-
Nutraceuticals are defined as food-derived products that offer both nutritional and medicinal advantages, such as aiding in the management of specific health issues. The use of therapeutics derived from plant sources is expanding in the beverage industry. Several claims are made about the dermatological benefits of botanical products, including anti-inflammatory, anti-aging, and whitening properties[1]. The rationale for utilizing herbal remedies lies in their composition, which is exclusively derived from herbs and shrubs. These herbal medicines not only supply the body with essential nutrients and beneficial elements but also offer the assurance of having no adverse effects on human health due to their natural components[2]. Proliv Essence-3, also known as PE3, is a botanical beverage made from three selected fruits; Momordica charantia (bitter gourd), Malus domestica (apple), and Psidium guajava (guava). It was formulated primarily to help with the restoration of healthy skin and body as well as to boost overall health and wellness. Each of these components has a long history of being promoted as an alternative therapy for several types of skin conditions, such as skin radiance, reversing anti-aging signs, and the general improvement of internal health[3−5].
M. charantia (Cucurbitaceae), often known as bitter melon or bitter gourd, is a potent nutrient-dense plant made up of a complex array of advantageous substances, including flavonoids, tannins, steroids, triterpenes glycoside, phenolic acid, all of which impart a bitter flavour[6]. Furthermore, a study has revealed that M. Charantia effectively treats inflammatory skin conditions[3]. On the other hand, M. domestica (Roseaceae), or apple, is among the fruits renowned for its exceptional medicinal and therapeutic properties[7]. It is also rich in dietary fibre, polyphenols, and antioxidants[8]. The benefits of apples in skin renewal and anti-aging have been documented in several studies[5,9].
P. Guava (Myrtaceae) is commonly grown in tropical and sub-tropical regions[10]. It is high in antioxidants and phytochemicals, including vitamin C, saponin, flavonoids, oleanolic acid, quercetin and many other compositions[11]. According to Naseer et al., guava exhibited the ability to modify the activity of the heme oxygenase-1 protein, suggesting its possible use as an anti-inflammatory remedy for skin-related problems[12]. The aforementioned studies have shown that each of the components used in the formulation of PE3 has been proven to possess therapeutic benefits. To maximize the benefits of PE3 as an oral supplement, we recommend that individuals incorporate it into their daily routine as part of a balanced diet. Consuming PE3 regularly may offer the best potential for experiencing the intended benefits for skin health and overall well-being. Therefore, the purpose of the present work is to substantiate the effectiveness and safety of PE3 as a nutraceutical product, employing scientific approaches to bolster the acquisition of scientific insights concerning PE3 and its key components.
-
Naturemedics Laboratories Sdn. Bhd., Terengganu, Malaysia provided the PE3 extract for the study. The ingredients of PE3 consisted of M. charantia, M. domestica and P. Guava fruits. Initially, these fruits were processed into juices and then subjected to filtration using Whatman filter paper and a vacuum pump. To produce a liquid extract, the filtered juices were centrifuged to remove any clumps. For powder extract, the moisture content was removed through the freeze-drying method before concentrating it. Both liquid and powder of PE3 extracts were stored in an airtight container at 4 °C.
Proximate and nutritional analysis
-
The proximate compositions, including protein, ash, moisture, fat, total carbohydrate, and calorie content, were analysed by Lotus Laboratory Services (M) Sdn. Bhd., Malaysia. While, other nutritional parameters such as vitamins, minerals, dietary fibers, and total sugar were analysed by KHTP Bio Analytical Laboratory Sdn. Bhd., Malaysia. All the parameters were analysed following the methods outlined in the Association of Official Analytical Chemists (AOAC).
2,2-diphenyl-2-picrylhydrazyl (DPPH) assay
-
DPPH solution (Sigma-Aldrich, USA) was mixed with 100 μL of PE3 liquid extract at various concentrations (1−1,000 μg/mL) and further incubated for 30 min at room temperature[13]. In the control group, ascorbic acid (Qrec, New Zealand) was used with a concentration of 0 to 10 μg/mL. The absorbance was measured in triplicate at 515 nm using the ELx800 Absorbance Microplate Reader (BioTek Instrument, USA). The DPPH scavenging activity was calculated using the formula:
DPPH activity (%) = [(Ablank − Asample) / Ablank] × 100
'A' represents the absorbance reading of the DPPH scavenging activity as a function of sample concentration was plotted, and the half-maximal inhibitory concentration (IC50) value was obtained.
2,2'-azino-bis(3-ethylbenzthiazoline-6-sulphonic acid (ABTS) radical assay
-
To generate the ABTS+ cation radical, a reaction was initiated by mixing 7 mM ABTS (Sigma-Aldrich, USA) with 2.45 mM potassium persulfate (Sigma-Aldrich, USA) in a 1:1 ratio. At room temperature, the mixture was then placed in a dark environment for 12−16 h[14]. Subsequently, the ABTS solution was diluted with methanol to achieve an absorbance reading of 0.70 at 750 nm. Then, 30 μL of PE3 liquid extract was mixed with 300 μL of the ABTS solution, and the mixture was allowed to incubate for 6 min under dark conditions. Absorbance measurements were acquired in triplicate at a wavelength of 750 nm utilizing an ELx800 Absorbance Microplate Reader (BioTek Instrument, USA). Trolox (0−100 μg/mL) was employed as a positive control. To calculate the percentage inhibition, the following formula was used:
Radical scavenging activity (%) = [1 − (Asample / Acontrol)] × 100
'A' represents the absorbance reading and a graph depicting the relationship between radical scavenging activity and sample concentration was constructed, enabling the determination of the half-maximal inhibitory concentration (IC50) value.
High-performance liquid chromatography (HPLC)
-
Agilent's 1260 series HPLC equipment (Agilent Technologies Inc., USA), equipped with an autosampler and a UV detector using column (150 mm × 4.6 mm) Luna 5 μm C18 (Phenomenex, Torrance, USA) was used for the detection of gallic acid. The powder of PE3 extract was dissolved in deionized water at a concentration of 1,000 μg/mL. To establish the calibration curve, a broad range of concentrations for the gallic acid standard (20, 40, 60, and 80 μg/mL) was prepared in methanol. Elution was conducted through a gradient elution process, with a flow rate of 1.0 mL/min. Eluent A was composed of water (99.9%) and formic acid (0.1%), while eluent B consisted of acetonitrile (99.9%) and formic acid (0.1%). Initially, at 0.0 min, the elution mixture consisted of eluent A (80%) and eluent B (20%), and at 10.0 min, it transitioned to eluent A (55%) and eluent B (45%). Detection was performed at 280 nm utilizing a UV detector[15].
Tyrosinase inhibition assay
-
The study investigated the inhibitory potential of PE3 against mushroom tyrosinase. For the tyrosinase inhibition assay, L-DOPA was employed as the substrate[16]. Since tyrosinase is an enzyme responsible for catalyzing the conversion of L-DOPA into dopachrome, enzymatic activity was tracked by monitoring dopachrome formation at 410 nm in the presence of a potent tyrosinase inhibitor. The reaction mixture consisted of 20 μL of L-DOPA, 40 μL of mushroom tyrosinase (10 μg/mL), and 140 μL of PE3 liquid extract at varying concentrations (0.001−1,000 μg/mL). Ascorbic acid was utilized as a positive control. The dopachrome was determined by measuring the absorbance at 410 nm in triplicate, employing the ELx800 Absorbance Microplate Reader (BioTek Instrument, USA). The median inhibitory concentration (IC50) was calculated from the dose-response curves by nonlinear regression analysis using GraphPad Prism Version 6 software (United States). The enzymatic activity was calculated using the formula:
Enzyme inhibition activity (%) = [1 − (Asample / Acontrol) × 100], where A is the absorbance reading.
Elastase inhibition assay
-
The experiment was conducted following the methodology outlined by Era et al.[17]. To initiate the experiment, porcine pancreatic elastase was dissolved in a Tris-Cl buffer at pH 8.0 to create an enzyme solution with a concentration of 0.5 units/mL. This enzyme solution was employed to monitor the release of p-nitroaniline during the cleavage of the substrate N-succ-(Ala)3-nitroanilide (SANA). The substrate solution was prepared at a concentration of 0.1 M by dissolving SANA in Tris-Cl buffer. Subsequently, 130 μL of the enzyme solution was incubated with 10 μL of PE3 liquid for 5 min, followed by the addition of 15 μL of the substrate solution after the incubation period. As a positive control, epigallocatechin gallate (EGCG) was utilized. The enzymatic activity was quantified by measuring the absorbance at 410 nm using an ELx800 Absorbance Microplate Reader (BioTek Instrument, USA). The median inhibitory concentration (IC50) was calculated from the dose-response curves by nonlinear regression analysis using GraphPad Prism Version 6 software (USA). The enzymatic activity was calculated using the formula:
Enzyme inhibition activity (%) = [Acontrol − Asample) / Acontrol × 100], where A is the absorbance reading.
In vitro scratch assay
-
The experiment aimed to assess in vitro cell migration using a protocol adapted from Fronza et al.[18]. Initially, the human skin fibroblast cell line HSF1184 (ATCC CL-193, USA) was seeded into a 12-well microplate and allowed to incubate for 24 h. After this incubation period, a uniform cell monolayer was formed and then a linear scratch was carefully created. Subsequently, different concentrations of PE3 extract were applied to the scratched area. As a positive control, platelet-derived growth factor (PDGF) was employed. The width of the scratch and the cell migration activity of HSF1184 were measured in triplicate.
Epidermal growth factor (EGF) cytokine analysis
-
The cytokine analysis was evaluated using an enzyme-linked immunosorbent assay (ELISA) (InvitrogenTM, USA) based on the manufacturer's procedure[18]. The cell lysate (0.001−10,000 μg/mL) was collected for cytokine analysis following 24 h of exposure to PE3 extract. EGF standard was prepared at various concentrations of 3.9 to 250 μg/mL. The standards were reconstituted using Standard Diluent Buffer before being subjected to a 10-min incubation period. Both the standards and the PE3 sample were allowed to incubate for 2 h at room temperature. Next, the mixture was combined with 100 μL of biotin conjugate, and this new mixture was further incubated for an additional hour at room temperature. Then, 100 μL of Streptavidin-HRP was added and the mixture was incubated for another 30 min before adding 100 μL of Stabilized Chromogen. Lastly, 100 μL of stop solution was added to the well. The color changes were observed and the absorbances were taken at 450 nm using ELx800 Absorbance Microplate Reader (BioTek Instrument, USA). Relative cytokine concentration was determined using the formula:
Relative cytokine concentration (%) = (Asample / A control) × 100, where A is the absorbance reading.
Neutral red uptake assay
-
A cytotoxicity test was conducted using the methodology described by Zofia et al.[19]. Two cell lines, HSF1184 (ATCC CL-193, USA) and BALB/c 3T3 (ECACC 90011883, UK), were cultured at 37 °C in a humidified incubator. The culture medium used was Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum, 5% fetal calf serum, and 1% Penicillin-Streptomycin. All the chemicals used in the experiment were procured from Gibco, Scotland. A density of 1 × 105 cells/mL of fresh media was used to seed the cells in 96-well plates. The medium was aspirated after the initial 24 h of preculture, and various concentrations of PE3 extracts (0.001−10,000 μg/mL) were applied to each well. The cultures were then continued for an additional 24 h. The control group was unexposed cells. Following exposure to the PE3 extract, the cells were incubated for 2 h with neutral red dye. After the treatment, the cells were washed with Phosphate Buffered Saline (PBS), and subsequently, 150 μL of a destaining solution containing ethanol and acetic acid was added to each well. The mixture was gently shaken for 10 min to facilitate the removal of neutral red from the cells, resulting in a homogeneous solution. For each concentration of the extract, triplicate tests were performed, and the absorbances were measured at 540 nm using the ELx800 Absorbance Microplate Reader (BioTek Instrument, USA). The IC50 value was then determined by creating a graph and utilizing GraphPad Prism Version 6 software (USA). The in vivo LD50 of acute oral toxicity was estimated from in vitro IC50 using the formula, logLD50 = 0.372 × logIC50 (μg/mL) + 2.024[20]. While relative cell viability was calculated using the formula:
Relative cell viability (%) = (Asample / Acontrol) × 100, where A is the absorbance reading.
Statistical analysis
-
The data were presented as mean ± standard deviation (SD), and the level of significance was assessed at a 5% probability (p < 0.05). Data analysis was performed using GraphPad Prism software (version 6, USA), and a one-way analysis of variance (ANOVA) was utilized for statistical evaluation.
-
Based on the proximate analysis in Table 1, high moisture content and low calories of PE3 were observed. The value of pH was slightly acidic, with an indication of bitter and sour taste. The nutritional analysis revealed a PE3 profile characterized by limited quantities of essential nutrients such as vitamin C, vitamin D, and dietary fiber. Additionally, the assessment identified a presence of 11 mg of calcium, a moderate level of 3 mg of iron, and 58 mg of potassium. The sodium content is moderate, warranting careful monitoring for individuals with specific health considerations.
Table 1. The proximate and nutritional analysis of PE3 extract.
Parameter Values (per 100 g) Protein, % (w/w) ND Calories, kcal/100 g 5.0 Carbohydrate, % (w/w) 7.0 Saturated fat, % (w/w) ND pH 4.45 Taste Bitter and sour Colour Brown Moisture, % (w/w) 92.4 Ash, % (w/w) 0.6 Dietary fibre (g) 0 Total sugar (g) 12.6 Vitamin C (mg) 0 Vitamin D (μg) 0 Calcium (mg) 11.0 Iron (mg) 3.0 Sodium (mg) 46.0 Potassium (mg) 58.0 ND = Not detected. Detection of gallic acid
-
The chromatographic fingerprint of the PE3 extract is shown in Fig. 1. Peak A indicated the presence of gallic acid in PE3 as it exhibited identical spectral characteristics and retention time with the standard marker, gallic acid. The concentration of gallic acid was estimated based on the linear calibration curves of the standard compound, and the result yielded 11.21 ± 0.07 mg/g of gallic acid.
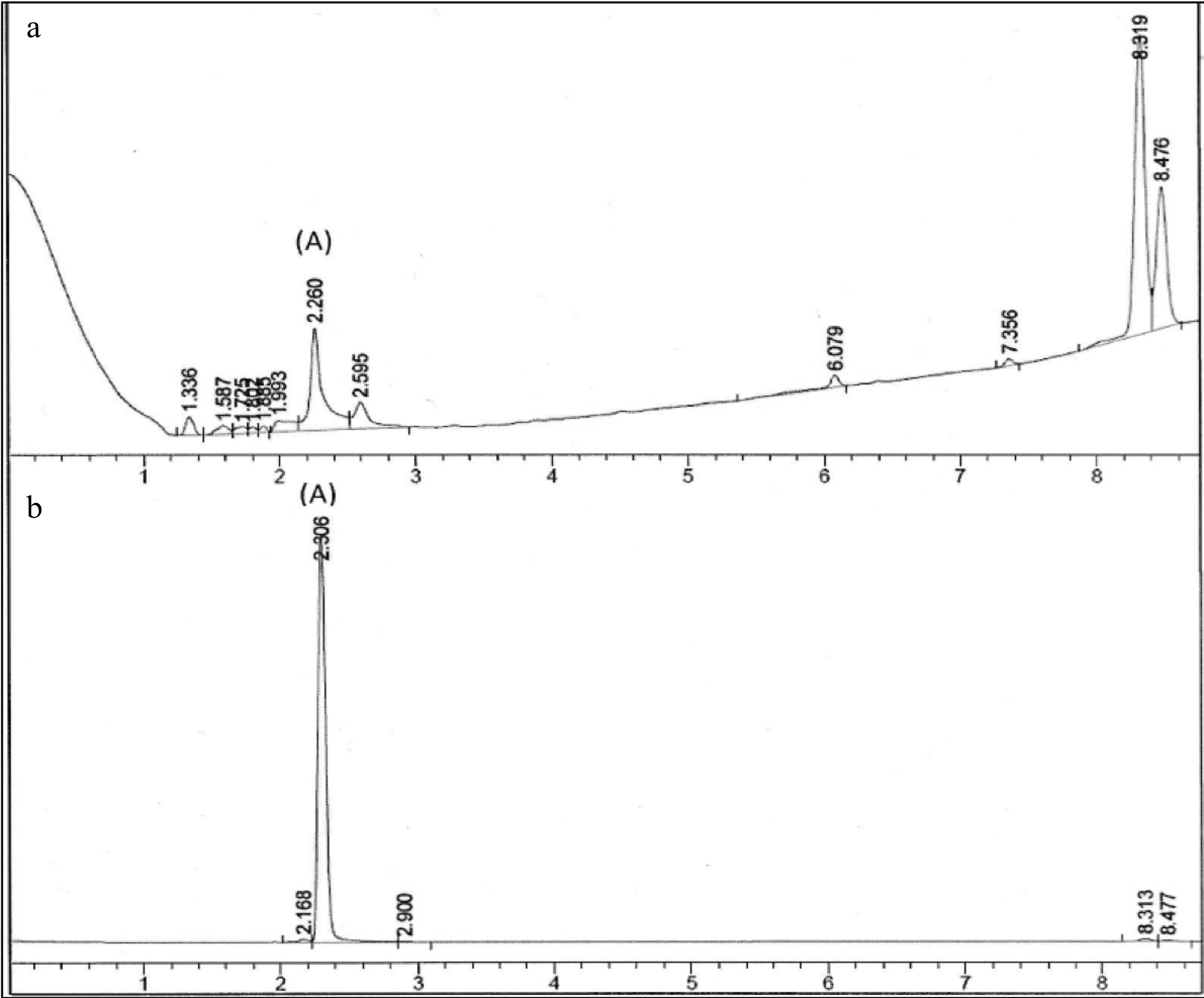
Figure 1.
Chromatographic fingerprint of (a) PE3 and standard marker, (b) gallic acid. Peak A indicated the presence of gallic acid.
Antioxidant activity
-
As shown in Table 2, the IC50 for PE3 extract was 148.60 ± 1.52 μg/mL, and the IC50 for ascorbic acid was 6.50 ± 1.04 μg/mL for DPPH assay. Based on the ABTS assay, PE3 extract exhibited 91.18 ± 1.15 μg/mL of IC50, while Trolox showed 6.07 ± 1.07 μg/mL of IC50. The scavenging activity percentage increased with concentrations of PE3 in both the DPPH and ABTS assays (Fig. 2). Interestingly, at the concentration of 10 μg/mL, PE3 produced 80% DPPH scavenging activity, while ascorbic acid had 90% of scavenging activity at the concentration of 1000 μg/mL. In the ABTS assay, 1,000 μg/mL of PE3 produced 14% scavenging activity, while Trolox revealed 90% scavenging activity at 10 μg/mL concentration.
Table 2. Antioxidant activity, elastase assay and tyrosinase assay of PE3 extract.
Assays Values Gallic acid content (mg/g) 11.22 ± 0.07 DPPH (IC50 μg/mL) 148.60 ± 1.52 ABTS (IC50 μg/mL) 91.18 ± 1.15 Tyrosinase assay (IC50 μg/mL) 160.20 ± 1.81 Elastase assay (IC50 μg/mL) 65.49 ± 0.38 All values represent the mean ± standard deviation of three replicates analyses. 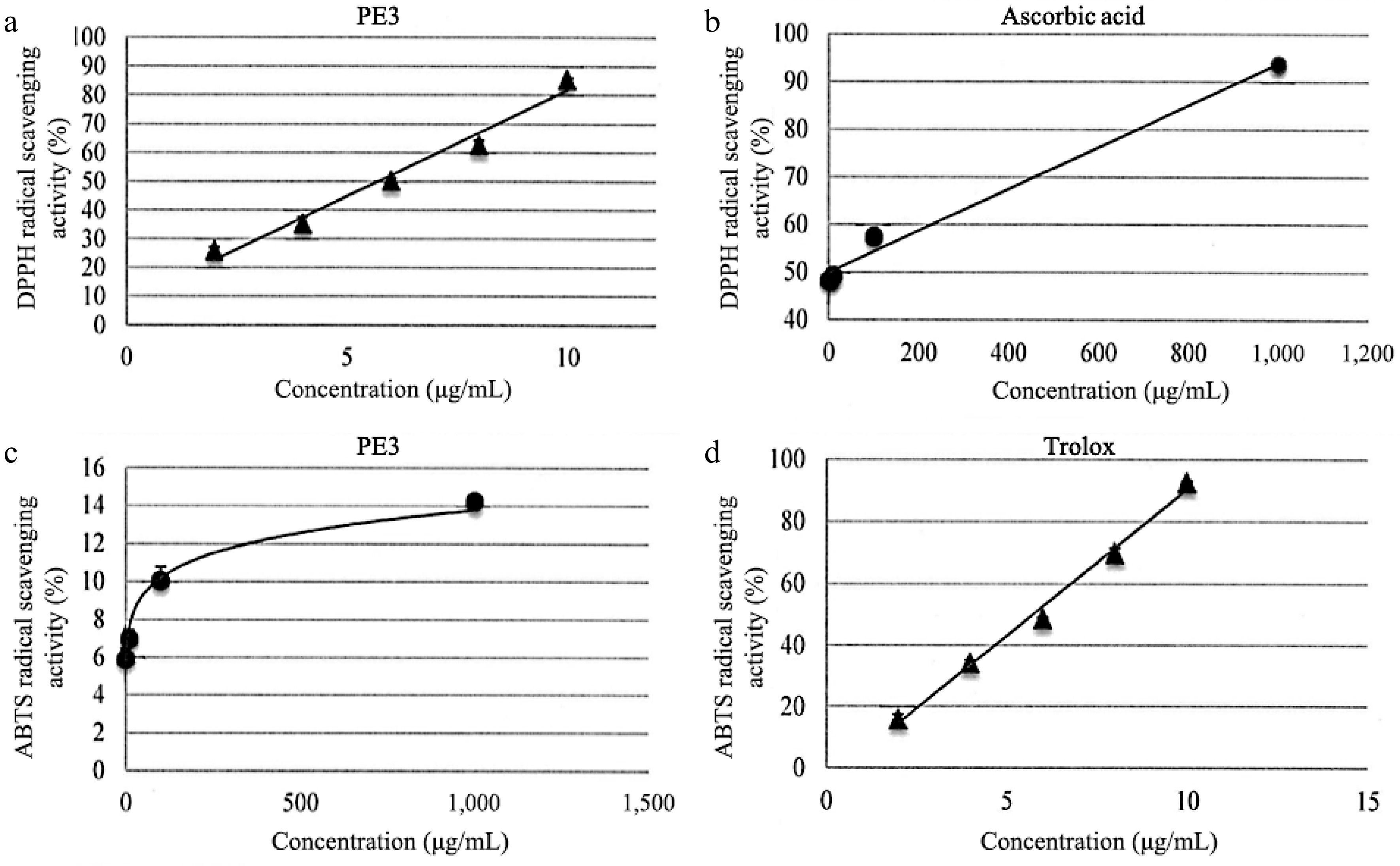
Figure 2.
DPPH scavenging activity of (a) PE3 extract and the standard, (b) ascorbic acid. ABTS scavenging activity of (c) PE3 extract and the standard, (d) Trolox. All values represent mean ± standard deviation.
Skin anti-aging and whitening properties
-
Skin whitening and rejuvenation properties of PE3 extract were evaluated using mushroom tyrosinase and elastase inhibition assays (Table 2). In the mushroom tyrosinase test, PE3 extract showed the highest significant inhibition (p < 0.05) at the concentration of 1,000 μg/mL (16%). PE3 extract also had an IC50 value of 160.20 ± 1.81 μg/mL, which was lower than the standard ascorbic acid (17.36 μg/mL). In the elastase inhibition activity assay, the enzyme was inhibited in a dose-dependent manner up to 1,000 μg/mL, which showed the highest inhibition (23%) of PE3 at the concentration of 1,000 μg/mL compared to the EGCG (33%). PE3 extract also exhibited a low IC50 value (65.49 ± 0.38 μg/mL), indicating a strong elastase inhibition capacity.
Skin cell growth and rejuvenation
-
The role of EGF in regulating the proliferation of HSF1184 cells was further investigated in cytokine analysis (Fig. 3). In HSF1184 cells treated with PE3, a substantial increase in the secretion of EGF was observed, ranging from 110% to 170% relative cytokine viability. As the extract concentration increased, the relative cytokine viability increased. The cell migration of fibroblast for PE3-treated cells (0.001−1,000 μg/mL) was approximately 100%, comparable to the cells treated with PDGF (Fig. 4). Cells with untreated medium showed 60 to 70% of migration activity. The confluent monolayer of HSF1184 cells treated with PE3 extract demonstrated a fast wound closure after 24 h, showing this extract is notably an efficient wound healing therapy.
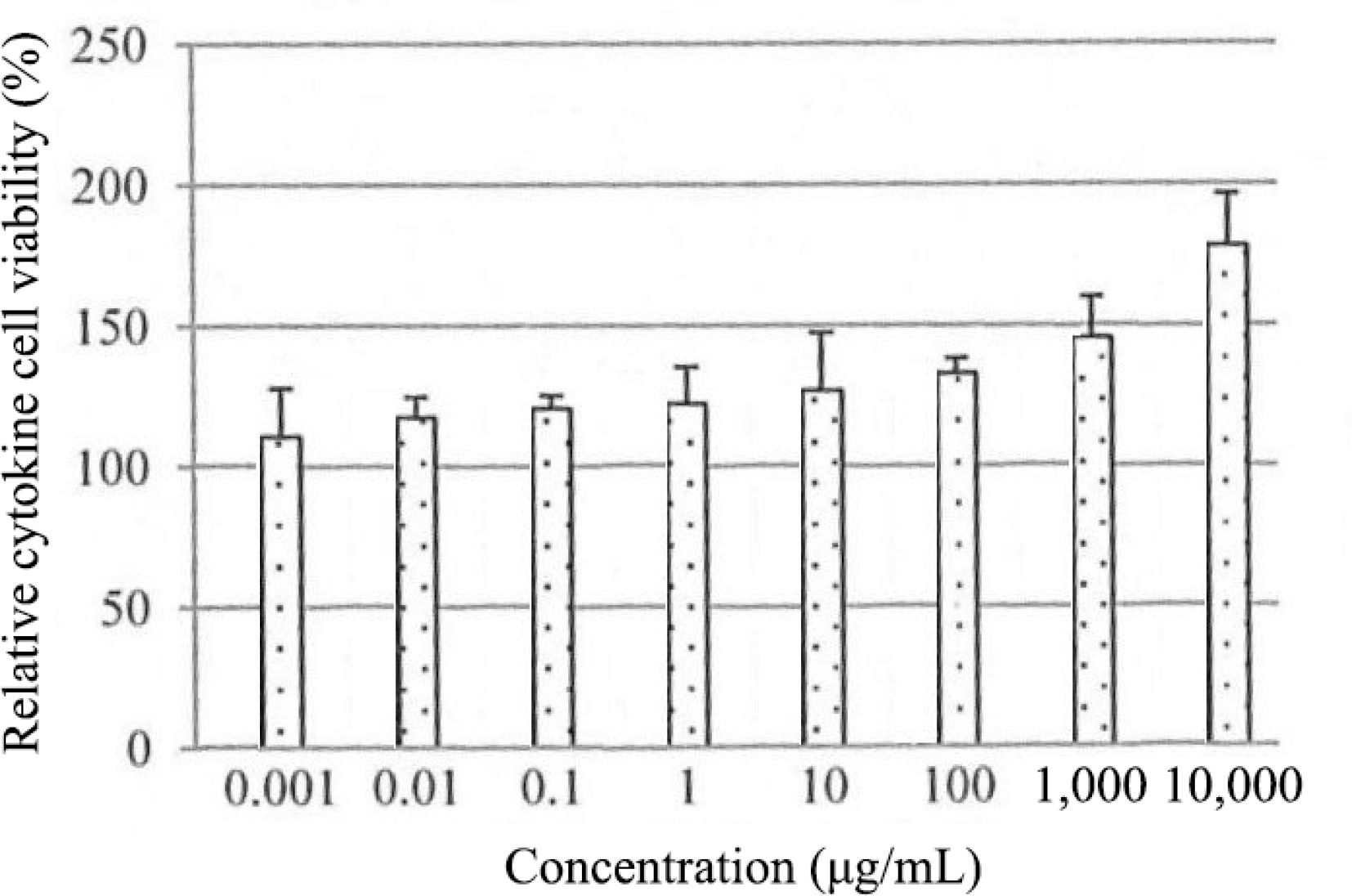
Figure 3.
Effect of PE3 extract on EGF secretion of HSF1184 cell within 24 h. Results are expressed as mean ± standard deviation.
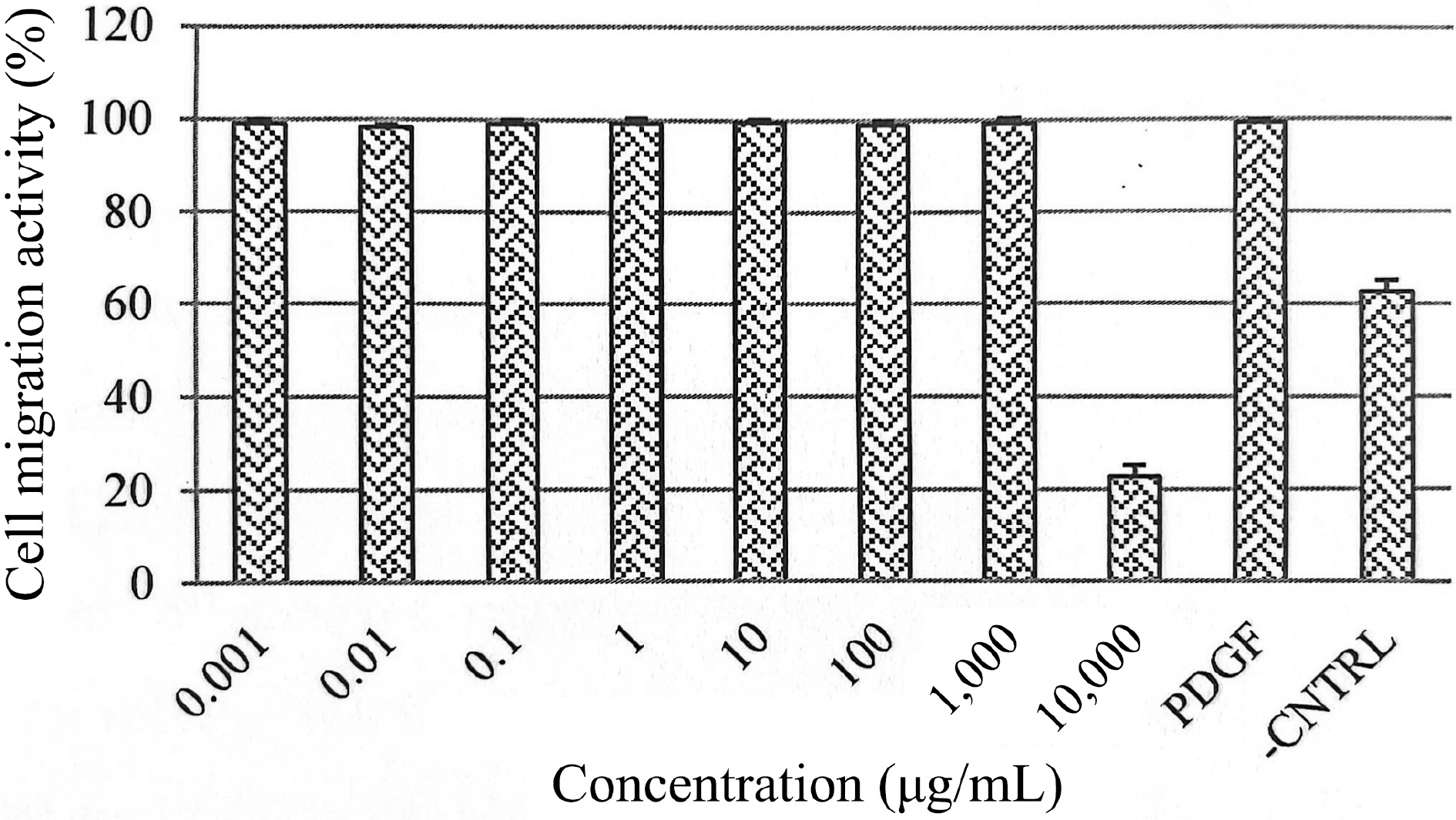
Figure 4.
Effects of PE3 extract at different concentrations on HSF1184 cell migration. Results are expressed as mean ± standard deviation.
Cytotoxicity test
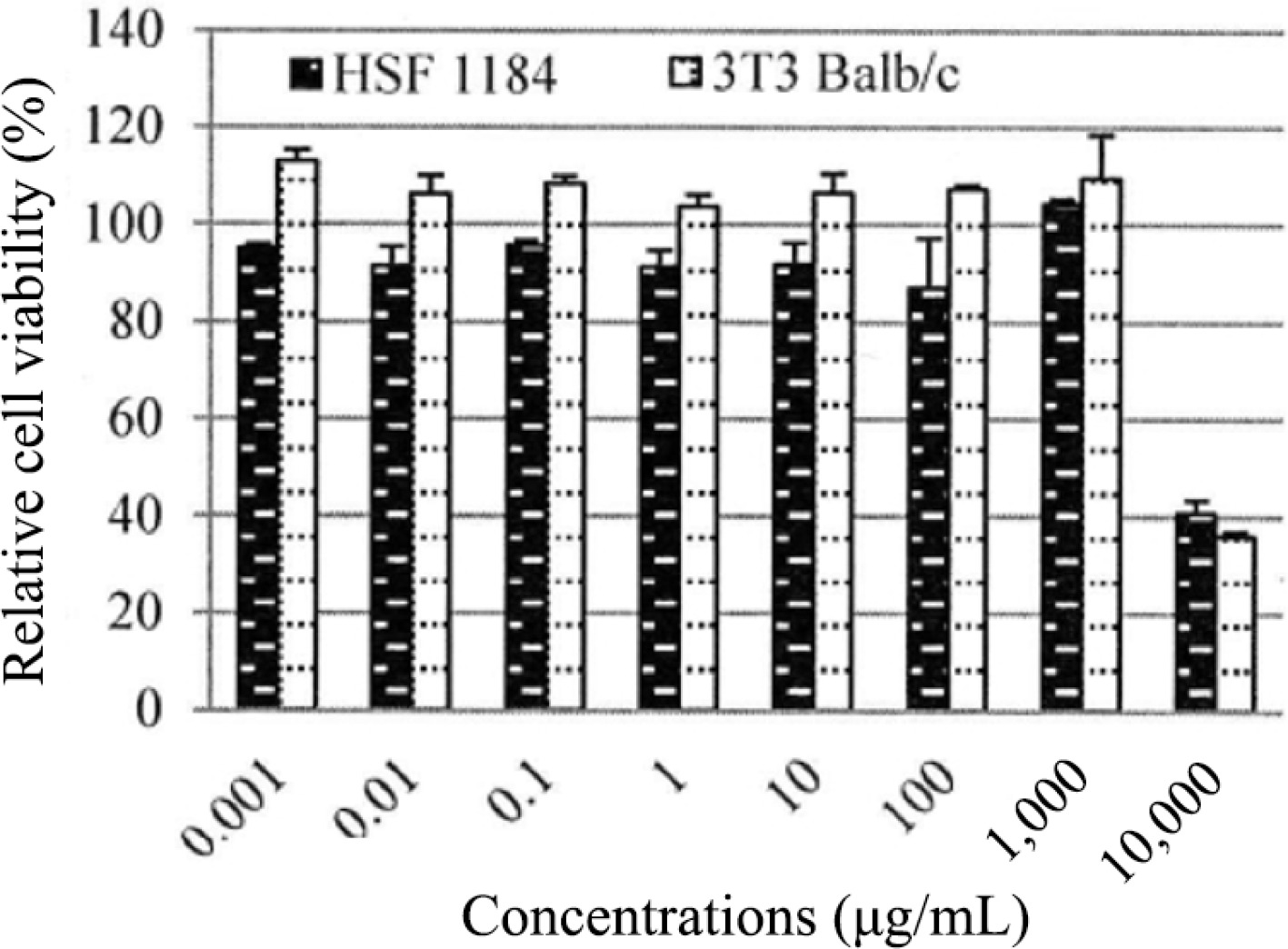
The biological activity of PE3 extract was also investigated on cells as an in vitro model. The test showed that PE3 extract was non-cytotoxic at concentrations below 1,000 μg/mL when exposed to HSF1184 and BALB/c 3T3 cell lines (Fig. 5). The result showed a consistent pattern in the mean of relative cell viability on both cell lines. Generally, the BALB/c 3T3 cell line produced higher cell viability than HSF1184 in all concentrations tested. The figure also shows that at 0.001 μg/mL (BALB/c 3T3) and 1,000 μg/mL (HSF1184), PE3 extract exhibited more than 100% cell viability in a dose-dependent manner for 24 h. Cell damage was observed at a concentration of 10,000 μg/mL, indicating that this dose is considered toxic when administered at levels exceeding 10,000 μg/mL. The calculated IC50 values were 3,141 μg/mL (HSF1184) and 3,145 μg/mL (BALB/c 3T3). Meanwhile, the calculated LD50 values were 2,113.05 μg/mL and 2,114.05 μg/mL for HSF1184 and BALB/c 3T3, respectively.
-
The demand for natural and plant-based nutricosmetics has consistently surged in recent years, driven by a growing awareness of the pivotal role these ingredients play in promoting health and beauty[21]. However, despite the considerable availability of natural nutricosmetic ingredients in the market, scientific validation and the tolerance profiles of many of these components remain limited. Preclinical trials are imperative to substantiate their effectiveness[22]. The increasing interest in nutrition that promotes both internal well-being and external appearance is also heightening consumer awareness. Consequently, this study holds great significance, given that PE3, as a dietary beverage, has the potential to provide essential nutrients to the body while also improving skin health concurrently. Its means of consumption are both straightforward and cost-effective, and its safety is assured due to its nature-derived and readily available raw materials.
Based on the proximate composition, PE3 extract has a unique flavour profile, comprising mainly bitter and sour acidic taste, as reflected in the low pH of the drink, making it suitable for bodily hydrating features. The sour taste in the formulation was attributed to the guava and green apple juices, while the bitter taste was derived from the bitter gourd juice. Research has shown that the sour and bitter taste may aid digestion, improve appetite, and help to keep sugar cravings at bay[23]. The nutritional analysis of PE3, focusing on its impact on skin health, indicated some noteworthy aspects. The degradation or loss of vitamins C and vitamin D in this study could be related to the processing method as these vitamins are sensitive to heat and light. Other than that, the formulation of the PE3 supplement, including how the ingredients are combined and processed, can influence the nutrient content. If certain ingredients or processing steps are chosen that lead to the loss of these vitamins, it would be reflected in the analysis results. While PE3 is found to be lacking in certain essential nutrients like vitamin C and vitamin D, which play crucial roles in skin health, it does contain moderate levels of beneficial minerals such as calcium, iron, and potassium. The presence of these minerals can contribute to overall skin well-being, supporting aspects like collagen production, oxygen transport, and cellular function[24]. The moderate sodium content in PE3 highlights the need for careful monitoring.
Previous studies have found gallic acid in M. charantia, M. domestica, and P. guava, which are the main ingredients of PE3, hence the presence of gallic acid in the PE3 concoction was verified in this study[25,26]. Furthermore, following extensive research, gallic acid has demonstrated significant antioxidant and anti-inflammatory capacities, with its most prominent advantages often linked to skin health. Zhang et al. validated the clinical efficacy of gallic acid in reducing free radical generation, controlling skin inflammation, inhibiting tyrosinase production, and preventing melanin transfer[27]. Therefore, gallic acid was selected as a targeted compound of interest in this study. Several studies have documented the quantification of gallic acid in herbal formulations. For example, in the research conducted by Borde et al.[28], 30 Ayurvedic herbs and formulations were screened for gallic acid by silica gel thin layer chromatography, and they found that nine Ayurvedic herbs contained gallic acid in the range of 0.091 to 27.36 mg/g. In a related study, the amount of gallic acid in the polyherbal Triphala churna formulation was determined using HPLC, and they discovered that it was present at a level of approximately 3.27% w/w which is lower than the findings of the present study[29]. The pharmacological features of the product, such as its anti-inflammatory, antibacterial, antioxidant, and antimutagenic actions, may have been influenced by the presence of gallic acid in the herbal formulation[30].
The antioxidant activity of PE3 was first assessed using the DPPH free radical scavenging assay. DPPH, which stands for 2,2-diphenyl-1-picrylhydrazyl, is a stable free radical molecule. It absorbs light in the range of 515 to 520 nm in its oxidized state. The DPPH test is a rapid and efficient method for evaluating free radical scavenging activity. DPPH can accept either an electron or a hydrogen radical, leading to the formation of a stable diamagnetic molecule[14]. This transformation results in a change in color from purple to yellow, indicating a decrease in DPPH radical absorption. The reducing power was determined by calculating the concentrations required to achieve 50% inhibition (IC50), representing the amount needed to scavenge 50% of DPPH free radicals. The lower the value of IC50, the higher the scavenging power of the antioxidant.
The DPPH solution undergoes reduction due to the presence of antioxidant compounds that contain hydrogen-donating groups, such as phenolic acids. This reduction leads to the formation of non-radical compounds. A study exploring the antioxidant properties of herbal tea revealed that the extracts displayed DPPH radical inhibition, with a range of 21.8% to 44.48%[31]. In contrast, the PE3 extract in our current study exhibited even higher antioxidant activity, ranging from 25 to 80%. Meanwhile, the IC50 value determined in our current study indicates that PE3 exhibits a significantly higher antioxidant capacity in comparison to the previous study conducted by Fatanah et al.[32]. In their research, they reported an IC50 value of 1055 μg/mL for herbal tea made from C. caudatus leaves.
To support the DPPH results, we conducted the ABTS assay, which is based on the decolorization of the blue-green color of the ABTS solution. This assay effectively demonstrates antioxidant capacity by reducing ABTS+ to ABTS. Our findings indicate that the PE3 extract serves as a potent source of antioxidants, capable of scavenging both ABTS and DPPH free radicals. The presence of gallic acid in the sample likely contributes to the observed antioxidant potential in PE3. Gallic acid, a phenolic compound commonly found in plants, plays an important role in preventing the detrimental effects of oxidative stress[33]. By mitigating oxidative stress, antioxidants can effectively slow down the skin aging process, diminishing the appearance of fine lines, wrinkles, and age spots, ultimately preserving a more youthful complexion[34].
Tyrosinase and elastase are pivotal enzymes in the context of skin aging. Tyrosinase plays a central role in melanin synthesis, influencing skin pigmentation. Overactivity of tyrosinase can lead to hyperpigmentation, contributing to age-related skin discoloration[35]. Addressing tyrosinase activity is a common strategy in skincare to manage uneven skin tone. On the other hand, elastase is responsible for breaking down elastin, a protein critical for maintaining skin elasticity and firmness. Increased elastase activity is associated with the degradation of elastin fibers, resulting in the loss of skin elasticity and the formation of wrinkles and sagging skin[36]. Therefore, inhibiting elastase is a targeted approach in skin-care formulations to preserve skin elasticity and mitigate visible signs of aging, contributing to a more youthful and resilient complexion.
Theoretically, tyrosinase will catalyse the production of brown pigment using L-dopa as a substrate. The capacity of the sample to inhibit the oxidation process will result in a reduction in the intensity of the brown pigment colour, which can be detected using a spectrophotometric approach[37]. The inhibitory activity of PE3 is comparable to what has been reported for the herbal formulation containing C. palala and U. micrantha, as well as their mixture, all of which exhibited less than 20% of tyrosinase inhibition[38]. Meanwhile, for elastase assay, as reported by Zofia et al.[19], several plants from the Apiaceae family revealed inhibition elastase activities ranging from 15% to 30% for 0.5% concentration of plant extracts. This action is primarily due to the presence of a significant amount of a variety of biologically active substances, such as polyphenols, which comprise, among other things, flavonoids, phenolic acids, tocopherols, and tannins[33]. In a recent study carried out by Raphaelli et al., the phenolic extracts of M. domestica showed potential uses in the treatment of melanoma and protection against harmful UV radiation which are ideal for the formulation of skin-related products[39].
The ability of PE3 extract to stimulate fibroblast proliferation and cell migration in enhancing wound healing activity was demonstrated using an in vitro wound scratch test. Skin fibroblast is one of the most crucial cells in tissue repair as its proliferation is involved in migration, contraction, and collagen production. Based on the results observed in this study, PE3 was proven to enhance wound healing. Significant secretion of EGF was observed in this study, indicating that it could promote cell proliferation and thus enhance skin rejuvenation. The wound-healing effect of PE3 could be related to the presence of gallic acid in the sample, which could induce fibroblast cell growth and proliferation to enhance the wound closure activity[40]. As supported by the previous study, gallic acid was found to increase fibroblast cell counts and decrease late inflammation along with increased TGF-β expressions in the wound healing process of Wistar rats[41].
The HSF1184 cell line is derived from the human skin fibroblast. Whereas, the BALB/c 3T3 cell line is a mouse fibroblast cell line that is widely used in cytotoxic studies. Fibroblasts are a type of connective tissue cells that play a crucial role in tissue repair and maintenance. BALB/c 3T3 cells, being fibroblasts, are often used as a representative cell type for studying cellular processes related to connective tissues[42]. Both HSF1184 and BALB/c 3T3 cells were used in the present study to ensure the reliability and increase robustness of the study's findings. Different cell lines have unique characteristics and responses to various substances, and using multiple cell lines assists in obtaining a more comprehensive understanding of the potential cytotoxic effects of PE3 extract.
PE3 extract exhibited no cytotoxic effects at concentrations below 1,000 μg/mL when tested both on HSF1184 and BALB/c 3T3 cell lines. If the non-cytotoxic effects are consistent across different cell lines, it strengthens the argument that the observed effects are not specific to a particular cell type but may have broader implications. The IC50 value for PE3 was considered high, which typically indicated low cytotoxicity[43]. Extracts with high IC50 values are preferable for work because they have fewer toxic effects on host cells. In a prior study, it was found that the leaf extract of R. communis displayed an IC50 value of 784 μg/mL when tested against the human Caucasian skin fibroblast cell line (Bud-8)[44]. In another study, a 5% concentration of Aegopodium podagraria extract demonstrated the highest proliferation, reaching 100% cell viability when treated with human foreskin fibroblast (BJ) cell lines for 24 h[19].
LD50 was also predicted in this study to assess the safety of PE3 extract. Traditionally, the safety and toxicity profiles are determined via in vivo acute oral toxicity tests, where the lethal dose for 50% of the tested animals (LD50) will be calculated[45]. However, in line with the recommendations of the Interagency Coordinating Committee on the Validation of Alternative Methods, the LD50 value for rats can be estimated using the lC50 value obtained from an in vitro cytotoxicity assay, employing a previously established regression formula. This innovative approach allows us to approximate a starting dose much closer to the actual LD50 value for subsequent in vivo acute oral toxicity studies. As a result, this approach not only reduces the number of animals required for preliminary range-finding experiments but also minimizes the unnecessary need to sacrifice animals due to toxicity concerns[46].
The research had limitations that involved a restricted focus primarily on in vitro assays, a lack of human clinical trials to substantiate the asserted health benefits, and an absence of extended-term studies to evaluate sustained effectiveness and safety. Future investigations may focus on the clinical trials for substantiating health claims, performing thorough chemical analyses, exploring long-term implication, and refining the formulation of the drink. While the present study provided valuable insights into the proximate composition, antioxidant activity, and biofunctionalities of PE3, it is also important to include further exploration of standardization on the formulated extract. Addressing these aspects comprehensively in future research is essential to ensure transparency, and further validate the quality and efficacy of PE3 in terms of its major constituents. Furthermore, continuous safety assessments are imperative to confirm its non-cytotoxic nature.
-
Botanical nutraceuticals are an efficient approach for sustaining health and combating nutritionally related illnesses, promoting optimal health, and quality of life. The findings from this study revealed that PE3 can serve as a botanical beverage due to the presence of bioactive ingredients, and its good antioxidant capacity which may aid in skin whitening and rejuvenation, wound healing activity, and overall consumer health. PE3 presents consumers with a cost-effective and practical alternative that not only offers a delightful flavor but also potential health benefits, making it a promising candidate to replace synthetic antioxidant beverages available in the market.
Author contributions
-
The authors confirm contribution to the paper as follows: study conception and design: Abdul Majid FA; experiment conduction and data analysis: Abdul Hamid SN; draft manuscript preparation and revision: Zakaria NH; manuscript revision: Fadhlina A, Anuar MNN, Tengku Aziz TN. All authors reviewed the results and approved the final version of the manuscript.
Data availability
-
All data generated or analyzed during this study are included in this published article.
Acknowledgement
-
The authors would like to acknowledge the Faculty of Chemical Engineering and Natural Resources, Universiti Teknologi Mara for providing the facilities to conduct the research and the Institute of Climate Adaptation and Marine Biotechnology, Universiti Malaysia Terengganu for the support. The authors are also thankful to Naturemedics Laboratories for providing the source material.
-
The authors declare that they have no conflict of interest.
- Copyright: © 2024 by the author(s). Published by Maximum Academic Press, Fayetteville, GA. This article is an open access article distributed under Creative Commons Attribution License (CC BY 4.0), visit https://creativecommons.org/licenses/by/4.0/.
-
About this article
Cite this article
Zakaria NH, Abdul Majid FA, Fadhlina A, Abdul Hamid SN, Anuar MNN, et al. 2024. Proliv Essence-3 (PE3): a nutricosmetic botanical blend as a dietary beverage for skin wellness and general health. Beverage Plant Research 4: e016 doi: 10.48130/bpr-0024-0008
Proliv Essence-3 (PE3): a nutricosmetic botanical blend as a dietary beverage for skin wellness and general health
- Received: 07 September 2023
- Revised: 26 December 2023
- Accepted: 25 January 2024
- Published online: 07 May 2024
Abstract: A natural product-based dietary approach could offer a safe and effective method for slowing down or preventing age-related deterioration in skin appearance and function, including issues like hyperpigmentation, dryness and wrinkles. Proliv Essence-3 (PE3) is a botanical beverage, meant for oral consumption with a unique formulation of three selected fruit extracts, Momordica charantia, Malus domestica, and Psidium guajava. Its purpose is to enhance both skin health and overall well-being from within. Proximate composition, presence of gallic acid and antioxidant activity of PE3 extract were determined. Elastase and tyrosinase inhibition assays were employed to investigate the anti-aging and whitening effects, respectively. The in vitro scratch assay and epidermal growth factor (EGF) assay were carried out to evaluate skin cell growth promotion and rejuvenation. The cytotoxicity analysis was carried out via neutral red uptake. The proximate analysis revealed that the product had a high moisture content and low amounts of calories. High-performance liquid chromatography (HPLC) analysis estimated 11.22 mg of gallic acid in 1 g of PE3 extract. PE3 exhibited a DPPH-IC50 value of 148.60 ± 1.52 μg/mL and an ABTS-IC50 value of 91.18 ± 1.15 μg/mL. The IC50 values for tyrosinase and elastase inhibition assays were 160.20 ± 1.81 μg/mL and 65.49 ± 0.38 μg/mL, respectively. PE3 was also discovered to be non-cytotoxic, and it enhanced the migration and proliferation of HSF1184 cells. EGF secretion was detected in PE3-treated HSF1184. This study provided preliminary evidence supporting the potential of PE3 as a nutricosmetical botanical beverage for promoting skin beautification and general health.
-
Key words:
- Proliv Essence-3 /
- Botanical beverage /
- Nutraceutical /
- Skin beautification /
- Anti-aging