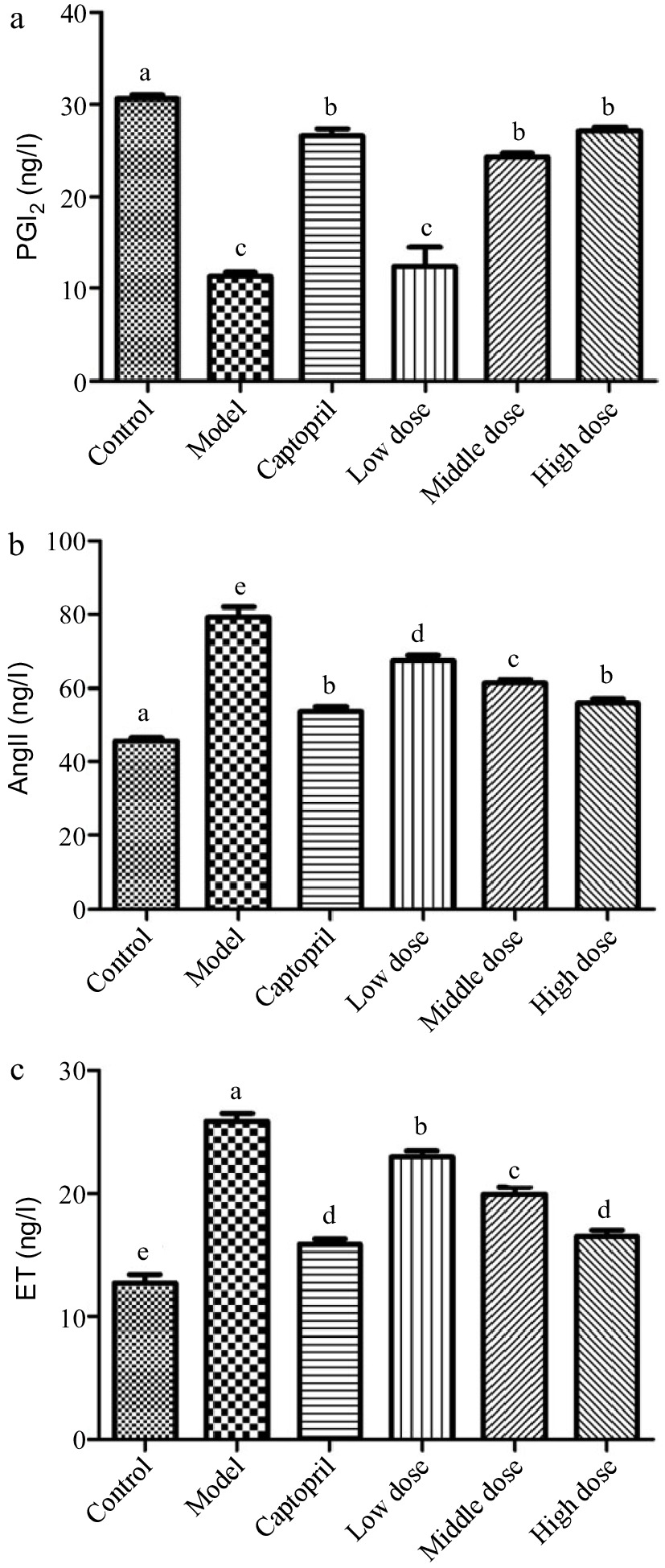
Figure 1.
The serum PGI2, AngII and ET levels in normal and spontaneously hypertensive rats treated with different Gaocha extract doses. (a) Serum PGI2 level, (b) Serum AngII level, and (c) Serum ET level. Different letters represent significant difference at a significant level of 0.05.
The Ang II level in the healthy control group rats was about 45 ng/L in this study, whereas the SHRs in the disease control group had an about 80 ng/L Ang II level (Fig. 1b). The captopril (positive control) administration significantly reduced the Ang II level to about 55 ng/L in the SHRs. Feeding the SHRs with the Gaocha extract resulted in a decrease on the Ang II level. Such a decrease was much more obvious with the higher dose administration.
In addition, the SHRs in the disease control group exhibited the ET level two times higher than the healthy control group rats (Fig. 1c). The captopril treatment significantly lowered the ET level in the SHRs. The Gaocha extract administration with different doses also resulted in an ET level decrease in the SHRs after the experiment and the high Gaocha extract dose appeared to result in the SHRs with the similar ET level as the positive control (captopril).
Gaocha extract effect on NO of SHRs
-
Nitric oxide (NO) is a key signaling messenger that is biosynthesized from L-arginine, oxygen, NADPH through nitric oxide synthase enzymes[33,34]. It has been reported that NO could relax the smooth muscle in the endothelial cells to widening the blood vessel and increasing blood flow[35,36]. Figure 2 shows the NO level of these rats after the whole animal study. The normal rats (healthy control group) were found to have a NO level around 8 ng/L, whereas spontaneously hypertension caused the rats to have the NO level less than 4 ng/L. After treating the SHRs with the positive control captopril for 16 d, the NO level of the SHRs was elevated to about 7 ng/L. It should be noted that the administration of the Gaocha extract for 16 d did improve the NO levels in the SHRs. The NO level in the SHRs treated with the low, mid, and high dose of the Gaocha extract appeared to be about 4, 5, and 7 ng/L, respectively. These indicated that the Gaocha extract could stimulate the biosynthesis of NO in the SHRs, which could enhance the vasodilation and reduce the blood pressure.
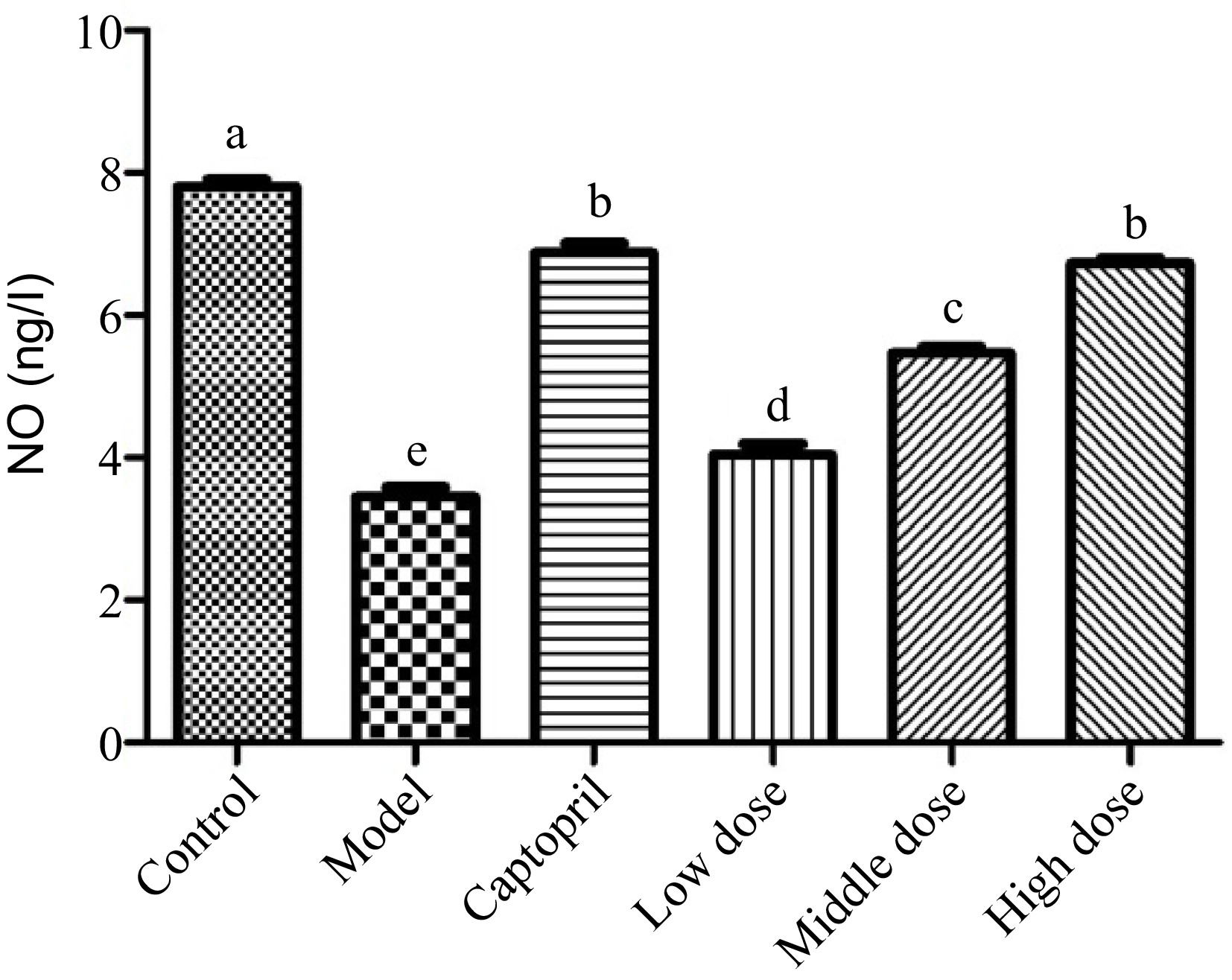
Figure 2.
The serum nitric acid (NO) levels in normal and spontaneously hypertensive rats treated with different Gaocha extract doses. Different letters represent significant difference at a significant level of 0.05.
Gaocha extract effect on IL-6 and TNF-α of SHRs
-
Interleukin 6 (IL-6) is an interleukin that could be produced as a pro-inflammatory cytokine by smooth muscle cells in the tunica media of blood vessels[37]. It has been reported that muscle contraction could stimulate the secretion of IL-6 into blood stream[18]. The spontaneous hypertension significantly increased the release of IL-6 level in the rats (Fig. 3a). The healthy control group rats had a IL-6 level of about 12 ng/L, whereas its level in the SHRs control group was higher than 25 ng/L. The SHRs treated with captopril for 16 d showed its IL-6 level at around 15 ng/L, indicating that captopril inhibited the biosynthesis and secretion of IL-6 in the SHRs. The SHRs gavaged with the Gaocha extract for 16 d also reduced the secretion of IL-6. For example, the IL-6 level in the SHRs with the mid and high dose of the Gaocha extract were much lower than the SHRs control group, and the high Gaocha extract dose treated SHRs showed a similar IL-6 level as the SHRs administrated with captopril.
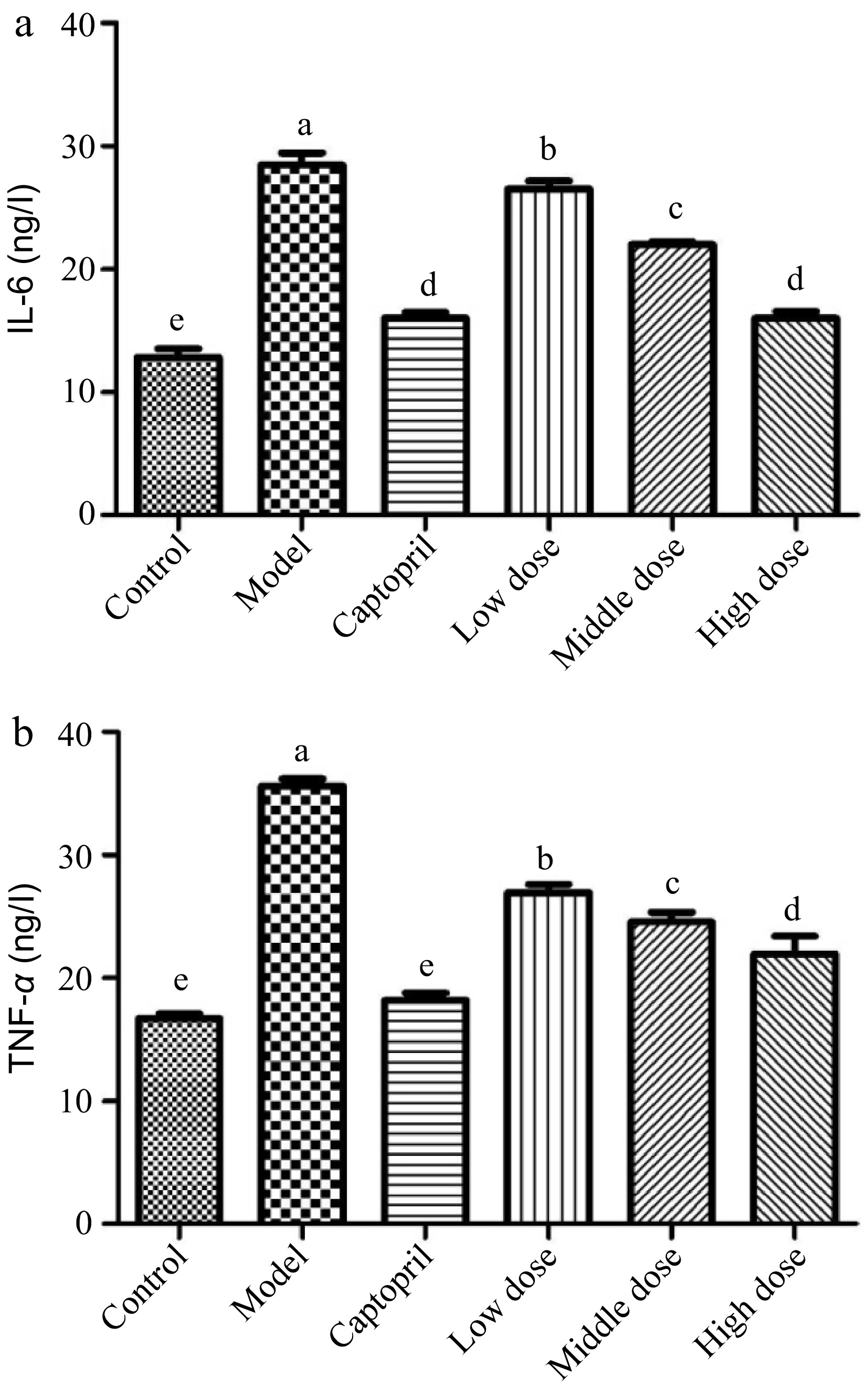
Figure 3.
The serum IL-6 and TNF-α levels in normal and spontaneously hypertensive rats treated with different Gaocha extract doses. (a) Serum IL-6 level and (b) Serum TNF-α level. Different letters represent significant difference at a significant level of 0.05.
Tumor necrosis factor-α (TNF-α) is a proinflammatory cytokine that can be elevated in an inflammatory state, such as hypertension[19,20]. It has been reported that the formation of TNF-α is associated with salt-sensitive hypertension and related renal injury, whereas Ang II increase could also promote the release of TNF-α[21,38]. In the present study, the SHRs control group showed a much higher level of TNF-α than the healthy control group rats (Fig. 3b). This indicated that hypertension stimulated the release of TNF-α in the SHRs. Both captopril and Gaocha extract inhibited the secretion of TNF-α in the SHRs after 16 d of treatment. For example, the captopril treated SHRs possessed a similar TNF-α level as the healthy control rats. The SHRs that were gavaged with Gaocha extract showed a significantly lower level of TNF-α compared to the control group. Furthermore, increasing the dose of the extract resulted in a greater inhibition of TNF-α release in the SHRs.
Gaocha extract effect on organ histomorphological features in SHRs
-
Regarding the rat heart histomorphological analysis, no thickening on the cardiac muscle fiber or fibrous hyperplasia was found in the healthy control group rats (Fig. 4a). However, the cardiac muscle fibers in the SHRs control group (disease control group) were significantly thickened with the muscle tissue hyperplasia (Fig. 4c). These demonstrated that spontaneous hypertension damaged the organs in the SHRs. In the captopril treated SHRs (positive control), their cardiac muscle fibers appeared to be normal without thickening, and their blood cells were visible in the cardiac cavities. Besides, no fibrous hyperplasia was found in the mesenchyme of the rats (Fig. 4b). The administration of the Gaocha extract improved the cardiac muscle fibers compared to the SHRs control although the thickening of the fibers still occurred to these Gaocha extract fed SHRs (Fig. 4d−f). It should be noted that the cardiac muscle fibers in the SHRs with the high Gaocha extract dose appeared to be much thinner than the low extract dose fed SHRs. Meanwhile, the blood cells were observed in the cardiac cavities of these SHRs treated with the Gaocha extract. More importantly, no obvious tissue hyperplasia was found in these Gaocha extract treated SHRs.
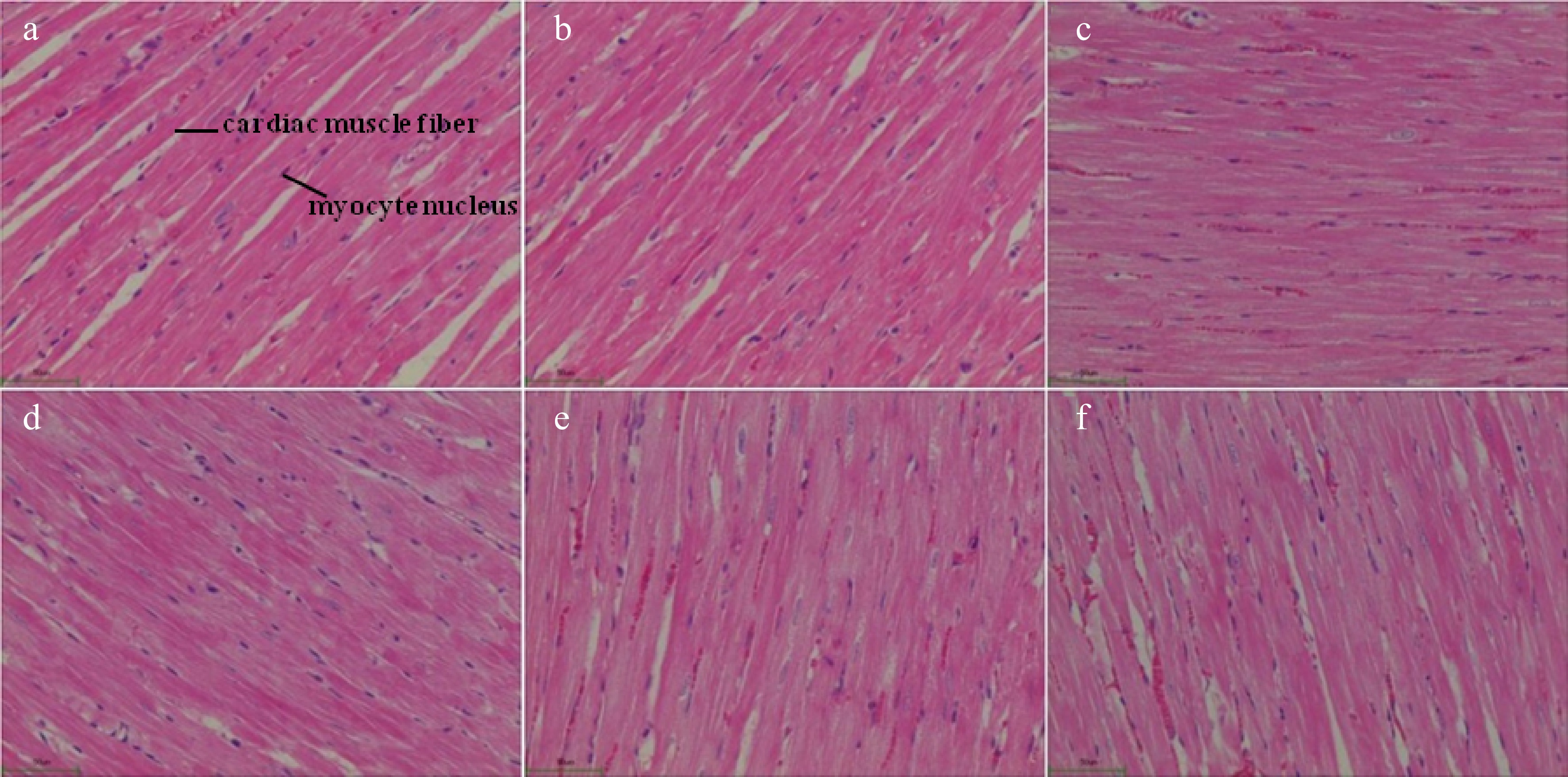
Figure 4.
Morphological feature of cardiac tissue in normal and spontaneously hypertensive rats treated with different Gaocha extract doses. (a) Normal rats in healthy control group.(b) Captopril treated spontaneously hypertensive rats. (c) Spontaneously hypertensive rats in disease control group. (d) Low Gaocha extract dose treated spontaneously hypertensive rats. (e) Mid Gaocha extract dose treated spontaneously hypertensive rats. (f) High Gaocha extract dose treated spontaneously hypertensive rats.
The healthy control group rats possessed normal renal tubules in the kidney histomorphological biopsy (Fig. 5a). However, a significant stenosis of the renal tubules happened in the SHRs (disease control group). Meanwhile, these SHRs were found to have fibrosis in the renal interstitium with a structural alteration on the glomerulus (Fig. 5c). The captopril treated SHRs improved the structure of the renal tubules and glomerulus and the histomorphological feature of the SHRs kidney was similar to that of the healthy rats. An improvement of the kidney structure on the SHRs treated with the Gaocha extract was also found. Among these treatments, the high Gaocha extract dose resulted in SHRs with normal renal tubules and glomerulus. Meanwhile, no obvious renal interstitial fibrosis or renal tubule stenosis was found in these SHRs.
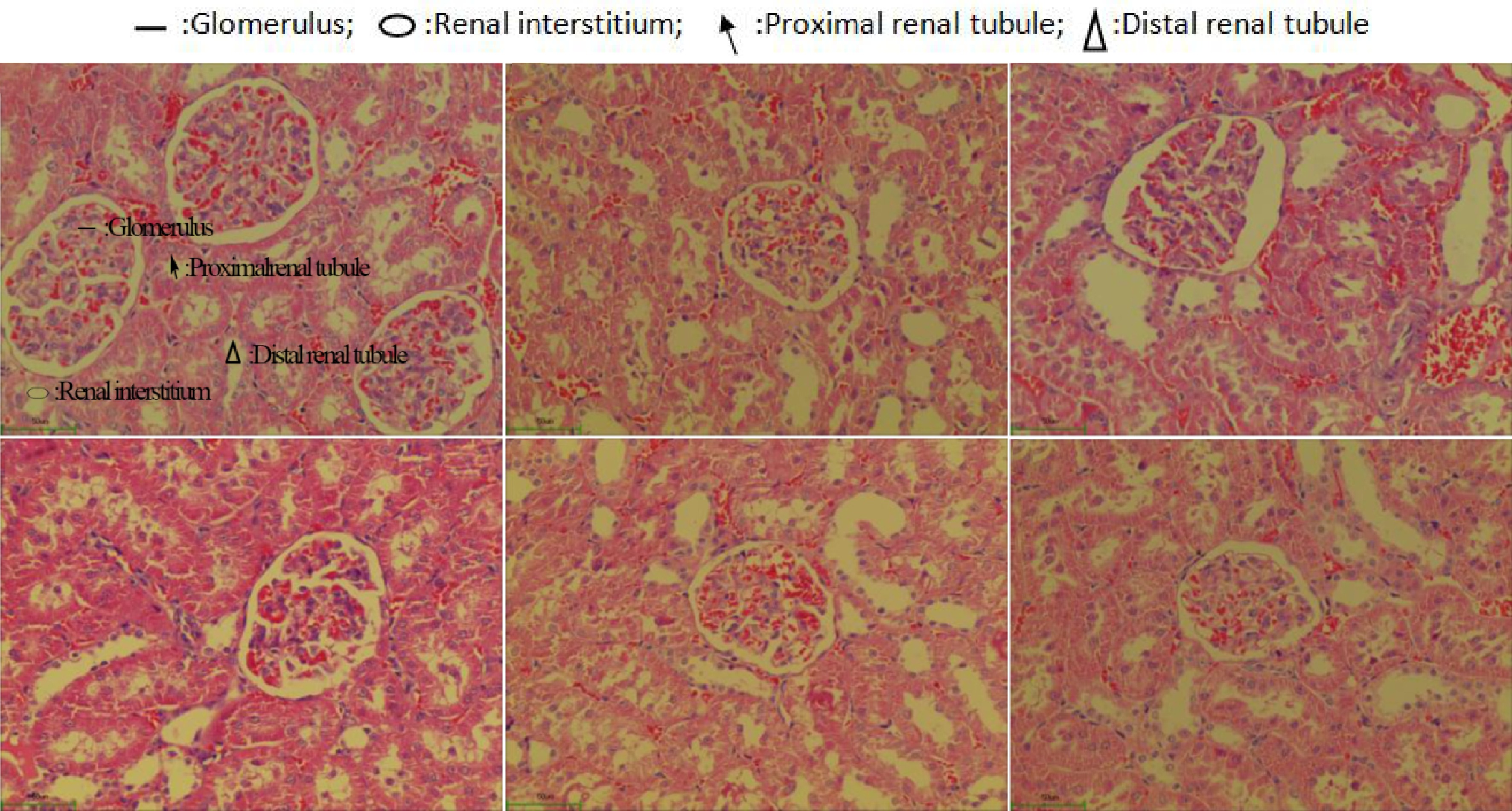
Figure 5.
Morphological feature of renal tissue in normal and spontaneously hypertensive rats treated with different Gaocha extract doses. (a) Normal rats in healthy control group.(b) Captopril treated spontaneously hypertensive rats. (c) Spontaneously hypertensive rats in disease control group. (d) Low Gaocha extract dose treated spontaneously hypertensive rats. (e) Mid Gaocha extract dose treated spontaneously hypertensive rats. (f) High Gaocha extract dose treated spontaneously hypertensive rats.
In this study, we explored the anti-hypertensive effects of Gaocha extract on spontaneously hypertensive (SHR) rats. It was found that the groups treated with Gaocha had lower systolic blood pressure in the tail artery compared to the control group. Additionally, the levels of certain biomarkers, such as PGI2, Angiotensin II, ET, and NO, were higher in the Gaocha-treated groups. After treatment, the levels of IL-6 and TNF-α were lower in the Gaocha-treated groups compared to the control group. Histomorphological examination of the heart and kidney indicated that Gaocha may possess a protective effect. Future research will aim to identify the active components and elucidate the associated health benefits.
-
Gaocha has been reported to contain numerous nutrients that could provide multiple benefits to human health[23−26]. Among these nutrients, polyphenols appear to be a major group for such beneficial activities. It has been confirmed that polyphenols are important antioxidants that could prevent the occurrence of many active and chronic diseases and these secondary metabolites could also regulate the secretion of signaling molecules in the human body[39,40]. Interestingly, our results showed that the content of EC was extremely high in Gaocha extracted solution. It was reported that the EC had a positive effect on vascular function in humans by inhibition of NO synthase[33]. Furthermore, it was also reported that epicatechin (EC) may in part contribute to the cardioprotective effects of cocoa and tea by improving insulin resistance[34]. These suggested that the high content of EC could be part of the reason for the vascular function of Gaocha. Moreover, Gallic acid (GA) was also reported as an active compound in Gaocha. Previous studies have demonstrated that GA can enhance NO production by augmenting the phosphorylation of eNOS. Additionally, GA has been found to impede the activity of ACE, leading to a reduction in the blood pressure of spontaneously hypertensive rats[35]. It can increase eNOS phosphorylation levels by influencing the Ca2+/CaM compounds, and also inhibit calcium influx into cells through the L-type calcium channel, resulting in the dilation of endothelial cells[36]. It is yet to be determined whether the GA lowers blood pressure, and this will be the focus of future research involving active ingredients, molecular and cellular studies.
Hypertension is a progressive cardiovascular condition, caused by different factors, which can cause changes in the heart and blood vessels' structure and function. It affects one in every three adults and an increase of 1 mm of mercury in hypertension patients increases the mortality rate by 1%. Therefore, research on molecular mechanisms is essential for the prevention and treatment of hypertension[12]. Hypertension is a multi-faceted medical condition, composed of several risk factors, such as atherosclerosis, endothelial cell damage, hyperlipidemia, and subclinical diseases, that can culminate in cardiovascular events. Its development involves endothelial dysfunction, vascular remodeling, inflammation, calcification, and increased vascular stiffness[11]. Endothelial dysfunction is believed to be an important factor in the pathogenesis of hypertension[7], as it has a genetic basis and can affect target organs. Furthermore, increased shear stress in the blood vessels due to hypertension can cause changes in the structure and function of vascular endothelium[41].Vascular endothelial cells are capable of releasing a variety of active factors that can cause blood vessels to relax or contract through autocrine and paracrine pathways, such as NO, PGI2, ET, and Ang II[11].
In hypertension, the contraction of blood vessels causes ischemia and hypoxia of the endothelial cells, leading to damage to their structure and function, and consequently, affects the production and release of NO. Oxidative stress has been identified as a major contributor to the processes of endothelial dysfunction, inflammation, hypertrophy, apoptosis, migration, fibrosis, angiogenesis, and hypertensive vascular remodeling[13]. Endothelin has a pronounced effect of decreasing vascular diameter, in comparison to Angiotensin II, which is produced by hyperpolarization of vascular endothelial cells and also has a vasoconstrictive action but with less intensity. Ang II is a peptide hormone that is basically converted from Ang I through angiotensin-converting enzyme (ACE)[42]. Ang II has been confirmed to act on venous and arterial smooth muscle to increase vasoconstriction. ACE inhibitors could restrain the conversion of Ang I to Ang II and thus improve the blood pressure[43]. Feeding the SHRs with the Gaocha extract resulted in a decrease in the Ang II level. Such a decrease was much more obvious with the higher dose administration. We speculated that phenolic compounds, such as epicatechin and epigallocatechin, in the Gaocha extract might inhibit the renin-angiotensin system (RAS) activation pathway to reduce the secretion of Ang II in the SHRs. In this way, the vascular wall of the SHRs could be protected.
In addition, ET is a peptide that is primarily present in endothelium and its overexpression could drive hypertension, heart diseases and other organ damages[44]. ET is a vasoconstrictor that interacts with smooth muscle endothelin receptors[45]. It has been reported that epicatechin could inhibit the interaction between ET and its endothelin receptors[46]. We hypothesized that the high level of epicatechin, found in the Gaocha extract, could inhibit endothelin receptors and thus prevent the excessive production of ET in SHRs.
PGl2 is an important prostaglandin member that could inhibit platelet activation. It is biosynthesized in endothelial cells and plays a vital role in preventing the platelet plug formation through primary hemostatsis[16]. In addition, PGl2 could exert as a vasodilator to stabilize cardiovascular homeostasis[47]. Physiologically, NO, PGI2, ET, and AngII are in a state of balance. When pathological factors disrupt this equilibrium of active substances in VEC, it can cause platelet activation and endothelial cell damage, which can contribute to or aggravate the occurrence of cardiovascular disease[14−16]. Recent research has revealed that endothelial damage and dysfunction are major factors in the onset and advancement of hypertension[48,49].
IL-6 and TNF-α are pro-inflammatory factors that can cause direct damage to vascular endothelium. TNF-α can disrupt the structure and function of vascular endothelial cells, leading to inflammatory reactions in the vascular wall[14]. IL-6 is a glycoprotein mainly produced by T cells and B cells, and it is involved in the regulation of immune response, acute phase response, and hematopoietic function. Additionally, IL-6 can activate platelet activating factors, which can enhance platelet aggregation and form thrombosis, thus damaging the endothelial cells of hematopoietic blood vessels[15].
The present study demonstrated that the Gaocha extract is capable of effectively reducing blood pressure in SHR rats, as well as decreasing the serum levels of Ang II and ET, while increasing the levels of NO and PGI2. This protective effect on vascular endothelial function is thought to be achieved by obstructing the activation pathway of RAS, thereby diminishing the amount of Ang II produced and its consequential damage to the vascular wall. Additionally, the Gaocha extract may bind to ET and receptors in vascular endothelial cells, resulting in the release of NO and PGI2, which leads to the relaxation of blood vessels and a decrease in blood pressure. The findings hold important clinical implications for the treatment of hypertension, suggesting that Gaocha extract may have the potential to serve as a natural alternative with potentially fewer side effects and greater safety compared to conventional chemical drugs. Furthermore, the study underscores the need for further research on Gaocha to elucidate its active components and associated mechanisms. Subsequent research will seek to identify the active components of Gaocha and explore their potential for treating hypertension using vascular endothelial cell lines. This will involve evaluating changes in pertinent biomarkers and utilizing Western blot or PCR techniques to assess the influence of these active components on gene and protein expression levels relevant to vascular activity, as well as their regulatory effects on genes linked to endothelial cell function. Meanwhile, conducting animal studies can provide insights into the effects of Gaocha extract on vascular endothelial function, vascular remodeling, inflammatory factors, and its regulatory influence on blood pressure. These efforts will enhance our understanding of the antihypertensive mechanisms of Gaocha extract and its potential for treating hypertension. Overall, the study highlights the promising antihypertensive effects of Gaocha extract and underscores the need for further research to unlock its full therapeutic potential for human health.
-
In this work, we discuss the effect of Gaocha extract treatment on spontaneously hypertensive (SHR) rats. Male SHR rats were randomly divided into the model (saline), positive control (captopril), low-dose Gaocha extract, medium-dose Gaocha extract and high-dose Gaocha extract groups, which were treated daily for two weeks. Tail artery systolic blood pressure (SBP) was measured before and 3, 7, 10, and 15 d after treatment. The Gaocha-treated groups had significantly lower SBP than the model group. Post-treatment abdominal aorta blood samples showed that the serum Angiotensin II, ET, IL-6, and TNF-α levels were significantly higher and the serum NO and PGI2 levels were significantly lower in the model group. Finally, haematoxylin-eosin staining of the heart and kidney showed that Gaocha extract had a protective effect. Thus, our findings indicate that Gaocha extract has obvious treatment benefits in SHR rats with regard to lowering SBP and protecting the vascular endothelium.
-
The animal study protocol was conducted in compliance with the Guide for the Care and Use of Laboratory Animals in Ministry of Science and Technology of the People's Republic of China. The protocol was reviewed and approved by the ethics committee of Anhui Agricultural University (Anhui, China).
-
The authors confirm contribution to the paper as follows: study conception and design: Wang H, Gao L; experiments performed: Wang W, Ma J, Ma Y, Bao Y, Long Z, Lei S, Xu Y, Dai Q; draft manuscript preparation: Wang H, Wang W. All authors reviewed the results and approved the final version of the manuscript.
-
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
This work was supported by the National Natural Science Foundation of China (Grant No. 31700608 and 32202551), Natural Science Foundation of Anhui Province (Grant No. 1708085MC58), the Natural Science Basic Research Program of Shaanxi (Grant No. 2022JQ-194) and the Anhui Agriculture University Pesident Fund (Grant No. 2014SKQJ020). We thank Professor Zijiang Long at the Function Experiment Center, Anhui University of Chinese Medicine for his assistance.
-
The authors declare that they have no conflict of interest.
-
# These authors contributed equally: Wenzhao Wang, Jing Ma
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Wang W, Ma J, Ma Y, Bao Y, Long Z, et al. 2024. Chemical constitutes and anti-hypertension potential of Gaocha (Acer ginnala Maxim) in spontaneously hypertensive rat. Beverage Plant Research 4: e011 doi: 10.48130/bpr-0024-0004
Chemical constitutes and anti-hypertension potential of Gaocha (Acer ginnala Maxim) in spontaneously hypertensive rat
- Received: 24 November 2023
- Revised: 03 January 2024
- Accepted: 17 January 2024
- Published online: 03 April 2024
Abstract: Gaocha is a kind of succedaneous tea beverage made by the tender shoots of Acer ginnala Maxim, widely distributed in China and Korea, which is used as a functional tea beverage with a long history dating back to 2000 BC. It has been used as folk medicine to depress blood pressure, however, little study has been reported on its antihypertensive mechanism. Thus, the present paper aimed at elucidating the bioactive constitutes in Gaocha extract and its effect of anti-hypertension on spontaneously hypertensive (SHR) rats. In this study, the phenolic compounds composition analysis showed that the total polyphenol content can reach 75.4 mg/g in dried Gaocha extract, and the content of epicatechin was 32.68 mg/g, indicating Gaocha may have many bioactivities. Therefore, in order to evaluate the anti-hypertension potential value of Gaocha extract, its effect on spontaneously hypertensive (SHR) rats was examined. Male SHR rats were randomly divided into the model (saline), positive control (captopril), low-dose Gaocha, medium-dose Gaocha and high-dose Gaocha groups. Every group was administered for 16 d. The results of tail artery systolic blood pressure (SBP) showed the Gaocha-treated groups had significantly lower SBP than the model group. Post-treatment abdominal aorta blood samples showed that the serum PGI2, Angiotensin II, ET, and NO levels in Gaocha-treated groups were significantly higher than these in the model group. After treating with Gaocha extract, the serum IL-6 and TNF-αlevels were significantly lower than these in the model group. The histomorphological examination of the heart and kidney also showed that Gaocha extract had a protective effect. Gaocha extract contained high levels of polyphenols with epicatechin as the predominant individual phenolic compound and has a significant improvement of anti-hypertension on spontaneously hypertensive (SHR) rats. The findings indicate that supplementation with Gaocha may contribute to preventing hypertension.
-
Key words:
- Gaocha /
- Spontaneously hypertensive rats /
- Anti-hypertension /
- Phenolic compounds.













